Abstract
Oral squamous cell carcinoma (OSCC) has high recurrence and mortality rates despite advances in diagnosis and treatment. Therefore, it is necessary to identify new biomarkers for early detection, efficient monitoring, and prognosis prediction. Since microRNA (miRNA) is stable and detectable in serum, it has been reported to inform the diagnosis and monitor disease progression through liquid biopsy. In this study, a circulating specific miRNA panel in OSCC patients was developed, and its usefulness as a dynamic monitor was validated. Small RNAs were extracted from the serum of OSCC patients (n = 4) and normal controls (n = 6) and profiled using next-generation sequencing. NGS identified 42 differentially expressed miRNAs (DEmiRNAs) in serum between patients with OSCC and healthy controls, with threefold differences (p < 0.05). Combining the 42 DEmiRNAs and The Cancer Genome Atlas (TCGA) databases OSCC cohort, 9 overlap** DEmiRNAs were screened out. Finally, 4 significantly up-regulated miRNAs (miR-92a-3p, miR-92b-3p, miR-320c and miR-629-5p) were identified from OSCC patients via validation in the Chungnam National University Hospital cohort. Application of the specific miRNA panel for distinguishing OSCC patients from healthy controls produced specificity and sensitivity of 97.8 and 74%, respectively. In addition, the serum levels of these 4 miRNAs significantly decreased after complete surgical resection and increased after recurrence. We suggest that circulating 4-miRNA panel might be promising non-invasive predictors for diagnosing and monitoring the progression of patients with OSCC.
Similar content being viewed by others
Introduction
Oral squamous cell carcinoma (OSCC) is the most common type of oral cancer, accounting for approximately 350,000–400,000 cases per year. OSCC is twice as common in male than female due to risk factors, such as tobacco, alcohol and HPV. According to statistics, OSCC is the 6th and 8th particularly for incidence and mortality in both men, respectively1. Due to the high occurrence of secondary carcinoma and tumor heterogeneity, OSCC is often diagnosed in an advanced state with a poor prognosis2. Even though most cases of OSCC could be managed with complete surgical resection alone or a combination of ionizing radiation or chemo-radiation therapy, a certain proportion of advanced OSCCs remain unresponsive to treatment or exhibit loco-regional recurrence, resulting in a mortality rate of 50%3,4.
In the diagnosis and prevention of OSCC, emphasis is placed on identifying potential malignant lesions of the oral mucosa and local diseases that promote chronic inflammation, mainly relying on objective clinical examinations and surgical biopsy5,6. Although surgical biopsy is the gold standard for the diagnosis of OSCC, it is somewhat invasive and can sometimes be harmful to patients7. Moreover, conventional biopsy is temporally and spatially limited and often provides a brief snapshot of a single region of a heterogeneous tumor8. Therefore, it is crucial to find promising non-invasive biomarkers for monitoring or patient surveillance and further illuminate the pathogenesis of OSCC regarding tumor behavior at the molecular level. Blood samples are relatively easy to collect in a minimally invasive manner, and increasingly many recent studies have suggested that circulating microRNAs (miRNAs) are promising as potential biomarkers for disease diagnosis and monitoring9,10.
miRNAs are small, non-coding RNAs of 18–25 nucleotides in length that have been linked to essentially all known pathological and physiological processes, including cancer. Recent studies have reported that miRNAs can not only be utilized for diagnosis and prognosis, but also play integral and convoluted roles in the regulatory network of cancer. miRNA have been reported as diagnostic biomarkers for many cancers, including head and neck cancer11,12. However, the approach of using tissue-derived miRNA in surveillance or prognosis is commonly invasive in nature which may impede the screening. Furthermore, previous studies have demonstrated that miRNA can be quite stable in serum due to its protection from endogenous RNase activity and that it is readily detected by various assays13,14, presents the possibility to exploiting circulating miRNAs as biomarkers for early-stage cancer. Therefore, serum miRNA panel signatures have recently been identified as promising candidate biomarkers for liquid biopsy. However, studies have rarely examined circulating miRNA expression in patients with OSCC, leading to little noticeable and reliable signatures.
The aim of this study was to explore and validate the possibility that circulating miRNAs could overcome the limitations of tissue biopsy and act as potential biomarkers in liquid biopsy for the early diagnosis and dynamic monitoring of disease progression in OSCC patients.
Materials and methods
Patient and sample collection
Serum samples from 27 patients with OSCC and 21 age- and sex-matched healthy individuals were obtained at the Chungnam National University Hospital (CNUH) (Daejeon, Republic of Korea), between January 2017 and December 2019. The clinical information of patients with OSCC were summarized in Table S1. Tumor tissues and adjacent non-tumor tissues were collected from 7 patients with OSCC. All patients with OSCC were enrolled at the initial diagnosis, and the pathological diagnoses were subsequently confirmed. The study participant provided an informed consent form before participating. The Institutional Review Board of CNUH approved this study (CNUH 2019-07-041). All methods were performed in accordance with the Institutional Review Board of CNUH guideline and regulation.
Next-generation sequencing and analysis
Serum samples from 4 OSCC patients and 6 age- and sex-matched healthy controls were selected from CNUH cohort for next-generation sequencing10. The clinical information of patients with OSCC for NGS were presented in Table S2. Whole-transcriptome next-generation sequencing was performed by Macrogen Inc. (Seoul, Republic of Korea). Briefly, extracted RNA samples were used to prepare small RNA libraries using SMARTer smRNA-Seq Kit protocol and sequenced using a HiSeq 2500 sequencer (Illumina, San Diego, CA, USA), following the HiSeq 2500 System User Guide Document #15035786 v02 HCS 2.2.70 protocol. After sequencing, the raw sequence reads were filtered based on quality determined by the phred quality score at each cycle (Table S3). Both the trimmed reads and non-adapter reads as processed reads were used, to do analyzing long target (≧ 50 bp).The processed reads were gathered forming a unique cluster. In order to eliminate the effect of large amounts of ribosomal RNA (rRNA) from this study, the read was aligned to the rRNA sequence. rRNA removed reads were sequentially aligned to reference genome (UCSC Homo sapiens reference genome (GRCh37/hg19)), miRBase v21 and non-coding RNA database, RNAcentral 10.0 to classify known miRNAs and other type of RNA such as tRNA, snRNA, snoRNA etc. Novel miRNA prediction was performed by miRDeep2. The read counts for each miRNA were extracted from mapped miRNAs, differentially expressed miRNAs (DEmiRNAs) were determined through comparing across conditions each miRNA using statistical methods. Detailed work flow of sequencing and analysis were additionally described in the supplementary material. Figure S1 represents the small RNA composition of each sample.
Bioinformatics
Differentially expressed miRNAs (DEmiRNAs) between the evaluated groups were estimated using DESeq2 and edgeR15. The screening criteria were a fold change > 3 and p < 0.05. All genomic data of OSCC from The Cancer Genome Atlas (TCGA) were obtained from a specific portal (https://tcga-data.nci.nih.gov) and cancer browser (https://genome-cancer.ucsc.edu). To select miRNA differentially expressed between patients with OSCC and normal controls, false discovery rate-adjusted p values (< 0.05) were used to correct, using the Benjamin-Hochberg method. A volcano map, heatmap, and cluster analysis were conducted using an online analysis tool (https://www.chiplot.online/), a free online platform for data analysis and visualization. The target genes of miRNAs were predicted with the TargetScan 8.0 database (www.targetscan.org). Functional annotation was performed using the Database for Annotation, Visualization and Integrated Discovery (https://david.ncifcrf.gov/), a web-accessible tool for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. A network analysis of miRNA-mRNA interactions was carried out using Cytoscape (version 3.7.1), an open bioinformatics software.
RNA extraction and quantitative real time polymerase chain reaction (qRT-PCR)
Circulating miRNA was isolated from 200 μL of serum for RNA purification using miRNeasy serum/plasma kits (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Total RNA was extracted from tissue samples using the TRIzol reagent (Invitrogen, Waltham, MA, USA). The SYBR Green qRT-PCR assay was used for miRNA quantification. Total miRNA was used as the template for cDNA synthesis with miScript II RT Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. For miRNA analysis, qRT-PCR was performed using the miScript SYBR Green PCR kit (Qiagen, Hilden, Germany) with the manufacturer-provided miScript assays, using the universal primer and miRNA-specific forward primers with the 7500 system. The miRNA-specific primers were obtained from the miScript primer assays, the miRCURY LNA miRNA PCR Assay (Qiagen, Hilden, Germany), and Bioneer (Daejeon, Korea). All primer sequences used for qRT-PCR are listed in Table S4. miR-16 and miR-423-5p were used as references for serum miRNA analysis, and U6 small nuclear (RNU6) was used as the reference for the tissue expression of miRNA. At the end of the PCR cycles, melting curve analyses were performed. Each sample was run in triplicate for analysis. The 2 − ΔΔCT method was used to analyze the expression levels of miRNA.
Statistical analysis
All statistical analyses were performed using SPSS for Windows version 26 (IBM Corp., Armonk, NY, USA) and GraphPad Prism 8 (GraphPad Software, La Jolla, CA, USA). All experiments in the CNUH OSCC patient cohort were repeated three times. The data on the expression differences of miRNA between patients with OSCC and healthy controls, and between the serum samples and tissue samples from the same patients were analyzed using Mann–Whitney U test or independent t-test. The data on the expression differences of miRNA between the same patients before and after surgery were analyzed by paired t-test. Data are expressed as means ± SD. *p < 0.05, **p < 0.01, ***p < 0.001. Receiver operating characteristic (ROC) curves were used to analyze the diagnostic value of DEmiRNAs. A logistic regression model was constructed to determine the predicted probability of the combination of the 4 miRNAs. Pearson correlation coefficients were used to compare the miRNA levels in serum and tissue. The independent t-test was used to identify possible associations between miRNA concentrations and clinicopathological features of OSCC patients. The levels of miRNA in each group were presented as mean ± standard deviation (SD). All p values were two-sided, and a p value < 0.05 was considered statistically significant.
Results
Serum miRNA profiling to identify differential expression between healthy individuals and OSCC patients
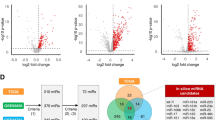
To identify potential circulating miRNA biomarkers of OSCC, we measured serum expression levels in 4 patients with OSCC and 6 healthy controls by NGS (small RNA-sequencing). The overview of the research workflow is illustrated in Fig. S2. In the initial screening, we identified 272 DEmiRNAs between patients with OSCC and healthy controls based on the exactTest using DESeq2 and edgeR. We screened out 42 DEmiRNAs, including 26 up- and 16 down-regulated DEmiRNAs, based on 2 criteria: (1) compared to the healthy group, the DEmiRNAs in the OSCC group had at least a threefold change in expression; and (2) the p values had statistical significance (p ≤ 0.05) with adjustment by the Benjamin-Hochberg procedure for multiple testing correction (Fig. 1A). The volcano plot directly presents the miRNA expression levels, and the most significantly up-regulated miRNA was shown to be miR-92a-3p, which had a log2 fold change of 2.46 (Fig. 1B). Using each sample’s normalized value, principal component analysis showed a circulating miRNA expression signature that segregated the serum samples of OSCC from those of healthy controls (Fig. 1C). We also identified the similarity between samples of the same group through Pearson correlation coefficients for the normalized values (Fig. 1D). Studies of microRNA in serum specific to cancer patients is based on the premise that it is expressed through the process of being released into the bloodstream from cancer tissue. The mechanism is not yet clear, but it is known to be a product of tumor cell death and dissolution or release from tumor-derived microsomes or exosomes16,17. Therefore, to discover specific candidate miRNAs, we combined our small RNA-sequencing results and data from TCGA, a large-scale tissue-derived database. Finally, 9 miRNAs were identified as candidates due to their differential expression in both the serum and tissue of OSCC patients. A Venn diagram (Fig. 1E) shows the screening pattern, and Fig. 1F and Table S5 present the 9 candidates, including 5 up- and 4 down-regulated DEmiRNAs. To further investigate the functions and pathways by which the dysregulation of the DEmiRNAs influences OSCC development, we predicted the target genes of the 9 DEmiRNAs and performed GO and KEGG pathway enrichment analysis. The target genes of the 9 DEmiRNAs participated in cancer progression-related processes, such as the PI3K-Akt signaling pathway and signaling pathways regulating choline metabolism (Fig. S3A-B). For biological processes, cellular components, and molecular function, the target genes of DEmiRNAs were significantly concentrated in cellular nitrogen compound metabolic process, organelle, ion binding, and biosynthetic process (Fig. S3C-D). Next, we identified downstream targets associated with DEmiRNA that could play a regulatory role in OSCC progression. The miRNA target predictions were performed using the TargetScan databases, and then we used Cytoscape software to visualize and analyze the predicted data for interactions in miRNA-mRNA regulatory networks (Fig. S3E-F). It was confirmed that one miRNA regulated the signal transduction pathway in association with several mRNAs and was also interconnected with other miRNAs.
miRNA profiling identifies differential expression in OSCC. (A) Heatmap of miRNAs that were differentially expressed between oral squamous cell carcinoma (OSCC) patients and normal controls (p < 0.05). Four pre-treatment OSCC serum samples and 6 normal serum samples are shown in the heatmap. (B) A volcano plot shows that many miRNAs were significantly different between normal and OSCC patients. Yellow: up-regulation with a fold change of more than 3; blue: down-regulation with a fold change of more than −3 (p > 0.05). (C) Principal component analysis (PCA). The fold change in expression between matched normal and tumor samples was used to perform PCA. (D) Heatmap of correlations based on the OSCC and normal serum samples. The correlogram shows correlation coefficients for all pairs of variables with coefficients colored based on their sign. (E) Venn diagram of differentially expressed miRNAs (DEmiRNAs) obtained from The Cancer Genome Atlas (TCGA) database and RNA sequencing results (fold change ≥ 3 or ≤ −3, p < 0.05). The Venn diagram shows that there are 9 overlap** DEmiRNAs. (F) Heatmap of one-way hierarchical clustering revealed 9 DEmiRNAs. The heatmap, volcano plot and PCA plot were conducted using an online tool ChiPlot (https://www.chiplot.online/).
Validation of the candidate miRNAs in the CNUH OSCC patient cohort by qRT-PCR
To search for and validate potential miRNA signatures to distinguish OSCC patients from healthy controls, we planned to validate the 9 candidate DEmiRNAs in the CNUH OSCC patient cohort. We collected serum samples from 23 OSCC patients with different subsites (tongue, n = 18; buccal mucosa, n = 2; retromolar trigone, n = 2; and floor of mouth mucosa, n = 1) and 15 healthy controls from the cohort for validation by qRT-PCR. The clinical information of patients with OSCC were summarized in Table S1.
Previous studies have shown that RNU6 can generally be used as an endogenous control (EC) to normalize the expression of miRNA in tissue or cells, but it is unstably expressed in the plasma and serum24. Beyond OSCC, it has been reported that miR-92a-3p also plays a role as an oncogenic component in gastric cancer25, esophageal squamous cell cancer The TCGA data presented in this study are openly available in a specific portal (https://tcga-data.nci.nih.gov) and cancer browser (https://genome-cancer.ucsc.edu). Further information is available from the corresponding author upon request. Area under the curve Biological processes Cellular components Confidence interval Chungnam National University Hospital Differentially expressed miRNAs Endogenous control Gene ontology Head and neck cancer Kyoto encyclopedia of genes and genomes Molecular function MicroRNA Next-generation sequencing Oral squamous cell carcinoma Principal component analysis Receiver operating characteristics Real-time polymerase chain reaction The cancer genome atlas Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J. Clin. 68(6), 394–424. https://doi.org/10.3322/caac.21492 (2018). Uz, U. & Eskiizmir, G. Association between interleukin-6 and head and neck squamous cell carcinoma: A systematic review. Clin. Exp. Otorhinolaryngol. 14(1), 50–60. https://doi.org/10.21053/ceo.2019.00906 (2021). Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249. https://doi.org/10.3322/caac.21660 (2021). Park, J. O. et al. Survival benefits from surgery for stage IVa head and neck squamous cell carcinoma: A multi-institutional analysis of 1,033 cases. Clin. Exp. Otorhinolaryngol. 14(2), 225–234. https://doi.org/10.21053/ceo.2020.01732 (2021). Ford, P. J. & Farah, C. S. Early detection and diagnosis of oral cancer: Strategies for improvement. J. Cancer Policy 1(1–2), e2–e7. https://doi.org/10.1016/j.jcpo.2013.04.002 (2013). Ahn, S. H. Usage and diagnostic yield of fine-needle aspiration cytology and core needle biopsy in thyroid nodules: A systematic review and meta-analysis of literature published by Korean authors. Clin. Exp. Otorhinolaryngol. 14(1), 116–130. https://doi.org/10.21053/ceo.2020.00199 (2021). Park, W. et al. Sentinel lymph node biopsy versus elective neck dissection: Long-term oncologic outcomes in clinically node-negative tongue cancer. Clin. Exp. Otorhinolaryngol. 15(1), 107–114. https://doi.org/10.21053/ceo.2020.02411 (2022). Bellairs, J. A., Hasina, R. & Agrawal, N. Tumor DNA: An emerging biomarker in head and neck cancer. Cancer Metastasis Rev. 36(3), 515–523. https://doi.org/10.1007/s10555-017-9685-x (2017). Nagasaka, M. et al. Liquid biopsy for therapy monitoring in early-stage non-small cell lung cancer. Mol. Cancer. 20(1), 82. https://doi.org/10.1186/s12943-021-01371-1 (2021). Umu, S. U. et al. A comprehensive profile of circulating RNAs in human serum. RNA Biol. 15(2), 242–250. https://doi.org/10.1080/15476286.2017.1403003 (2018). Sethi, N., Wright, A., Wood, H. & Rabbitts, P. MicroRNAs and head and neck cancer: Reviewing the first decade of research. Eur. J. Cancer (Oxford, England : 1990) 50(15), 2619–2635. https://doi.org/10.1016/j.ejca.2014.07.012 (2014). Kabzinski, J., Maczynska, M. & Majsterek, I. MicroRNA as a novel biomarker in the diagnosis of head and neck cancer. Biomolecules https://doi.org/10.3390/biom11060844 (2021). Patrick, S., Mitchell, R. K. P. & Kroh, E. M. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 105(30), 10513–10518. https://doi.org/10.1073/pnas.0804549105 (2008). Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin. Chem. 56(11), 1733–1741. https://doi.org/10.1373/clinchem.2010.147405 (2010). Seyednasrollah, F., Laiho, A. & Elo, L. L. Comparison of software packages for detecting differential expression in RNA-seq studies. Brief Bioinform. 16(1), 59–70. https://doi.org/10.1093/bib/bbt086 (2015). Chen, X. et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 18(10), 997–1006. https://doi.org/10.1038/cr.2008.282 (2008). Yu, S. et al. Circulating microRNA profiles as potential biomarkers for diagnosis of papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. 97(6), 2084–2092. https://doi.org/10.1210/jc.2011-3059 (2012). **ang, M. et al. U6 is not a suitable endogenous control for the quantification of circulating microRNAs. Biochem. Biophys. Res. Commun. 454(1), 210–214. https://doi.org/10.1016/j.bbrc.2014.10.064 (2014). Li, A. et al. MicroRNA array analysis finds elevated serum miR-1290 accurately distinguishes patients with low-stage pancreatic cancer from healthy and disease controls. Clin. Cancer Res. 19(13), 3600–3610. https://doi.org/10.1158/1078-0432.Ccr-12-3092 (2013). Ragni, E. et al. Identification of miRNA reference genes in extracellular vesicles from adipose derived mesenchymal stem cells for studying osteoarthritis. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20051108 (2019). Kopanska, M. et al. MiRNA expression in the cartilage of patients with osteoarthritis. J. Orthop. Surg. Res. 12(1), 51. https://doi.org/10.1186/s13018-017-0542-y (2017). Kang, K., Peng, X. & Luo, J. Identification of circulating miRNA biomarkers based on global quantitative real-time PCR profiling. J. Anim. Sci. Biotechnol. https://doi.org/10.1186/2049-1891-3-4 (2012). Hsu, M., Chang, Y. C. I. & Hsueh, H. M. Biomarker selection for medical diagnosis using the partial area under the ROC curve. BMC Res. Notes https://doi.org/10.1186/1756-0500-7-25 (2014). Salazar-Ruales, C. et al. Salivary microRNAs for early detection of head and neck squamous cell carcinoma: A case-control study in the high altitude mestizo ecuadorian population. Biomed. Res. Int. 2018, 9792730. https://doi.org/10.1155/2018/9792730 (2018). Zhang, G. et al. LncRNA MT1JP functions as a ceRNA in regulating FBXW7 through competitively binding to miR-92a-3p in gastric cancer. Mol. Cancer. 17(1), 87. https://doi.org/10.1186/s12943-018-0829-6 (2018). **n Li, S. G., Min, L., Guo, Q. & Zhang, S. miR-92a-3p promotes the proliferation, migration and invasion of esophageal squamous cell cancer by regulating PTEN. Int. J. Mol. Med. 44(3), 973–981. https://doi.org/10.3892/ijmm.2019.4258 (2019). Liu, S. et al. miR-92a-3p promoted EMT via targeting LATS1 in cervical cancer stem cells. Front. Cell Dev. Biol. 9, 757747. https://doi.org/10.3389/fcell.2021.757747 (2021). **ghua, H. et al. MicroRNA miR-92a-3p regulates breast cancer cell proliferation and metastasis via regulating B-cell translocation gene 2 (BTG2). Bioengineered 12(1), 2033–2044. https://doi.org/10.1080/21655979.2021.1924543 (2021). Hu, J. L. et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer. 18(1), 91. https://doi.org/10.1186/s12943-019-1019-x (2019). Long, M. et al. miR-92b-3p acts as a tumor suppressor by targeting Gabra3 in pancreatic cancer. Mol. Cancer. 16(1), 167. https://doi.org/10.1186/s12943-017-0723-7 (2017). Wu, Z. B. et al. The miR-92b functions as a potential oncogene by targeting on Smad3 in glioblastomas. Brain Res. 1529, 16–25. https://doi.org/10.1016/j.brainres.2013.07.031 (2013). Gong, L., Ren, M., Lv, Z., Yang, Y. & Wang, Z. miR-92b-3p promotes colorectal carcinoma cell proliferation, invasion, and migration by inhibiting FBXW7 in vitro and in vivo. DNA Cell Biol. 37(5), 501–511. https://doi.org/10.1089/dna.2017.4080 (2018). Wang, C. et al. MicroRNA-92b-3p is a prognostic oncomiR that targets TSC1 in clear cell renal cell carcinoma. Cancer Sci. 111(4), 1146–1155. https://doi.org/10.1111/cas.14325 (2020). Li, C., Huo, B., Wang, Y. & Cheng, C. Downregulation of microRNA-92b-3p suppresses proliferation, migration, and invasion of gastric cancer SGC-7901 cells by targeting Homeobox D10. J. Cell Biochem. 120(10), 17405–17412. https://doi.org/10.1002/jcb.29005 (2019). Li, M. et al. Exosomal miR-92b-3p promotes chemoresistance of small cell lung cancer through the PTEN/AKT pathway. Front. Cell. Dev. Biol. 9, 661602. https://doi.org/10.3389/fcell.2021.661602 (2021). Uotani, K. et al. Circulating .icroRNA-92b-3p as a novel biomarker for monitoring of synovial sarcoma. Sci. Rep. 7(1), 14634. https://doi.org/10.1038/s41598-017-12660-5 (2017). Li Yh, Y., Zhou, C., Jiang, P.-C. & Pan, W. MiR-320c prevents the malignant development of cervical cancer by regulating GABRP level. Eur. Rev. Med. Pharmacol. Sci. 24(17), 8731–8739. https://doi.org/10.26355/eurrev_202009_22810 (2022). Wang, H. Y. et al. Profiling plasma microRNA in nasopharyngeal carcinoma with deep sequencing. Clin. Chem. 60(5), 773–782. https://doi.org/10.1373/clinchem.2013.214213 (2014). Wang, J. et al. Circulating exosomal miR-125a-3p as a novel biomarker for early-stage colon cancer. Sci. Rep. 7(1), 4150. https://doi.org/10.1038/s41598-017-04386-1 (2017). Shen, Y. et al. Identification of novel circulating miRNA biomarkers for the diagnosis of esophageal squamous cell carcinoma and squamous dysplasia. Cancer Epidemiol. Biomark. Prev. 28(7), 1212–1220. https://doi.org/10.1158/1055-9965.EPI-18-1199 (2019). Liu, Y. et al. MiR-629–5p promotes prostate cancer development and metastasis by targeting AKAP13. Front Oncol. 11, 754353. https://doi.org/10.3389/fonc.2021.754353 (2021). Tao, X. et al. miR-629-5p promotes growth and metastasis of hepatocellular carcinoma by activating beta-catenin. Exp. Cell Res. 380(2), 124–130. https://doi.org/10.1016/j.yexcr.2019.03.042 (2019). Li, Y. et al. MiR-629-5p promotes the invasion of lung adenocarcinoma via increasing both tumor cell invasion and endothelial cell permeability. Oncogene 39(17), 3473–3488. https://doi.org/10.1038/s41388-020-1228-1 (2020). Cheng, G. Circulating miRNAs: Roles in cancer diagnosis, prognosis and therapy. Adv. Drug Deliv. Rev. 81, 75–93. https://doi.org/10.1016/j.addr.2014.09.001 (2015). Xu, L., Cai, Y., Chen, X., Zhu, Y. & Cai, J. Circulating MiR-1290 as a potential diagnostic and disease monitoring biomarker of human gastrointestinal tumors. BMC Cancer 21(1), 989. https://doi.org/10.1186/s12885-021-08729-0 (2021). Gandellini, P., Giovannetti, E. & Nicassio, F. MicroRNAs in cancer management: big challenges for small molecules. Biomed. Res. Int. https://doi.org/10.1155/2015/982156 (2015). Salloum-Asfar, S., Satheesh, N. J. & Abdulla, S. A. Circulating miRNAs, small but promising biomarkers for autism spectrum disorder. Front. Mol. Neurosci. 12, 253. https://doi.org/10.3389/fnmol.2019.00253 (2019). Tiberio, P., Callari, M., Angeloni, V., Daidone, M. G. & Appierto, V. Challenges in using circulating miRNAs as cancer biomarkers. Biomed. Res. Int. https://doi.org/10.1155/2015/731479 (2015). Wang, K. et al. Comparing the microRNA spectrum between serum and plasma. PLoS ONE 7(7), e41561. https://doi.org/10.1371/journal.pone.0041561 (2012). This research was supported by Chungnam National University Hospital Research Fund 2019 [to BS Koo], Chungnam National University Sejong Hospital Research Fund 2021 [to HR Won], and the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning [grant numbers 2019R1A2C1084125 to BS Koo, 2022R1C1C1008265 to HR Won and 2021R1C1C1014142 to JW Chang], and by the Korea Health Technology R&D Project through the Korea health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HR20C0025 and HR22C1734), and by Korea Medical Device Development Fund (Ministry of Science and ICT, Ministry of Trade, Industry and Energy, Ministry of Health & Welfare, Ministry of Food and Drug Safety; Project Number: 1711138229, KMDF_PR_20200901_0124). The work reported in the paper has been performed by the authors, unless clearly specified in the text. Y.P., contributed to conception, design, data acquisition, and interpretation, drafted and critically revised the manuscript; S.-N.J., M.A.L., J.W.C. contributed to design and data acquisition, C.O., Y.L.J., H.J.K., N.Q.K. contributed to data analysis; H.-R.W., B.S.K. contributed to conception, design and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work. The authors declare no competing interests. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. Piao, Y., Jung, SN., Lim, M.A. et al. A circulating microRNA panel as a novel dynamic monitor for oral squamous cell carcinoma.
Sci Rep 13, 2000 (2023). https://doi.org/10.1038/s41598-023-28550-y Received: Accepted: Published: DOI: https://doi.org/10.1038/s41598-023-28550-yData availability
Abbreviations
References
Funding
Author information
Authors and Affiliations
Contributions
Corresponding authors
Ethics declarations
Competing interests
Additional information
Publisher's note
Supplementary Information
Rights and permissions
About this article
Cite this article