Abstract
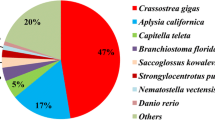
The shells of window pane oyster Placuna placenta are very thin and exhibit excellent optical transparency and mechanical robustness. However, little is known about the biomineralization-related proteins of the shells of P. placenta. In this work, we report the comprehensive transcriptome of the mantle tissue of P. placenta for the first time. The unigenes of the mantle tissue of P. placenta were annotated by using the public databases such as nr, GO, KOG, KEGG, and Pfam. 24,343 unigenes were annotated according to Pfam database, accounting for 21.48% of the total unigenes. We find that half of the annotated unigenes of the mantle tissue of P. placenta are consistent to the annotated unigenes from pacific oyster Crassostrea gigas according to nr database. The unigene sequence analysis from the mantle tissue of P. placenta indicates that 465,392 potential single nucleotide polymorphisms (SNPs) and 62,103 potential indel markers were identified from 60,371 unigenes. 178 unigenes of the mantle tissue of P. placenta are found to be homologous to those reported proteins related to the biomineralization process of molluscan shells, while 18 of them are highly expressed unigenes in the mantle tissue. It is proposed that four unigenes with the highest expression levels in the mantle tissue are very often related to the biomineralization process, while another three unigenes are potentially related to the biomineralization process according to the Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR) analysis. In summary, the transcriptome analysis of the mantle tissue of P. Placenta shows the potential biomineralization-related proteins and this work may shed light for the shell formation mechanism of bivalves.
Similar content being viewed by others
Introduction
The biological exoskeletons are usually composed of highly ordered hierarchical micro- and nanostructures occluded with organic molecules and exhibit superior mechanical, optical, thermal, magnetic properties 1,2,3. Calcium carbonate is one of the most abundant biominerals in nature, and the biomineralization and bioinspired mineralization of calcium carbonate have been the focus of investigation for many decades. The molluscan shells have well defined micro- and nanostructures and excellent mechanical properties, composed of 95% CaCO3 and less than 5 wt % of organic macromolecules (proteins, glycoproteins, polysaccharides and lipids) 4,5. Generally, the polymorphs of CaCO3 include aragonite, calcite, vaterite, calcite monohydratre and calcium carbonate hemihydrate 6. Biogenic CaCO3 exhibits complex microstructures such as prismatic, nacreous, foliate, and cross-lamellar microstructures 7,8. Importantly, protein components in the molluscan shells are responsible for the nucleation and crystal growth of biogenic CaCO3 and can enhance the mechanical properties of the shells 9. The matrix proteins occluded in the molluscan shells were extracted, sequenced and identified as biomineralization-related proteins which were found to have functions including stabilization of amorphous calcium carbonate, inhibition or acceleration of nucleation of calcium carbonate 10,11,12,13. A few examples of biomineralization-related proteins include mantle protein N25 14 , perlucin, perlustrin 15, aspein 16, prisilkin-39 17, Shematrin-2 18, PfN44 19, and enzymes such as carbonic anhydrase 20. Most of matrix proteins in molluscan shells are specifically secreted by the mantle tissue of molluscans. In another word, the proteins from the mantle tissues of molluscans can tune the formation of different structural layers of the shells 12,S5). The numbers of transition (261,292) and transversion (202,192) mutations accounted for 56.14% and 43.45% in SNP types, respectively. The most abundant type of base variations is C/T polymorphism (70,283; 15.1%), because C in CG base is often methylated and transfers to thymine after spontaneous deamination. Another abundant type of base variations is A/T polymorphism, such as G to A (70,176; 15.08%), A to G (61,051; 13.12%) (Fig. 7). 62,103 potential indels in total were also identified among 21,630 unigenes with frequency of about 2.87 indels per unigene (Excel S6). The numbers of unigenes decrease gradually with the increase of indel length from 1 to 10 bp (Fig. S1). The indel lengths are mainly 1, 2 and 3 bp. The numbers of deletion are larger than that of insertion when the indel length is more than 10 bp (Fig. S1).
In addition, 21,640 unigenes containing 26,048 potential SSR markers were identified (Fig. 8). Among these unigenes, the most abundant type of repeats is p1 (mono-nucleotide) (15,224; 58.4%), followed by p2 (di-nucleotide) (6227; 23.9%), p3 (tri-nucleotide) (2700; 10.3%), p4 (tetra-nucleotide) (361; 1.3%) repeats. Furthermore, there are some c- types (complex repeat motifs), accounting for 5.8% (1524), different from the above mentioned nucleotides in these SSR markers. The frequencies of SSRs with different numbers of repeat units were calculated. Among the p1 repeat units, the counts of the repeat units between 9 and 12 are dominant, accounting for more than 50%, followed by 13–16 repeat units (17.74%) and 17–20 repeat units (12.85%). More than half of 5–8 repeat units were assigned to the p2 repeat units, followed by 9–12 repeat units. However, only 5–8 repeat units are present in both p3 and p4 repeats (Fig. 8). The A/T repeat units of mono-nucleotide are dominant among these types, accounting for 55.57%. AT/TA (9.42%) was dominant in the p2 repeat units, followed by TG/CA (4.94%), GA/GT (4.16%) (Fig. S2). ATG/GAT (2.2%) was dominant in the p3 repeat units, followed by ATC/TGA (1.71%), CAT/TCA (1.2%). The remaining types of motifs have more complicated types but less numbers (c-type, complex repeat motifs), accounted for 7.28% in total. To further test the SSR markers, forward and reverse primer pairs were obtained using Primer 3.0 (Excel S7).
Identification of genes involved in biomineralization process
To better understand the proteins related to the shell formation process, the annotated unigenes of mantle tissues of P. placenta were compared with the sequences of proteins known to be associated with biomineralization process in molluscan shells using nr database. The annotated unigene with lowest E-value was selected as the representative unigene while several annotated unigenes were assigned to the same reference unigene. 178 homologous unigenes of 51 shell matrix proteins, such as calmodulin, perlucin, ferritin and carbonic anhydrase were found to be related to biomineralization process in the transcriptome of mantle tissue of P. placenta (Excel S10). This is the first time to report the potential biomineralization-related unigenes in the mantle tissue of P. placenta, as far as we know. A lot of researchers have focused on the identification of genes related to the shell formation of molluscan animals, and an increasing number of genes have been identified 55,56,57,58. 259 proteins were identified from oyster C. gigas shells by proteomic analysis 56. In comparison with the proteomic data of shell matrix of C. gigas, we identified a set of 158 unigenes that are probably related to shell formation, including house-kee** protein elongation factor 1a, and extracellular matrix protein collagen (Excel S9). Many of the shell-formation related proteins are enzymes such as glutathione peroxidase, hemicentin and tyrosinase, that may be involved in matrix construction or modification 57,58. In this study, only one enzyme tyrosinase (three unigenes) was found to be related to the shell formation process in the transcriptome of P. placenta (Excel S9). Twenty-one proteins with 66 homologous unigenes were identified to be related to calcitic shell formation of P. placenta (Table 4). Furthermore, eighteen of the above 66 unigenes were identified to be highly expressed in mantle tissue of P. placenta. A few highly expressed unigenes with fragments per kilobase of feature per million mapped reads (FPKM) values of > 15 are carbonic anhydrase, calreticulin, ferritin, perlucin, gigasin-2, and tyrosinase-like proteins.
Comparison of the mantle transcriptomes of different molluscans
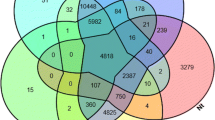
The shells of scallops P. yessoensis, C. farreri and P. placenta are composed of foliated calcite minerals. The mantle transcriptomes of scallop P. yessoensis, C. farreri and P. placenta were compared to find the similarities and differences of the biomineralization related proteins of these molluscan organisms 36,37. We find 117 biomineralization-related unigenes in the mantle of P. placenta, much less than those in P. yessoensis (162 unigenes), but much more than those in C. farreri (42 unigenes) (Fig. 9). There are six biomineralization-related unigenes expressed in the mantles of the three species, including sarcoplasmic calcium-binding protein, calcineurin a, calmodulin-like protein, perlucin, alkaline phosphatase, tyrosinase-like protein tyr-3. In comparison to C. farreri, P. placenta and P. yessoensis have more similar homologous biomineralization-related unigenes, 31 unigenes in total, about 27% of the biomineralization related unigenes of P. placenta. For example, collagen, chitin synthase 1/2, carbonic anhydrase-like, heat shock protein 70 and calmodulin were found in both P. yessoensis and P. placenta, potentially indicating their functional similarities for their biomineralization processes. There are probably more mineralization proteins to be discovered in the unigenes of the mantle tissue of P. placenta (Fig. 9, Excel S10).
Comparison of mantle transcriptomes of three scallop shells composed of folicated calcite crystals. Blue circle: 168 biomineralization-related unigenes (124 exclusive) discovered from P. yessoensis mantle 36. Yellow circle: 42 biomineralization-related unigenes (25 exclusive) discovered from C. farreri mantle 37. Green circle: 117 biomineralization-related unigenes (84 exclusive) discovered from P. placenta mantle.
Quantitative Real-Time PCR (qRT-PCR) analysis
Ten selected potential biomineralization-related unigenes in the different tissues such as adduction muscle, gonad, gill, mantle, hepatopancreas, mouthparts, and intestine of P. placenta were examined by qRT-PCR (Fig. 10). In general, four of ten unigenes have much higher expression in mantle tissue than those in the other tissues, which include c76266_g1 (gigasin-2), c73086_g1 (tyrosinase-like), c66761_g1 (pif-like) and c59513_g1 (teneurin-2) unigenes (Fig. 10a–d). Pif is an important macromolecule for in vivo shell formation of nacre 44. In addition, it was found that pif can induce the formation of aragonite and vaterite crystals in the in vitro system 77. Tyrosinases are abundant in shells and their high expression in mantle of the pacific oyster C. gigas indicates that their functions are probably related to shell formation 56. Gigasin-2 and teneurin-2 were identified for the first time from the shell of C. gigas via shell matrix proteome characterization 78,79. According to the Pfam database analysis, teneurin-2 in P. placenta is predicted to be epidermal growth factor (EGF) domain and gigasin-2 has zona pellucida (ZP) domain (Excel S11). EGF domains are mostly found in the SMPs as tandem repeats and only present in the prismatic (calcitic) layers but not in the nacreous layer of Pinctada80. The EGF domain is a calcium-binding motif composed of 45 amino acids arranged in two small \(\upbeta\)-sheets with six conserved cysteine residues 81. Both EGF-like and ZP domains have been reported in the shells of Lottia gigantea 82 and C. gigas 78. ZP domains are present in a range of extracellular filament or matrix proteins from a wide variety of eukaryotic organisms, and are characterized by eight conserved cysteine residues, which are involved in protein polymerization processes 83. We propose that the above mentioned four unigenes are primarily biomineralization-related unigenes. However, the expression levels of the other six unigenes in the mantle tissue are not as high as their expression levels in the other tissues, indicating that they probably have other functions except for the biomineralization process (Fig. 10e–j). We would discuss more about the functions of these ten unigenes in the discussion part.
Differential expression of gigasin-2 (c76266_g1) (a), tyrosinase-like (c73086_g1) (b), pif-like(c66761_g1) (c), teneurin-2 (c59513_g1) (d), perlucin (c67461_g2) (e), calmodulin-like, (c81494_g1) (f), carbonic anhydrase-like (c84941_g5) (g), caltractin-like (c70548_g1) (h), calreticulin-like (c84621_g1) (i), and insoluble matrix shell protein 6 (c69385_g1) (j) in adduction muscle, gonad, gill, mantle, hepatopancreas, mouthparts, and intestine tissues of P. placenta, were determined using real-time PCR, The error bars represent the standard error of three biological replicates, statistical significance was considered at p < 0.05.
Pfam database analysis
Pfam database search is important for understanding the possible biomineralization-related functions of the shell matrix proteins of molluscan animals. In molluscan animals, shell matrix proteins are very often repetitive, highly conserved, low complex domains. The functions of these protein domains involved in biomineralization have been studied by many groups 13,84. For example, the nacre protein perlucin contains a C-type lectin domain and has a broad carbohydrate-binding feature, which was supposed to facilitate calcium-dependent glycoprotein-protein interactions within the skeletal matrix 23. In molluscan animals, the pif-like proteins contain von willebrand factor type A (VWA), chitin-binding and laminin G domains, which can bind chitin framework and accelerate CaCO3 precipitation inside the chitin membrane, and then regulate their vertical alignment 85,86. In bivalves, most EF-hand proteins from the mantle tissue of bivalves are Ca2+ sensors or signal modulators, which may induce conformational change by binding with Ca2+, such as calmodulin, troponin C and myosin light chains 87.
In this study, the sequences of potential biomineralization genes were obtained from the transcriptomics analysis of P. placenta mantle. Based on these amino acid sequences, the information of their domain was obtained by using Pfam. Finally, we speculate on the function of potential biomineralization-related proteins of P. placenta based on the characteristics of the domain found in biomineralization proteins of molluscan. Identification of the possible biomineralization-related functions of the unigenes expressed in the mantle tissue of P. placenta were carried out by keyword searching according to the domain which were reported to be related to biomineralization in the Pfam database. Many potential biomineralization-related unigenes are proposed to be involved in the shell formation from the transcriptomes of the mantle tissue of P. placenta according to the Pfam database analysis. (Excel S11).
Calmodulin, calponin and mucin proteins are supposed to be associated with molluscan shell formation. Among those proteins, calponin was highly expressed in the mantle of P. placenta, with FPKM values > 551, but the expression values of most of the other two proteins are relatively low. Only c81494_g1 unigene (calmodulin-like protein) had a high expression level (FPKM values in between 38.17 and 76.44) ( Excel S11), but it expressed a higher quantity in mouth parts than that in the other tissues (Fig. 10f), suggesting that calmodulin may play other roles in P. placenta. The insoluble matrix shell protein 6 shows a higher expression in the tissues of hepatopancreas and intestine than that of the other tissues in P. placenta (Fig. 10j).
Cadherin and collagen proteins contain enriched amount of von willebrand factor type A and epidermal growth factor domains, indicating that they were derived from the extracellular matrix 88. However, most of them show a low expression level in the mantle tissue of P. placenta. Only c83310_g1 (collagen alpha-1) shows a relatively high expression level (FPKM values in between 11.18 and 32.32) (Excel S11). Perlucin extracted from abalone nacre contains a functional C-type lectin domain which can increase the precipitation rate of calcium carbonate from a saturated solution, indicating that it may promote the nucleation and/or the growth of calcium carbonate crystals 15,23. Among the proteins containing perlucin domains, only the c67461_g2 unigene (perlucin) shows a high expression level (FPKM values: 65.55–100.36) in the mantle tissue of P. placenta (Excel S11). However, c67461_g2 unigene shows higher expression in tissues of hepatopancreas than that of the other tissues by qRT-PCR (Fig. 10e). Carbonic_anhydrase domain was reported to be involved in the formation of both nacreous and prismatic layers 20,24,89. The expression values of all the carbonic anhydrase unigenes are relatively high, with FPKM values \(>\) 27, such as c71108_g1, c85950_g1, c81423_g3 and c84941_g5 unigenes (Excel S11). At the mean time, the expression of c84941_g5 was higher in gill than that in the other tissues (Fig. 10g). On the other hand, some extracellular enzymes or inorganic ion-binding proteins such as chitinase, hemicentin and peroxidasin with relatively high expression levels in the mantle tissue of P. placenta are probably involved in the shell formation (Excel S11). Chitinase in shell matrix may reconstruct the chitinaceous scaffold and promote the interaction between chitin and chitin binding protein 82. In comparison to the other ion-binding proteins, the c71155_g1 unigene (chitinase 1) has a relatively high expression level (FPKM values in between 96.85 and 125.15) (Excel S11), but we didn't know yet how specific this gene is in each tissue of P. placenta.
Discussion
Shell formation is a very complicated process that involves a series of proteins and genes, while living organisms produce biominerals with superior mechanical properties under biological control 1. The main objective of this study is to identify unigenes involved in biomineralization. Our study indicates that a high-coverage expression profile can be produced by using short-read Illumina sequencing technology and some effective sequence assembly tools such as Trinity and ESTScan softwares. A total of 113,325 unigenes with an average length of 697 bp was generated from the m antle tissue of P. placenta by using Illumina HiSeqTM 4000 sequencing technology, while 66.39% of the above unigenes (76,237) have lengths less than 500 bp (Fig. 1). The lengths of the unigene sequences from the mantle tissue of P. placenta are larger than those generated in the reported unigene sequences of P. penguin 34. These unigene sequences are similar to the reports for the yesso scallop P. yessoensis (93,204 unigenes; 733 bp) 36 and the freshwater pearl mussel C. plicata (98,501 unigenes; 689 bp) 33. However, the number and mean length of the unigenes from the mantle tissue of P. placenta are larger than that from the zhikong scallop C. farreri (77,975 unigenes; 538 bp) 37 and the pearl oyster P. maxima (108,704 unigenes; 407 bp) 90. This difference may be attributed to the use of different sequencing platforms. A large quantity of genomic data is available for many bivalve species, but only 18.99% of the mantle tissue of P. placenta were annotated in nr database. This means that more than half of the unigenes of the mantle tissue of P. placenta have no known homologous unigenes. The low rate of annotated unigenes of the mantle tissue of P. placenta could be a result of limitations in the genomic information available for P. placenta, which is the case in many of other bivalve species 33,36.
The SNPs of the mantle tissue of P. placenta were obtained from the sequencing errors as well as the true SNPs. In our data, the obtained SNP density was 0.0059 SNPs/bp (0.59%) in the mantle tissue of P. placenta (Excel S5, Table 2), which was significantly larger than the sequencing error (0.01%) (Table 1). This result confirmed the reliability of the obtained SNPs data. The SNPs are potentially useful for genetic linkage map** and for the analysis of quantitative traits of the P. placenta. The transcriptome we present here provides the most comprehensive polymorphism for the P. placenta to date, as far as we know. The SNP density of the eastern oyster C. virginica is 0.042 SNPs/bp 54, which was more polymorphic than that of the mantle of P. placenta (0.0059 SNPs/bp) (Excel S5, Table 2). 465,392 potential SNPs were constructed from 60,371 unigenes, with frequency of about 7.56 SNPs per unigene in the mantle tissue of P. placenta, which was consistent with the reported result of the mantle tissue of pearl oyster P. martensii 35. The indel density of the mantle tissue of P. placenta was 2.87 indels per unigene (Excel S6), which was much lower than that indel density of the mantle tissue of P. martensii 35. As is well known, the Illumina sequencing is considered to be robust against homopolymer errors and therefore it may be suited well for identification of indels 91.
The variations in unigene expression between different tissues have been shown to be correlated with shell formation in molluscans such as C. gigas 22, P. penguin 34, and T. pyramis 32. The exact biomineralization functions of proteins such as perlucin 15 and pif 85 have been investigated by in vitro and in vivo mineralization studies. According to the nr database, 178 potential biomineralization-related unigenes were identified in the mantle transcriptome of P. placenta in this work. Among these unigenes, ten selected potential biomineralization-related unigenes (FPKM values \(>\) 15) were examined in the different tissues of P. placenta by qRT-PCR. In the present study, four of ten unigenes (gigasin-2, tyrosinase-like, pif-like and teneurin-2) have much higher expression in the mantle tissue than those in the other tissues, indicating that they are very often related with the biomineralization process of P. placenta shell (Fig. 10a–d). Three gigasin-2 isoforms were identified in water soluble matrix of C. gigas shell, which are proposed to be involved in bone remodeling processes and could be responsible for the biocompatibility between bone and nacre grafts 79. Meanwhile, gigasin-2 was highly expressed in the mantle tissue of C. gigas 56. However, homologous proteins have not been identified in other species since gigasin-2 was reported in 2012 56. It is known that tyrosinase family are potentially involved in melanin biosynthetic pathway in various organisms. Moreover, it was reported that tyrosinases from molluscans are secreted from the mantle and transported to the prismatic layer of the shell, while they contribute to melanin biosynthesis and shell pigmentation 56,92,93. In this study, there is no pigmentation in the mantle tissue and the transparent shell of P. placenta. Therefore, the high expression of tyrosinase in the mantle tissue indicates that their functions are not only related to melanin biosynthesis, but also related to the shell formation. After injection of pif dsRNA, both of calcite laths of the C. cigas shell and nacreous layer of the P. fucata shell grew to disordered structure in vivo, indicating that pif protein might be essential for the normal growth of the prismatic and nacreous layer 44,77. The teneurin-2 was first identified from diverse shell matrix proteome and had signal peptides in C. gigas, and it was proposed that it was secreted from the mantle into the shell 78. According to the above discussion about the expressions of the four unigenes and their functions for biomineralization process in the other molluscan species in the literatures, we propose that gigasin-2, tyrosinase-like, pif-like and teneurin-2 may play important roles for the biomineralization process. It would be important to extract these four proteins and investigate their functions for biomineralization via in-vitro crystallization process of CaCO3 in the future. We consider to study the full-length cDNA sequences, gene expression and recombinant proteins of these four unigenes to understand their functions for in vivo and in vitro crystallization of CaCO3.
Six of the remaining unigenes don’t exhibit high expression in the mantle tissue, in comparison to the other tissues of P. placenta. Thus, it is hard to tell whether they participate in biomineralization or not based on the qRT-PCR analysis. These findings are somewhat similar from those of previous studies. For example, the researchers identified six types of perlucin and discovered their different expression levels in different tissues of T. pyramis. Some of the perlucin proteins were expressed at the highest levels in the digestive gland, while the others were expressed at high levels in the mantle or the gonad of T. pyramis 32. The perlucin was isolated from the nacreous layer of the marine snail Haliotis laevigata and it could promote the nucleation of CaCO3 crystals on the calcite surface in the in vitro experiments 94. In addition, calmodulin-like protein can induce the nucleation of aragonite through binding with the 16-kDa protein and regulates the growth of calcite in the prismatic layer of pearl oyster P. fucata 95. This expression pattern and the in vitro crystallization experiments suggest that perlucin family may play important roles in both of the biomineralization process and digestive process 32,94. Similar to the perlucin, two types of calmodulin were expressed at the lowest level in the mantle than the other tissues in T. pyramis 32. Calmodulin-like protein was expressed with the highest level in the mantle tissue of P. fucata species and has a potentially high affinity for calcium 96. The carbonic anhydrase family were expressed in the mantle and associated with the shell formation in the european abalone Haliotis tuberculata 97. Nacrein containing carbonic anhydrase domain was expressed in both the nacreous layer and the prismatic layer of P. fucata 24. Meanwhile, it was also highly expressed in the mantle of T. pyramis and P. penguin 32,34. However, the researchers analyzed the expression levels of the five studied carbonic anhydrase isoforms in different tissues and found that four of them were more highly expressed in the hemocytes than in the gills or the mantle in C. gigas 22. In P. placenta, one of the perlucin family was identified by qRT-PCR and showed a highest expression in the digestive gland than that in the other tissues (Fig. 10e). The expression level of c81494_g1 unnigene (calmodulin-like) was the highest in the mouthpart, moderate in the gills, very low in the mantle tissue (Fig. 10f). The expression of carbonic anhydrase-like was the highest level in the gill tissue, medium in the mantle in P. placenta (Fig. 10g). Based on the qRT-PCR and in vitro crystallization results in the reported literature, we conclude that the above mentioned three unigenes, perlucin, calmodulin-like and carbonic anhydrate-like unigenes are potentially related to the biomineralization process of P. placenta.
As a member of the calmodulin subfamily of EF-hand Ca2+-binding proteins, Caltractin was first identified in C. gigas 56,98. Another unigene calreticulin is also a calcium-binding protein, it was primarily involved in the unfolded protein response to cellular stress (temperature, salinity, air exposure and heavy metals) in the endoplasmic reticulum 14,56,99. Both of the two calcium-binding proteins, calreticulin-like and caltractin-like exhibit relatively low expression in the mantle tissue according to the qRT-PCR results (Fig. 10h,i). However, we consider that they may probably have some kind of function for biomineralization process since they are calcium-binding proteins 100,101,102.
Conclusions
In conclusion, the transcript dataset of the mantle tissue of P. placenta was investigated in details by using Illumina HiSeqTM 4000 platform and public unigene databases. The identified and annotated unigenes provide valuable genomic resources for the understanding of the biomineralization mechanism. More than half of the annotated unigenes of the mantle tissue of P. placenta are consistent with those proteins from the pacific oyster C. gigas according to nr database. The transcripts of mantle tissue of P. placenta were identified with SNP, SSR and indel markers. These SNP markers, SSR markers and primers may be used in the construction of a genetic linkage map and gene-based association studies. 66 homologous unigenes of 21 shell matrix proteins in the transcriptome of mantle tissue of P. placenta were found to be related to the calcitic shell formation, while eighteen of the above unigenes are highly expressed with FPKM larger than 15 in the mantle tissue. Furthermore, qRT-PCR analysis for ten of highly expressed homologous unigenes (FPKM > 50) related to biomineralization from six different tissues of P. placenta indicate that seven of them are potentially related to the biomineralization process of the calcitic shells of P. placenta. Especially, the qRT-PCR analysis shows that four of ten examined unigenes including teneurin-2, gigasin-2, pif-like, tyrosinase-like unigenes have the highest expression levels in the mantle tissue than the levels in the other tissues, indicating their primary functions for biomineralization process. This study can contribute to the understanding of the molecular mechanisms and the functional components of the proteins that involve the biomineralization process of the calcite foliated plates of P. placenta. The transcriptomic data generated in this study provide a basis for further studies of P. placenta genome. Moreover, the comparison of potential biomineralization genes also reveals the similarities and differences between shell formation matrix of different molluscan animals.
Data availability
All relevant data reported here are included in the main section of the manuscript or in the supplementary materials. All described materials are available upon request.
Abbreviations
- qRT-PCR:
-
Quantitative real-time PCR
- SSR:
-
Simple sequence repeat
- SNP:
-
Single nucleotide polymorphisms
- Indel:
-
Insertion–deletion mutations
- KOG:
-
EuKaryotic Ortholog Groups
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GO:
-
Gene Ontology
- CDSs:
-
Coding regions
- nr:
-
NCBI non-redundant protein sequences
- Pfam:
-
Protein family
- SMPs:
-
Shell matrix proteins
- VWA:
-
Von willebrand factor type A
- FPKM:
-
Fragments per kilobase of feature per million mapped reads
References
Talham, D. R. Biomineralization: Principles and concepts in bioinorganic materials chemistry Stephen Mann. Cryst. Growth Des. 2, 675–675 (2002).
Zeichner-David, M. et al. Biosynthesis and characterization of rabbit tooth enamel extracellular-matrix proteins. Biochem. J. 251, 631–641 (1988).
Gize, A. On biomineralization. Mineral. Mag. 54, 344–345 (1990).
Jackson, D. et al. A rapidly evolving secretome builds and patterns a sea shell. BMC. Biol. 4, 40 (2006).
Marin, F. & Luquet, G. Molluscan shell proteins. C. R. Palevol. 3, 469–492 (2004).
Zou, Z. et al. A hydrated crystalline calcium carbonate phase: Calcium carbonate hemihydrate. Science 363, 396–400 (2019).
Furuhashi, T., Schwarzinger, C., Miksik, I., Smrz, M. & Beran, A. Molluscan shell evolution with review of shell calcification hypothesis. Comp. Biochem. Phys B. 154, 351–371 (2009).
Chateigner, D., Hedegaard, C. & Wenk, H.-R. Mollusc shell microstructures and crystallographic textures. J. Struct. Geol. 22, 1723–1735 (2000).
Belcher, A. et al. Control of crystal phase switching and orientation by soluble mollusc-shell proteins. Nature 381, 56–58 (1996).
Li, S. et al. Transcriptome and biomineralization responses of the pearl oyster Pinctada fucata to elevated CO2 and temperature. Sci. Rep. 6, 18943 (2016).
Gardner, L. D., Mills, D., Wiegand, A., Leavesley, D. & Elizur, A. Spatial analysis of biomineralization associated gene expression from the mantle organ of the pearl oyster Pinctada maxima. BMC Genomics 12, 455 (2011).
Zhang, C. & Zhang, R. Matrix proteins in the outer shells of molluscs. Mar. Biotechnol. 8, 572–586 (2006).
Kocot, K., Aguilera, F., McDougall, C., Jackson, D. & Degnan, B. Sea shell diversity and rapidly evolving secretomes: Insights into the evolution of biomineralization. Front. Zool. 13, 23 (2016).
Yang, Y. et al. The ER-localized Ca2+-binding protein calreticulin couples ER stress to autophagy by associating with microtubule-associated protein 1A/1B light chain 3. J. Biol. Chem. 294, 772–782 (2019).
Weiss, I. M., Kaufmann, S., Mann, K. & Fritz, M. Purification and characterization of perlucin and perlustrin, two new proteins from the shell of the Mollusc Haliotis laevigata. Biochem. Biophys. Res. Commun. 267, 17–21 (2000).
Tsukamoto, D., Sarashina, I. & Endo, K. Structure and expression of an unusually acidic matrix protein of pearl oyster shells. Biochem. Biophys. Res. Commun. 320, 1175–1180 (2004).
Kong, Y. et al. Cloning and characterization of Prisilkin-39, a novel matrix protein serving a dual role in the prismatic layer formation from the oyster Pinctada fucata. J. Biol. Chem. 284, 10841–10854 (2009).
Chen, Y. et al. Transcriptional regulation of the matrix protein Shematrin-2 during shell formation in pearl oyster. J. Biol. Chem. 293, 17803–17816 (2018).
Pan, C. et al. A novel acidic matrix protein, PfN44, stabilizes magnesium calcite to inhibit the crystallization of aragonite. J. Biol. Chem 289, 2776–2787 (2014).
Miyamoto, H. et al. A carbonic anhydrase from the nacreous layer in oyster pearls. Proc. Natl. Acad. Sci. 93, 9657–9660 (1996).
Kong, W., Li, S., **ang, L., **e, L. & Zhang, R. Calcium carbonate mineralization mediated by in vitro cultured mantle cells from Pinctada fucata. Biochem. Biophys. Res. Commun. 463, 1053–1058 (2015).
Ivanina, A. V., Falfushynska, H. I., Beniash, E., Piontkivska, H. & Sokolova, I. M. Biomineralization-related specialization of hemocytes and mantle tissues of the Pacific oyster Crassostrea gigas. J. Exp. Biol. 220, 3209–3221 (2017).
Mann, K., Weiss, I. M., Andre, S., Gabius, H. J. & Fritz, M. The amino-acid sequence of the abalone (Haliotis laevigata) nacre protein perlucin—Detection of a functional C-type lectin domain with galactose/mannose specificity. Eur. J. Biochem. 267, 5257–5264 (2000).
Miyamoto, H., Miyoshi, F. & Kohno, J. The carbonic anhydrase domain protein nacrein is expressed in the epithelial cells of the mantle and acts as a negative regulator in calcification in the mollusc Pinctada fucata. Zool. Sci. 22, 311–315 (2005).
Gotliv, B. A. et al. Asprich: A novel aspartic acid-rich protein family from the prismatic shell matrix of the bivalve Atrina rigida. ChemBioChem 6, 304–314 (2005).
Carpenter, K. E. & Niem, V. H. The living marine resources of the Western Central Pacific. F. A. O. 19, 359–363 (1998).
Runnegar, B. Crystallography of the foliated calcite shell layers of bivalve molluscs. Alcheringa 8, 273–290 (1984).
Checa, A. G., Esteban-Delgado, F. J. & Rodriguez-Navarro, A. B. Crystallographic structure of the foliated calcite of bivalves. J. Struct. Biol. 157, 393–402 (2007).
Li, L. & Ortiz, C. Biological design for simultaneous optical transparency and mechanical robustness in the shell of Placuna placenta. Adv. Mater. 25, 2344–2350 (2013).
Li, L. & Ortiz, C. Pervasive nanoscale deformation twinning as a catalyst for efficient energy dissipation in a bioceramic armour. Nat. Mater. 13, 501–507 (2014).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Shi, Y. et al. Transcriptome analysis of mantle tissues reveals potential biomineralization-related genes in Tectus pyramis Born. Comp. Biochem. Phys D. 29, 131–144 (2019).
Wang, X., Liu, Z. & Wu, W. Transcriptome analysis of the freshwater pearl mussel (Cristaria plicata) mantle unravels genes involved in the formation of shell and pearl. Mol. Genet. Genomics. 292, 343–352 (2017).
Li, H. et al. Characterization of transcriptome and identification of biomineralization genes in winged pearl oyster (Pteria penguin) mantle tissue. Comp. Biochem. Phys D. 21, 67–76 (2017).
Shi, Y. et al. Characterization of the pearl oyster (Pinctada martensii) mantle transcriptome unravels biomineralization genes. Mar. Biotechnol. 15, 175–187 (2013).
Sun, X., Yang, A., Wu, B., Zhou, L. & Liu, Z. Characterization of the mantle transcriptome of yesso scallop (Patinopecten yessoensis): identification of genes potentially involved in biomineralization and pigmentation. PLoS ONE 10, 4 (2015).
Shi, M. et al. Characterization of the Zhikong Scallop (Chlamys farreri) mantle transcriptome and identification of biomineralization-related genes. Mar. Biotechnol. 15, 706–715 (2013).
Freer, A., Bridgett, S., Jiang, J. & Cusack, M. Biomineral proteins from Mytilus edulis mantle tissue transcriptome. Mar. Biotechnol. 16, 34–45 (2014).
McGinty, E., Zenger, K., Jones, D. & Jerry, D. Transcriptome analysis of biomineralisation-related genes within the pearl sac: Host and donor oyster contribution. Mar. genomics. 5, 27–33 (2012).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Iseli, C., Jongeneel, C. & Bucher, P. ESTScan: A program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Int. Conf. Intell. Syst. Mol. Biol. 99, 138–148 (1999).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome. Res. 20, 1297–1303 (2010).
Suzuki, M. et al. Characterization of Prismalin-14, a novel matrix protein from the prismatic layer of the Japanese pearl oyster (Pinctada fucata). Biochem. J. 382, 205–213 (2004).
Suzuki, M. et al. An acidic matrix protein, pif, is a key macromolecule for nacre formation. Science 325, 1388–1390 (2009).
Su, J. et al. Structural characterization of amorphous calcium carbonate-binding protein: An insight into the mechanism of amorphous calcium carbonate formation. Biochem. J. 453, 179–186 (2013).
Kalamajski, S., Aspberg, A., Lindblom, K., Heinegård, D. & Oldberg, Å. Asporin competes with decorin for collagen binding, binds calcium and promotes osteoblast collagen mineralization. Biochem. J. 423, 53–59 (2009).
Punta, M. et al. The Pfam protein families database. Nucleic. Acids. Res. 40, D290–D301 (2012).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: Interactive sequence similarity searching. Nucleic. Acids. Res. 39, W29–W37 (2011).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Liu, J. et al. Microarray: a global analysis of biomineralization-related gene expression profiles during larval development in the pearl oyster, Pinctada fucata. BMC Genomics 16, 15 (2015).
Wang, X. T. et al. Oyster shell proteins originate from multiple organs and their probable transport pathway to the shell formation front. PLoS ONE 8, 6 (2013).
Mount, A. & Pickering, C. M. Testing the capacity of clothing to act as a vector for non-native seed in protected areas. J. Environ. Manag. 91, 168–179 (2009).
Mount, A. S., Wheeler, A. P., Paradkar, R. P. & Snider, D. Hemocyte-mediated shell mineralization in the eastern oyster. Science 304, 297–300 (2004).
Zhang, L. & Guo, X. Development and validation of single nucleotide polymorphism markers in the eastern oyster Crassostrea virginica Gmelin by mining ESTs and resequencing. Aquaculture 302, 124–129 (2010).
Song, X. R., Liu, Z. Q., Wang, L. L. & Song, L. S. Recent advances of shell matrix proteins and cellular orchestration in marine molluscan shell biomineralization. Front. Mar. Sci. 6, 16 (2019).
Zhang, G. F. et al. The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490, 49–54 (2012).
Ramos-Silva, P. et al. The skeletal proteome of the coral Acropora millepora: The evolution of calcification by co-option and domain shuffling. Mol. Biol. Evol. 30, 2099–2112 (2013).
Drake, J. L. et al. Proteomic analysis of skeletal organic matrix from the stony coral Stylophora pistillata. Proc. Natl. Acad. Sci. 110, 3788–3793 (2013).
Luo, Y.-J. et al. The Lingula genome provides insights into brachiopod evolution and the origin of phosphate biomineralization. Nat. Commun. 6, 8301 (2015).
Nong, W. et al. Genome of the ramshorn snail Biomphalaria straminea-an obligate intermediate host of schistosomiasis. GigaScience. 11 (2022).
Sarashina, I. Skeletal matrix proteins of invertebrate animals: Comparative analysis of their amino acid sequences. Paleont. Res. 10, 311–336 (2009).
Gorbushin, A. M. & Borisova, E. A. Lectin-like molecules in transcriptome of Littorina littorea hemocytes. Dev. Comp. Immunol. 48, 210–220 (2015).
McGinty, E. L., Zenger, K. R., Jones, D. B. & Jerry, D. R. Transcriptome analysis of biomineralisation-related genes within the pearl sac: Host and donor oyster contribution. Mar. Genom. 5, 27–33 (2012).
Zakrzewski, A. C. et al. Early divergence, broad distribution, and high diversity of animal chitin synthases. Genome Biol. Evol. 6, 316–325 (2014).
Zhang, S.-M. & Loker, E. Representation of an immune responsive gene family encoding fibrinogen-related proteins in the freshwater mollusc Biomphalaria glabrata, an intermediate host for Schistosoma mansoni. Gene 341, 255–266 (2004).
Zheng, L. et al. Ferritin has an important immune function in the ark shell Scapharca broughtonii. Dev. Comp. Immunol 59, 15–24 (2016).
**e, J. et al. Molecular and functional characterization of ferritin in abalone Haliotis diversicolor supertexta. Acta. Oceanologica. Sinica. 31 (2012).
Munro, H. et al. The ferritin genes: structure, expression, and regulation. Ann. N. Y. Acad. Sci. 526, 113–123 (2006).
Lin, J. Y., Ma, K. Y., Bai, Z. Y. & Li, J. L. Molecular cloning and characterization of perlucin from the freshwater pearl mussel, Hyriopsis cumingii. Gene 526, 210–216 (2013).
Asada, T., Ashiba, G. & Watanabe, S. Calcium sensitivy of foot muscle myosin from clam (Meretrix lusoria). J. Biochem. 85, 1543–1546 (1979).
Collins, J., Johnson, J. & Szent-Györgyi, A. Purification and characterization of a scallop sarcoplasmic calcium-binding protein. Biochemistry 22, 341–345 (1983).
Wang, S. et al. Scallop genome provides insights into evolution of bilaterian karyotype and development. Nat. Ecol. Evol. 1, 120 (2017).
Yang, J. et al. Transcriptome analysis of growth heterosis in pearl oyster Pinctada fucata martensii. FEBS Open Bio 8, 1794–1803 (2018).
Megrabian, Z. B. & Nalbandian, R. M. Copper-containing amine oxidases of blood vessels (review of the literature). Vopr. Med. Khim. 33, 13–18 (1987).
Piomelli, D., Shapiro, E., Feinmark, S. J. & Schwartz, J. H. Metabolites of arachidonic acid in the nervous system of Aplysia: possible mediators of synaptic modulation. J. Neurosci. 7, 3675–3686 (1987).
Voolstra, C. R. et al. Comparative analysis of the genomes of Stylophora pistillata and Acropora digitifera provides evidence for extensive differences between species of corals. Sci. Rep. 7, 17583 (2017).
Wang, X. et al. Evolution and functional analysis of the Pif97 gene of the Pacific oyster Crassostrea gigas. Curr. Zool. 59, 109–115 (2013).
Feng, D., Li, Q., Yu, H., Kong, L. & Du, S. Identification of conserved proteins from diverse shell matrix proteome in Crassostrea gigas: characterization of genetic bases regulating shell formation. Sci. Rep. 7, 45754 (2017).
Oliveira, D. V., Silva, T. S., Cordeiro, O. D., Cavaco, S. I. & Simes, D. C. Identification of proteins with potential osteogenic activity present in the water-soluble matrix proteins from Crassostrea gigas nacre using a proteomic approach. Sci. World. J. 2012, 765909–765909 (2012).
Marie, B., Le Roy, N., Zanella-Cléon, I., Becchi, M. & Marin, F. Molecular evolution of mollusc shell proteins: Insights from proteomic analysis of the edible mussel Mytilus. J. Mol. Evol. 72, 531–546 (2011).
Stenflo, J., Stenberg, Y. & Muranyi, A. Calcium-binding EGF-like modules in coagulation proteinases: Function of the calcium ion in module interactions. Biochim. Biophys. Acta. 1477, 51–63 (2000).
Marie, B. et al. The shell-forming proteome of Lottia gigantea reveals both deep conservations and lineage specific novelties. Febs. J. 280, 214–232 (2012).
Jovine, L., Darie, C., Litscher, E. & Wassarman, P. Zona pellucida domain proteins. Annu. Rev. Biochem. 74, 83–114 (2005).
Bjarnmark, N. A. et al. Transcriptomics provides insight into Mytilus galloprovincialis (Mollusca: Bivalvia) mantle function and its role in biomineralisation. Mar. Genomics. 27, 37–45 (2016).
Suzuki, M., Iwashima, A., Kimura, M., Kogure, T. & Nagasawa, H. The molecular evolution of the PIF family proteins in various species of mollusks. Mar. Biotechnol. 15, 145–158 (2013).
Bahn, S. Y., Jo, B. H., Choi, Y. S. & Cha, H. J. Control of nacre biomineralization by Pif80 in pearl oyster. Sci. Adv. 3, e1700765 (2017).
Nelson, M. R. & Chazin, W. J. Structures of EF-hand Ca2+-binding proteins: Diversity in the organization, packing and response to Ca2+ binding. Biometals 11, 297–318 (1998).
Özbek, S., Balasubramanian, P., Chiquet-Ehrismann, R., Tucker, R. & Adams, J. The evolution of extracellular matrix. Mol. Biol. Cell. 21, 4300–4305 (2010).
Miyashita, T., Takagi, R., Miyamoto, H. & Matsushiro, A. Identical carbonic anhydrase contributes to nacreous or prismatic layer formation in Pinctada fucata (Mollusca: Bivalvia). Veliger 45, 250–255 (2002).
Deng, Y. et al. De novo assembly, gene annotation, and simple sequence repeat marker development using Illumina paired-end transcriptome sequences in the pearl oyster Pinctada maxima. Biosci. Biotechnol. Biochem. 78, 1685–1692 (2014).
Minoche, A. E., Dohm, J. C. & Himmelbauer, H. Evaluation of genomic high-throughput sequencing data generated on illumina HiSeq and genome analyzer systems. Genome Biol. 12, R112 (2011).
Nagai, K., Yano, M., Morimoto, K. & Miyamoto, H. Tyrosinase localization in mollusc shells. Comp. Biochem. Phys B. 146, 207–214 (2007).
Aguilera, F., McDougall, C. & Degnan, B. M. Evolution of the tyrosinase gene family in bivalve molluscs: independent expansion of the mantle gene repertoire. Acta. Biomater. 10, 3855–3865 (2014).
Blank, S. et al. The nacre protein perlucin nucleates growth of calcium carbonate crystals. J. Microsc. 212, 280–291 (2003).
Yan, Z. et al. Biomineralization: functions of calmodulin-like protein in the shell formation of pearl oyster. Biochim. Biophys. Acta. 1770, 1338–1344 (2007).
Li, S., **e, L., Ma, Z. & Zhang, R. cDNA cloning and characterization of a novel calmodulin-like protein from pearl oyster Pinctada fucata. Febs. J. 272, 4899–4910 (2005).
Roy, N. L. E. et al. Identification of two carbonic anhydrases in the mantle of the European Abalone Haliotis tuberculata (Gastropoda, Haliotidae): phylogenetic implications. J. Exp. Zool. B 318, 353–367 (2012).
Veeraraghavan, S. et al. Structural independence of the two EF-hand domains of caltractin *. J. Biol. Chem. 277, 28564–28571 (2002).
Fan, W., Hu, Y., Li, C., **e, L. & Zhang, R. Cloning, Characterization, and expression analysis of Calreticulin from pearl oyster Pinctada fucata. Tsinghua. Sci. Technol. 13, 466–473 (2008).
Hattan, S. J., Laue, T. M. & Chasteen, N. D. Purification and characterization of a novel calcium-binding protein from the extrapallial fluid of the mollusc, Mytilus edulis. J. Biol. Chem. 276, 4461–4468 (2001).
Duplat, D. et al. Identification of calconectin, a calcium-binding protein specifically expressed by the mantle of Pinctada margaritifera. Febs. Lett. 580, 2435–2441 (2006).
Liu, H. L. et al. Identification and characterization of a biomineralization related gene PFMG1 highly expressed in the mantle of Pinctada fucata. Acs. Biochem. 46, 844–851 (2007).
Acknowledgements
This research was funded by National Natural Science Foundation of China (Grant No. 21877009), and Bei**g Institute of Technology Research Fund Program for Young Scholars. This work was also supported by the facilities at the Analysis & Testing Center, Bei**g Institute of Technology.
Author information
Authors and Affiliations
Contributions
N.S. performed the main experiments, data analysis, and wrote the paper. Y.M. designed the experiments and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, N., Li, J., Li, B. et al. Transcriptome analysis of the bivalve Placuna placenta mantle reveals potential biomineralization-related genes. Sci Rep 12, 4743 (2022). https://doi.org/10.1038/s41598-022-08610-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-08610-5
- Springer Nature Limited
This article is cited by
-
Transcriptome sequencing of the endangered land snail Karaftohelix adamsi from the Island Ulleung: De novo assembly, annotation, valuation of fitness genes and SSR markers
Genes & Genomics (2024)
-
A review of the endangered mollusks transcriptome under the threatened species initiative of Korea
Genes & Genomics (2023)