Abstract
This is the first study to compare genome-wide DNA methylation profiles of sorted blood cells from myelofibrosis (MF) patients and healthy controls. We found that differentially methylated CpG sites located to genes involved in ‘cancer’ and ‘embryonic development’ in MF CD34+ cells, in ‘inflammatory disease’ in MF mononuclear cells, and in ‘immunological diseases’ in MF granulocytes. Only few differentially methylated CpG sites were common among the three cell populations. Mutations in the epigenetic regulators ASXL1 (47%) and TET2 (20%) were not associated with a specific DNA methylation pattern using an unsupervised approach. However, in a supervised analysis of ASXL1 mutated versus wild-type cases, differentially methylated CpG sites were enriched in regions marked by histone H3K4me1, histone H3K27me3, and the bivalent histone mark H3K27me3 + H3K4me3 in human CD34+ cells. Hypermethylation of selected CpG sites was confirmed in a separate validation cohort of 30 MF patients by pyrosequencing. Altogether, we show that individual MF cell populations have distinct differentially methylated genes relative to their normal counterparts, which likely contribute to the phenotypic characteristics of MF. Furthermore, differentially methylated CpG sites in ASXL1 mutated MF cases are found in regulatory regions that could be associated with aberrant gene expression of ASXL1 target genes.
Similar content being viewed by others
Introduction
The chronic myeloproliferative neoplasms (MPNs) include the classical diseases myelofibrosis (MF), polycythemia vera (PV), and essential thrombocythemia (ET), with MF patients having the highest morbidity and mortality1. In addition to the expansion of one or more of the myeloid lineages, MF is characterized by progressive bone marrow fibrosis leading to extramedullary hematopoiesis and hepatosplenomegaly2.
The most commonly observed mutation in MF is JAK2V617F, which is found in 60% of MF patients3. Eight to 11% of JAK2V617F negative MF patients carry MPL mutations4, and both JAK2 and MPL mutations cause constitutive activation of the JAK/STAT pathway that promotes cell survival and proliferation5. More recently, mutations were identified in CALR that are mutually exclusive to JAK2 and MPL mutations in the majority of patients6, 7. In addition to causing constitutive activation of the JAK/STAT pathway7, mutated CALR lose the ability to bind calcium and retrieve and retain chaperone proteins to the endoplasmic reticulum6, 7. Although mutations in JAK2, MPL, and CALR are recurrent in MPN, they alone explain neither the pathogenesis nor the clinical manifestations associated with the distinctive MPN subgroups.
Mutations in epigenetic regulators, including ASXL1, TET2, DNMT3A, EED, EZH2, IDH1/2, JARID2, and SUZ12 have also been observed in MF8, 9, and expansion of the ASXL1 mutated clone has been associated with leukemic transformation10. However, despite high frequency of mutations in some of these genes, little is known about their impact on epigenetic regulation in MF. Few studies have investigated the genome-wide methylation patters in MF11, 12, and none of them have compared different MF cell populations.
Both TET2 and ASXL1 mutations have been associated with increased DNA methylation levels when analyzing neutrophils11, and unsorted cells from bone marrow and peripheral blood12. In addition, ASXL1 mutations were associated with a distinct DNA methylation signature11. Disruption of ASXL1 is frequent in myeloid malignancies with a prevalence of 20–30%13,14,15. In vivo analysis shows that hematopoiesis-specific loss of Asxl1 causes multi lineage cytopenia and dysplasia13 indicating its pivotal role in hematopoiesis. ASXL1 and BAP1 constitute a deubiquitination complex, where BAP1 catalyze the deubiquitination of H2AK119Ub16, 17. H2AK119Ub is a repressive histone mark deposited by the Polycomb Repressive Complex 1 (PRC1)18, both in a PRC2-dependent19 and independent manner20. Moreover, it was recently observed that H2AK119Ub could recruit components of the PRC2 complex to catalyze H3K27me321.
Since MF is a disease affecting several hematopoietic cell lineages, we investigated the genome-wide DNA methylation profiles of CD34+ cells, mononuclear cells, and granulocytes from 16 MF patients and 3 healthy age-matched controls. We further aimed to investigate whether distinct DNA methylation profiles are related to genetic aberrations of any of the epigenetic modifiers ASXL1, TET2, DNMT3A, IDH1, and IDH2.
Results
By comparison of individual MF cell populations to their normal counterparts isolated from healthy donors, we initially identified differentially methylated CpG sites within MF granulocytes, MF mononuclear cells and MF CD34+ cells, respectively.
MF granulocytes are hypomethylated relative to MF CD34+ cells and MF mononuclear cells
Based on the 504 most differentially methylated CpG sites with a standard deviation (SD) > 0.3 across all samples, a hierarchical cluster analysis clearly distinguished individual samples of MF mononuclear cells and MF CD34+ cells from MF granulocytes (Figure S1). In general, granulocytes were characterized by an overall low methylation level, which correlates to previous findings22. A single MF granulocyte sample (F16) clustered together with the mononuclear cells and CD34+ cells due to a higher overall methylation level. Three distinct clusters were observed in the granulocyte population; however, this could not be explained by mutations in any of the genes investigated.
Each MF cell compartment has a specific DNA methylation profile
A Venn diagram was used to illustrate the overlap of differentially methylated CpG sites between the MF granulocytes, MF mononuclear cells and MF CD34+ cells. The 200 most significantly differentially methylated CpG sites were included, and only a minor overlap was observed between the three MF cell populations (Fig. 1).
Venn diagram showing the overlap of differentially methylated CpG sites between MF cell populations. The CD34+ cell population is blue, the MF granulocyte population is yellow, and the MF mononuclear cell population is red. Five differentially methylated CpG sites overlapped between the three cell populations.
Aberrantly methylated genes in the MF CD34 + cell population
In the MF CD34+ cells, 1628 CpG sites annotated to 739 genes were differentially methylated (FDR p < 0.05; |Δβ| ± 0.2) when compared to their healthy counterparts (Table S2). Ingenuity pathway analysis revealed that differentially methylated CpG sites were annotated to genes involved in ‘cancer’ (e.g. WT1, BCL2, BIN1, GATA6, RUNX2, EGFR) and ‘embryonic development’ (e.g. WT1, BMP4, FOXC1, GATA4), ‘cell death and survival’ (e.g. BCL2, EGFR, BMP4) ‘hematopoiesis’ (e.g. BMP4), ‘cell cycle’ (e.g. EGFR, BMP4), and ‘hematological diseases’(e.g. JAK2) (Figure S2A).
Aberrantly methylated genes in the MF mononuclear cells
In MF mononuclear cells, 213 CpG sites annotated to 121 genes were differentially methylated (FDR p < 0.05; |Δβ| ± 0.2) when compared to their healthy counterparts (Table S3). Ingenuity pathway analysis revealed that differentially methylated CpG sites were annotated to genes involved in ‘cell cycling’ (e.g. NDRG1, NEDD1, and MAD1L1), ‘inflammatory diseases’ (e.g. PRTN3), and ‘cancer’ (PCDHA6, MUC4, and ATP2C2) (Figure S2B).
Aberrantly methylated genes in the MF granulocyte population
In the MF granulocytes, 519 CpG sites annotated to 303 genes were differentially methylated (FDR p < 0.05; |Δβ| ± 0.3) when compared to their healthy counterparts (Table S4). Ingenuity pathway analysis revealed that differentially methylated CpG sites were annotated to genes involved in ‘cancer’ (e.g. WT1, BIN1, PCDHA6, RIPK4, SOCS3, KTN1), ‘cellular growth and proliferation’ (e.g. mir-146, WT1, FOXP1, CEBPE, IGF2BP1, IGF1R, CASP8), ‘immunological diseases’ (e.g. BCL2L1 and MICA) and in ‘cell death and survival’ (e.g. SOCS3, mir-146, UHRF1, and CASP8) (Figure S2C).
Investigation of differentially methylated CpG sites in the validation cohort
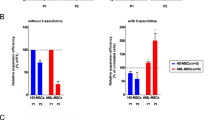
We selected four genes (LEP, TRIM59, WT1, and ZNF577) with at least two differentially methylated CpG sites in close proximity to the transcription start site for further validation. Differential methylation was confirmed using pyrosequencing for all genes in the MF validation cohort comprising 30 MF whole blood samples compared to 11 whole blood samples from healthy individuals (Fig. 2).
Validation of the methylated genes in a validation MF cohort. The DNA methylation level of 2–8 CpG sites annotated to four genes (ZNF577, WT1, LEP, and TRIM59) was validated in a validation MF cohort consisting of 30 MF patients where DNA had been isolated from whole blood. Hypermethylation of the ZNF577, LEP, and TRIM59 promoter regions and the WT1 gene body was verified using pyrosequencing (P ≤ 0.001 for all genes analyzed, Mann-Whitney test).
Somatic mutations in the MF cases
The mutational status of JAK2 was determined for all 16 MF patients (Table 1), whereas the mutational status of TET2, ASXL1, DNMT3A, IDH1, IDH2, CALR, and MPL was only determined in 15 patients due to limited material (Table 2). The most frequent mutations in the epigenetic regulators were nonsense mutations predicted to cause premature termination in ASXL1, which was observed in six patients (no. 1, 3, 5, 10, 14, and 15). Patient 7 had a missense mutation in ASXL1 causing the p.N986S substitution. Truncating mutations in the TET2 gene were observed for three patients (no. 1, 5, and 12), whereas two patients (no. 8 and 16) carried a previously unreported missense variant (c.1162 T > A) causing p.S388T. A skin biopsy from patient 8 was positive for the c.1162 T > A variant indicating its germ-line origin (data not shown). No mutations were identified in DNMT3A, IDH1 or IDH2.
Activating mutations of JAK2 were detected in 11/16 patients (no. 1, 4, 5, 7, 8, 9, 11, 12, 13, 14, and 15) whereas the frameshift CALR mutation p.L367fs*46, predicted to cause a C-terminal truncation, was observed in three patients (no. 3, 6, and 16). The activating MPL mutation p.W515L was detected in the two patients without JAK2 and CALR mutations (no. 2 and 10).
Unsupervised cluster analysis did not reveal a genome-wide specific DNA methylation profile associated with ASXL1 or TET2 mutations in MF granulocytes or CD34+ cells
An unsupervised clustering of granulocytes and CD34+ cells did not identify differential methylation signatures associated with ASXL1 and TET2 mutated cases (Figs 3 and S3). RPMM clustering of the 519 CpG sites differentially methylated among MF granulocyte samples and their healthy age-matched controls show three distinct clusters (Fig. 3). Cluster one (light blue) included samples from three patients, whereas cluster two (pink) included the three healthy age-matched controls and a single MF sample (F9). Cluster three (grey) included the remaining 12 MF samples. ASXL1 mutations were observed in one of two analyzed cases in cluster one and in 50% of patients in cluster 3, indicating that ASXL1 mutations do not seem to correlate with a specific DNA methylation profile using an unsupervised approach, which is in contrast to a previous study of 12 patients11.
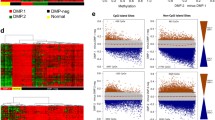
RPMM clustering of the granulocytes and their healthy age-matched counterparts with overlaid mutational status. Fifteen samples were analyzed for mutations in ASXL1, TET2, IDH1, IDH2, DNMT3A, CALR, JAK2, and MPL, while sample 13 was only analyzed for JAK2 mutations. The upper purple panel: Dynamic International Prognostic Scoring System (DIPSS). *The blast count was not available for MF patient F14. The middle black and red panel: Mutational status (black represents a mutation). Mutations were found for ASXL1, TET2, JAK2, CALR, and MPL. Lower panel: Hierarchical clustering of methylation levels in granulocytes from MF patients and healthy age-matched controls. β values range from 0 (blue; unmethylated) to 1 (red; methylated). Columns represent samples and rows represent differentially methylated CpG sites. Euclidean distance and complete linkage were used to study the cluster pattern of differential methylated probes. None of the mutations analyzed were associated with a DNA methylation-based subgrou**.
ASXL1 mutations are associated with differential DNA methylation of tumor suppressors and oncogenes in MF CD34+ cells
We next performed a supervised cluster analysis to investigate the association between mutated ASXL1 cases and aberrant DNA methylation in MF CD34+ cells. We identified 308 differentially methylated CpG sites (with FDR p < 0.05; |Δβ| ± 0.2) annotated to 174 genes (Table S5) associated with mutated ASXL1, which we named the “ASXL1 methylation signature” (Fig. 4). Of the 308 CpG sites 124 were hypermethylated while 184 were hypomethylated. In the granulocyte population a supervised cluster analysis identified 281 differentially methylated CpG sites (with FDR p < 0.05; |Δβ| ± 0.3) annotated to 137 genes (Table S6) associated with mutated ASXL1. Of the 281 CpG sites 105 were hypermethylated while 176 were hypomethylated. Several tumor suppressors and oncogenes including RASSF1, miR-663, ARID5B, FIP1L1, BCL6, TRPM2, ADORA1, ADORA2A, TFA2PA, and DIRC2 were differentially methylated in the ASXL1 mutated cases (Table S7).
Hierarchical clustering of 308 differentially methylated CpG sites in MF CD34+ cells associated with ASXL1 mutations using Pearson correlation and Average distance. Green indicates hypomethylated CpG sites and red indicates hypermethylated CpG sites. Columns represent the 15 MF patients analyzed. Rows represent differentially methylated CpG sites in MF CD34+ cells in ASXL1 mutated (n = 7) and ASXL1 non-mutated (_NM) (n = 8) cases.
ASXL1 mutations correlate with differential methylation in CpG rich regions in MF CD34+ cells
We mapped the 308 “ASXL1 methylation signature” CpG sites according to genomic regions. In samples with ASXL1 mutations, 36% of the differentially methylated CpG sites mapped to promoter regions, 31% to gene bodies, 30% to intergenic regions, and 3% to 3′UTRs (Fig. 5A). The majority of these regions were CG rich with 36% of the affected areas categorized as CpG islands and 37% as CpG shores (Fig. 5B).
The relative distribution of the differentially methylated CpG sites associated with ASXL1 mutations in CD34 + MF cells. (A) Functional genomic distribution in gene body, 3’UTR, intergenic, and promoter; and (B) map** according to the CpG density to islands, shelf, shore, and others/open sea. The majority of CpG sites were mapped to CpG shores (37%) and CpG islands (36%). (C) ASXL1 associated differentially methylated CpG sites were enriched in regions enriched for the histone marks H3K4me1 (P = 0.004), H3K27me3 (P = 1.10E-05), and the bivalent mark H3K27me3 plus H3K4me3 (P = 2.00E-03) in healthy CD34+ cells.
ASXL1 methylation signature probes are enriched in regions that carry H3K27me3, H3K4me1, and H3K27me3 plus H3K4me3 in CD34+ cells
Regions enriched with the repressive mark H3K27me3 and the bivalent histone mark H3K27me3 plus H3K4me3 were found to have a significantly higher number of differentially methylated CpG sites in ASXL1 mutated cases (Fig. 5C). Both hypo- and hypermethylation of CpG sites were observed in regions enriched for H3K27me3 and H3K27me3 plus H3K4me3 histone marks (Table S5). In addition, regions enriched with H3K4me1 in CD34+ cells were also found to have a significant higher number of differentially methylated CpG sites in patients with ASXL1 mutations compared to non-mutated ASXL1 cases (Fig. 5C) of which the majority of the CpG sites (93/120) were hypomethylated (Table S5).
Discussion
As MF involves both hematopoietic progenitors and more mature cells, a broad spectrum of cells throughout the myeloid compartment may be affected, but the contribution of the individual cell types to MF pathogenesis has not previously been explored. A previous study has shown correlation of ASXL1 mutations to a higher overall DNA methylation level and leukemic transformation in MF, whereas TET2 mutations correlated with increased DNA methylation levels of a distinct set of genes11, however, that study was based on the analyses of only 12 cases and needs confirmation in a larger cohort.
In our study, DNA methylation profiling of sorted MF cells and normal counterparts revealed that all three cell populations studied were characterized by distinct differential DNA methylation patterns. Interestingly, we observed that the majority of differentially methylated CpG sites were only differentially methylated in particular cellular compartments. This is likely indicative that, within each individual cell type, different DNA methylation patterns have a specific contribution to MF pathogenesis, rather than just being associated with lineage. To validate the genome-wide DNA methylation data a set of 4 genes (LEP, TRIM59, ZNF577, and WT1), that were found hypermethylated in the CD34 + compartment, were analyzed in a validation cohort of 30 MF patients where DNA had been isolated from whole blood. The fact that hypermethylation was confirmed in the validation cohort underline the presence of a MF specific methylation pattern, and opens up the potential of DNA methylation-based biomarkers for clinical purposes. Of the four genes analyzed in our validation cohort WT1 is especially interesting as it is found upregulated in myelofibrosis23, which corresponds to our finding of increased gene body methylation. WT1 has been shown to contribute to the plasticity of DNA methylation by recruiting TET2 to target genes causing site-specific demethylation24. A functional role of the remaining three genes LEP, TRIM59, and ZNF577 in MF pathogenesis still needs to be established and will require functional studies.
The MF CD34+ population had most differentially methylated CpG sites and, according to the pathway analyses, the genes with differentially methylated CpG sites were involved in ‘hematopoietic differentiation’, ‘cell-cycle’, ‘cell death and survival’, and ‘cancer’, probably contributing to the increased proliferation and dedifferentiation observed in these cells. The mononuclear cells had the lowest number of differentially methylated CpG sites, but interestingly, differentially methylated genes were associated with ‘immunological disease’, ‘cell death and survival’, and ‘cancer’. These aberrations may at least to some extent be linked to the high level of inflammation observed in MPN25, 26. Further sorting of the mononuclear cells could potentially reveal a more profound understanding of the contribution from the different subtypes. Genes that were found differentially methylated in the granulocytes were involved in ‘inflammatory disease’, ‘cell cycle’, ‘hematological disease’, and ‘cancer’. These data imply that specific characteristics of the malignant clones in the individual cellular compartments may contribute to different aspects of the MF phenotype.
Since DNA methylation has been associated with mutations in epigenetic regulators we next analyzed the mutational status of epigenetic regulators in our MF cohort. Sequencing analyses showed that ASXL1 was mutated in 7/15 (47%) patients, where a premature stop codon predicted to result in truncation in six cases, indicating that normal ASXL1 function may be lost. TET2 mutations were observed in 3/15 (20%). Thus, the frequency of mutations observed in our cohort is consistent with that of others, who report ASXL1 mutations in 20–55%14, 27, 28, and TET2 mutations in 14–20%29, 30. Mutations of IDH1, IDH2 and DNMT3A in MF are infrequent ranging from 0–7%14, 29,30,31,32.
With 7/15 patients having ASXL1 mutations, we aimed to investigate the ASXL1 mutation associated DNA methylation signature in MF. In contrast to a previous study of 12 patients11, we did not find ASXL1 mutations to be associated with an overall higher level of DNA methylation, or a distinct DNA methylation profile in either the granulocytes or the CD34+ cells using an unsupervised approach. However, when using supervised clustering analysis in the CD34+ cells a subset of differentially methylated CpG sites, frequently located in tumor suppressors and oncogenes, were found in ASXL1 mutated cases.
The majority of differentially methylated CpG sites associated with the “ASXL1 methylation signature” in CD34+ cells were enriched in regions with the repressive histone mark H3K27me3, whereas a minor proportion of the differentially methylated CpG sites were enriched in regions with the bivalent histone mark H3K27me3 plus H3K4me3. This is remarkable because ASXL1 has been suggested to regulate histone H3K27 methylation through interactions with the Polycomb-repressive complex 2 (PRC2). However, the association between ASXL1 mutation, H3K27me3, and differential DNA methylation is not straight forward and warrants further study. Differentially methylated CpG sites in ASXL1 mutated cases were also found in regions with the active histone mark H3K4me1, found at enhancer regions, which is likely to influence transcription of nearby genes. Most of the CpG sites overlap** with H3K4me1 were hypomethylated, and may thus possibly be associated with enhancer activation. Several genes previously recognized as tumor suppressors and oncogenes in other cancers including e.g. RASSF1, miR-663, ARID5B, FIP1L1, BCL6, TRPM2, ADORA1, ADORA2A, TFA2PA, and DIRC2 were among the “ASXL1 methylation signature genes”. Our data indicate that truncated ASXL1 is associated with methylation changes of a distinct set of cancer related genes that may be involved in disease progression, although no direct link between ASXL1 and DNA methylation has yet been established. Extending these analyses to TET2 mutated cases had been interesting but with only three MF patients carrying a TET2 mutation, of which two also had an ASXL1 mutation, we decided to focus on ASXL1 only. In addition, it would have been interesting to extent these analyses to JAK2 as JAK2 has been shown to influence the chromatin directly by phosphorylation of histone H3 tyrosine 4133. Indirectly, through the phosphorylation of PRMT5, mutated JAK2V617F has been shown to result in reduced methylation at histone H2A or H4 at R334. A direct link between JAK2 and DNA methylation is, however, missing and previous studies have not shown any association between JAK2 mutations and DNA methylation in myelofibrosis11, 12.
Taken together we found that aberrant and variable methylation patterns are present in the different myeloid cell compartments of MF patients. The differentially methylated CpG sites are annotated to several tumor suppressor genes and oncogenes, but also to genes involved mainly in inflammation and immunological diseases. Thus, the MF phenotype is likely a result of the aberrant function of distinct cell types throughout the myeloid lineages. In addition, we found that ASXL1 mutations are associated with DNA methylation changes in regulatory regions of cancer associated genes, not previously associated with MF. In future studies it shall be interesting to explore if there is a direct link between ASXL1 mediated gene regulation and aberrant methylation of these genes in malignant myelopoiesis.
Methods
Patient material and controls
This study is based on a primary cohort of 16 MF patients and three healthy age-matched controls and a validation cohort of 30 MF patients and 11 healthy controls. Clinical characteristics of the primary MF cohort including the Dynamic International Prognostic Scoring System (DIPSS) are shown in Table 1. For the primary cohort peripheral blood was separated into granulocytes and mononuclear cells using a Ficoll gradient. As a consequence of fibrotic bone marrow and extramedullary hematopoiesis, MF CD34+ cells could be isolated from the fraction of mononuclear cells using a CD34 + positive selection kit on a RoboSepTM platform (Stemcell Technologies, Grenoble, France). CD34+ cells from bone marrow, peripheral blood granulocytes, and mononuclear cells from three healthy age-matched individuals were used as controls. DNA was extracted using the AllPrep DNA/RNA Mini Kit (Qiagen, Hilden, Germany).
DNA from whole blood was extracted from the validation cohort using the Autopure LS (Qiagen) instrument and the Gentra Puregene Blood Kit (Qiagen), respectively.
The study was approved by the regional ethical committee (De Videnskabsetiske Komitéer Region Hovedstaden, Journal: H-C-2008–079) and all experiments were performed in accordance with the approved guidelines and regulations. All patients included had given a written informed consent.
Genome-wide DNA methylation profiling
Genome-wide DNA methylation profiling was performed using the 450 K Infinium array (Illumina Inc, San Diego, USA) platform as desribed previously35. This platform interrogates the methylation status of more than 480,000 CpGs in the human genome corresponding to 99% of NCBI RefSeq genes, which include CpGs in the promoters, enhancers, and gene bodies among others. In addition, the array covers CpG islands, shores and shelves of CpG islands. After hybridization and scanning of BeadChips, IDAT files were extracted to calculate the DNA methylation score (β values) ranging from 0 (non-methylated) to 1 (fully methylated) as described previously35.
Data filtering and normalization of DNA methylation data: Measurements in which the fluorescence intensity was not statistically significant above background signal were removed from the data set. Through an initial filtering process, probes corresponding to X and Y chromosomes and those containing a single nucleotide polymorphism (SNP) within five base pairs of targeted CpG sites were excluded. Probes with a repetitive element in the probe sequence within five bases of the targeted CpG site were also excluded. In total 361974 probes were used for further analysis.
Differential methylation between the MF samples and healthy control samples of individual CpG sites for each cell type was calculated. The probes with a FDR p < 0.05 in t-test and Δβ > ± 0.2 were considered to be differentially methylated as previously described35, with the exception of the granulocytes for which a mean Δβ > ± 0.3 was used with an adjusted p value < 0.01. For visualization, RPMM (recursively partitioned mixture model) and hierarchical clustering approaches were used. Hierarchial clustering using Euclidean distance and average linkage was used to classify samples into various groups as described previously36. All statistical analyses and clustering were performed using a R-statistical packages (https://www.r-project.org/) as described previously35, 37.
Pathway analysis
Functional interpretation of genes with one or more significantly differentially methylated CpG sites annotated was analyzed in the context of gene ontology and molecular networks by using Ingenuity pathway software (IPA; www.ingenuity.com) as described previously35.
Enrichment analysis
To investigate whether differentially methylated CpG sites were enriched in regions with distinct histone modifications, including H3K4me1, H3K4Me3, H3K4me3 plus H3K27me3, and H3K27me3, ChIP-seq data from human CD34+ cells was downloaded from NIH roadmap Epigenomics map** consortium (http://www.roadmapepigenomics.org/) and GSE36994, and the coordinates of ChIP-seq peaks were mapped to the 450 K probe locations. A hypergeometric test was used to evaluate possible enrichment of differentially methylated CpG sites in regions with distinct histone modifications.
Validation of differentially methylated sites using pyrosequencing
Four genes (LEP, TRIM59, WT1, and ZNF577) with at least two differentially methylated CpG sites in close proximity to the transcription start site (Table S1) were further analyzed in whole blood from a validation cohort of 30 MF patients and 11 healthy controls. Methylation independent (MIP) assays38 were designed using the PyroMark Assay Design 2.0 (Qiagen). The PCR amplicons were pyrosequenced on the PyroMark Q24 (Qiagen) instrument using the PyroMark Gold Q24 reagents (Qiagen) according to manufacturers’ instructions. For each of the four genes the DNA methylation level is calculated as the median DNA methylation level of the CpG sites included in the assay (Table S1). Primer sequences and PCR conditions are given in Table S1.
Mutation analysis
Mutation analyses were performed on DNA extracted from MF granulocytes. The primer sequences and assay conditions for the mutation analyses of the genes of interest have previously been published; TET2, IDH1, IDH2, DNMT3A35, JAK239, CALR7, and MPL40. ASXL1 exon 12 was analyzed for mutations as previously described15 with modifications for two assays (Table S1). M13 tagged primers were used for CALR, ASXL, and MPL.
Change history
20 November 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Spivak, J. L. & Silver, R. T. The revised World Health Organization diagnostic criteria for polycythemia vera, essential thrombocytosis, and primary myelofibrosis: an alternative proposal. Blood 112, 231–239, doi:10.1182/blood-2007-12-128454 (2008).
Barosi, G. Myelofibrosis with myeloid metaplasia: diagnostic definition and prognostic classification for clinical studies and treatment guidelines. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 17, 2954–2970 (1999).
Tefferi, A. & Pardanani, A. Myeloproliferative Neoplasms: A Contemporary Review. JAMA Oncol 1, 97–105, doi:10.1001/jamaoncol.2015.89 (2015).
Pardanani, A. D. et al. MPL515 mutations in myeloproliferative and other myeloid disorders: a study of 1182 patients. Blood 108, 3472–3476, doi:10.1182/blood-2006-04-018879 (2006).
Chaligne, R. et al. New mutations of MPL in primitive myelofibrosis: only the MPL W515 mutations promote a G1/S-phase transition. Leukemia 22, 1557–1566, doi:10.1038/leu.2008.137 (2008).
Nangalia, J. et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. The New England journal of medicine 369, 2391–2405, doi:10.1056/NEJMoa1312542 (2013).
Klampfl, T. et al. Somatic mutations of calreticulin in myeloproliferative neoplasms. The New England journal of medicine 369, 2379–2390, doi:10.1056/NEJMoa1311347 (2013).
Score, J. et al. Inactivation of polycomb repressive complex 2 components in myeloproliferative and myelodysplastic/myeloproliferative neoplasms. Blood 119, 1208–1213, doi:10.1182/blood-2011-07-367243 (2012).
Milosevic, J. D. & Kralovics, R. Genetic and epigenetic alterations of myeloproliferative disorders. International journal of hematology 97, 183–197, doi:10.1007/s12185-012-1235-2 (2013).
Ferrer-Marin, F. et al. Leukemic transformation driven by an ASXL1 mutation after a JAK2V617F-positive primary myelofibrosis: clonal evolution and hierarchy revealed by next-generation sequencing. Journal of hematology & oncology 6, 68, doi:10.1186/1756-8722-6-68 (2013).
Nischal, S. et al. Methylome profiling reveals distinct alterations in phenotypic and mutational subgroups of myeloproliferative neoplasms. Cancer Res 73, 1076–1085, doi:10.1158/0008-5472.CAN-12-0735 (2013).
Perez, C. et al. Aberrant DNA methylation profile of chronic and transformed classic Philadelphia-negative myeloproliferative neoplasms. Haematologica 98, 1414–1420, doi:10.3324/haematol.2013.084160 (2013).
Abdel-Wahab, O. et al. Deletion of Asxl1 results in myelodysplasia and severe developmental defects in vivo. The Journal of experimental medicine 210, 2641–2659, doi:10.1084/jem.20131141 (2013).
Brecqueville, M. et al. Mutation analysis of ASXL1, CBL, DNMT3A, IDH1, IDH2, JAK2, MPL, NF1, SF3B1, SUZ12, and TET2 in myeloproliferative neoplasms. Genes, chromosomes & cancer 51, 743–755, doi:10.1002/gcc.21960 (2012).
Gelsi-Boyer, V. et al. Mutations of polycomb-associated gene ASXL1 in myelodysplastic syndromes and chronic myelomonocytic leukaemia. British journal of haematology 145, 788–800, doi:10.1111/j.1365-2141.2009.07697.x (2009).
Scheuermann, J. C. et al. Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature 465, 243–247, doi:10.1038/nature08966 (2010).
Balasubramani, A. et al. Cancer-associated ASXL1 mutations may act as gain-of-function mutations of the ASXL1-BAP1 complex. Nature communications 6, 7307, doi:10.1038/ncomms8307 (2015).
Wang, H. et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature 431, 873–878, doi:10.1038/nature02985 (2004).
Levine, S. S. et al. The core of the polycomb repressive complex is compositionally and functionally conserved in flies and humans. Molecular and cellular biology 22, 6070–6078 (2002).
Tavares, L. et al. RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell 148, 664–678, doi:10.1016/j.cell.2011.12.029 (2012).
Blackledge, N. P. et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 157, 1445–1459, doi:10.1016/j.cell.2014.05.004 (2014).
Reinius, L. E. et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PloS one 7, e41361, doi:10.1371/journal.pone.0041361 (2012).
Guglielmelli, P. et al. Molecular profiling of CD34+ cells in idiopathic myelofibrosis identifies a set of disease-associated genes and reveals the clinical significance of Wilms’ tumor gene 1 (WT1). Stem Cells 25, 165–173, doi:10.1634/stemcells.2006-0351 (2007).
Rampal, R. et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep 9, 1841–1855, doi:10.1016/j.celrep.2014.11.004 (2014).
Hasselbalch, H. C. Chronic inflammation as a promotor of mutagenesis in essential thrombocythemia, polycythemia vera and myelofibrosis. A human inflammation model for cancer development? Leukemia research 37, 214–220, doi:10.1016/j.leukres.2012.10.020 (2013).
Hasselbalch, H. C. The role of cytokines in the initiation and progression of myelofibrosis. Cytokine & growth factor reviews 24, 133–145, doi:10.1016/j.cytogfr.2013.01.004 (2013).
Ricci, C. et al. ASXL1 mutations in primary and secondary myelofibrosis. British journal of haematology 156, 404–407, doi:10.1111/j.1365-2141.2011.08865.x (2012).
Stein, B. L. et al. Disruption of the ASXL1 gene is frequent in primary, post-essential thrombocytosis and post-polycythemia vera myelofibrosis, but not essential thrombocytosis or polycythemia vera: analysis of molecular genetics and clinical phenotypes. Haematologica 96, 1462–1469, doi:10.3324/haematol.2011.045591 (2011).
Brecqueville, M. et al. Rare mutations in DNMT3A in myeloproliferative neoplasms and myelodysplastic syndromes. Blood cancer journal 1, e18, doi:10.1038/bcj.2011.15 (2011).
Abdel-Wahab, O. et al. Concomitant analysis of EZH2 and ASXL1 mutations in myelofibrosis, chronic myelomonocytic leukemia and blast-phase myeloproliferative neoplasms. Leukemia: official journal of the Leukemia Society of America, Leukemia Research Fund, U.K 25, 1200–1202, doi:10.1038/leu.2011.58 (2011).
Tefferi, A. et al. IDH1 and IDH2 mutation studies in 1473 patients with chronic-, fibrotic- or blast-phase essential thrombocythemia, polycythemia vera or myelofibrosis. Leukemia: official journal of the Leukemia Society of America, Leukemia Research Fund, U.K 24, 1302–1309, doi:10.1038/leu.2010.113 (2010).
Abdel-Wahab, O. et al. DNMT3A mutational analysis in primary myelofibrosis, chronic myelomonocytic leukemia and advanced phases of myeloproliferative neoplasms. Leukemia: official journal of the Leukemia Society of America, Leukemia Research Fund, U.K 25, 1219–1220, doi:10.1038/leu.2011.82 (2011).
Dawson, M. A. et al. JAK2 phosphorylates histone H3Y41 and excludes HP1alpha from chromatin. Nature 461, 819–822, doi:10.1038/nature08448 (2009).
Liu, F. et al. JAK2V617F-mediated phosphorylation of PRMT5 downregulates its methyltransferase activity and promotes myeloproliferation. Cancer Cell 19, 283–294, doi:10.1016/j.ccr.2010.12.020 (2011).
Asmar, F. et al. Genome-wide profiling identifies a DNA methylation signature that associates with TET2 mutations in diffuse large B-cell lymphoma. Haematologica 98, 1912–1920, doi:10.3324/haematol.2013.088740 (2013).
Houseman, E. A. et al. Model-based clustering of DNA methylation array data: a recursive-partitioning algorithm for high-dimensional data arising as a mixture of beta distributions. BMC Bioinformatics 9, 365, doi:10.1186/1471-2105-9-365 (2008).
Andersen, C. L. et al. Whole-exome sequencing and genome-wide methylation analyses identify novel disease associated mutations and methylation patterns in idiopathic hypereosinophilic syndrome. Oncotarget 6, 40588–40597, doi:10.18632/oncotarget.5845 (2015).
Kristensen, L. S. & Hansen, L. L. PCR-based methods for detecting single-locus DNA methylation biomarkers in cancer diagnostics, prognostics, and response to treatment. Clinical chemistry 55, 1471–1483, doi:10.1373/clinchem.2008.121962 (2009).
Andersen, C. L. et al. A phase II study of vorinostat (MK-0683) in patients with polycythaemia vera and essential thrombocythaemia. British journal of haematology 162, 498–508, doi:10.1111/bjh.12416 (2013).
Chen, X. et al. Detection of MPL exon10 mutations in 103 Chinese patients with JAK2V617F-negative myeloproliferative neoplasms. Blood cells, molecules & diseases 47, 67–71, doi:10.1016/j.bcmd.2011.04.004 (2011).
Acknowledgements
We would like to thank you Konstantinos Dimopoulos for analyzing data and Anja Pedersen for technical assistance. This work was supported by University of Aarhus to HMN., The Danish Council for Strategic Research (to Danstem), the Novo Nordisk Foundation, the Danish Cancer Society and the van Andel Research Institute, Stand Up to Cancer, Epigenetics Dream Team to KG.
Author information
Authors and Affiliations
Contributions
H.M. Nielsen, L.S. Kristensen, and K. Grønbæk conceived and designed the experiments. C.L. Andersen, T.A. Kruse, M. Thomassen, T.S. Larsen, V. Skov, O.W. Bjerrum, L.L. Hansen and H.C. Hasselbalch provided, patient material, clinical informations and resources. H.M. Nielsen, M. Westman, and F. Asmar performed the experiments. H.M. Nielsen, M. Westman, L.S. Kristensen, F. Asmar, V. Punj, and K. Grønbæk were involved in data analysis. H.M.Nielsen, V. Punj, and K. Grønbæk provided a first draft of the manuscript. All authors participated in the writing and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
VP is consultant in bioinfreg.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nielsen, H., Andersen, C., Westman, M. et al. Epigenetic changes in myelofibrosis: Distinct methylation changes in the myeloid compartments and in cases with ASXL1 mutations. Sci Rep 7, 6774 (2017). https://doi.org/10.1038/s41598-017-07057-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07057-3
- Springer Nature Limited
Keywords
This article is cited by
-
Genome-wide DNA methylation profiling is able to identify prefibrotic PMF cases at risk for progression to myelofibrosis
Clinical Epigenetics (2021)
-
Droplet digital PCR for the quantification of Alu methylation status in hematological malignancies
Diagnostic Pathology (2018)