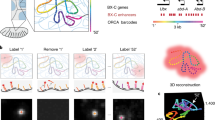
Abstract
DNA fluorescence in situ hybridization (FISH) has been a central technique in advancing our understanding of how chromatin is organized within the nucleus. With the increasing resolution offered by super-resolution microscopy, the optimal maintenance of chromatin structure within the nucleus is essential for accuracy in measurements and interpretation of data. However, standard 3D-FISH requires potentially destructive heat denaturation in the presence of chaotropic agents such as formamide to allow access to the DNA strands for labeled FISH probes. To avoid the need to heat-denature, we developed Resolution After Single-strand Exonuclease Resection (RASER)-FISH, which uses exonuclease digestion to generate single-stranded target DNA for efficient probe binding over a 2 d process. Furthermore, RASER-FISH is easily combined with immunostaining of nuclear proteins or the detection of RNAs. Here, we provide detailed procedures for RASER-FISH in mammalian cultured cells to detect single loci, chromatin tracks and topologically associating domains with conventional and super-resolution 3D structured illumination microscopy. Moreover, we provide a validation and characterization of our method, demonstrating excellent preservation of chromatin structure and nuclear integrity, together with improved hybridization efficiency, compared with classic 3D-FISH protocols.






Similar content being viewed by others
Data availability
Figures 2, 3 and 6 have associated raw image data plus one dataset. All raw data files are archived in Figshare: Fig. 2 at https://doi.org/10.6084/m9.figshare.16778899, Fig. 3 at https://doi.org/10.6084/m9.figshare.16778902 and Fig. 6 at https://doi.org/10.6084/m9.figshare.16755394. Source data are provided with this paper.
References
Benabdallah, N. S. et al. Decreased enhancer–promoter proximity accompanying enhancer activation. Mol. Cell 76, 473–484 e477 (2019).
Cattoni, D. I. et al. Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions. Nat. Commun. 8, 1753 (2017).
Fabre, P. J. et al. Nanoscale spatial organization of the HoxD gene cluster in distinct transcriptional states. Proc. Natl Acad. Sci. USA 112, 13964–13969 (2015).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
van de Corput, M. P. et al. Super-resolution imaging reveals three-dimensional folding dynamics of the beta-globin locus upon gene activation. J. Cell Sci. 125, 4630–4639 (2012).
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science https://doi.org/10.1126/science.aau1783 (2018).
Cardozo Gizzi, A. M. et al. Microscopy-based chromosome conformation capture enables simultaneous visualization of genome organization and transcription in intact organisms. Mol. Cell 74, 212–222.e215 (2019).
Mateo, L. J. et al. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 568, 49–54 (2019).
Su, J.-H., Zheng, P., Kinrot, S. S., Bintu, B. & Zhuang, X. Genome-scale imaging of the 3D organization and transcriptional activity of chromatin. Cell 182, 1641–1659.e1626 (2020).
Takei, Y. et al. Integrated spatial genomics reveals global architecture of single nuclei. Nature 590, 344–350 (2021).
Markaki, Y. et al. The potential of 3D-FISH and super-resolution structured illumination microscopy for studies of 3D nuclear architecture: 3D structured illumination microscopy of defined chromosomal structures visualized by 3D (immuno)-FISH opens new perspectives for studies of nuclear architecture. Bioessays 34, 412–426 (2012).
Hausmann, M., Lee, J. H., Sievers, A., Krufczik, M. & Hildenbrand, G. COMBinatorial Oligonucleotide FISH (COMBO-FISH) with uniquely binding repetitive DNA probes. Methods Mol. Biol. 2175, 65–77 (2020).
Lee, J.-H. et al. COMBO-FISH: a versatile tool beyond standard FISH to study chromatin organization by fluorescence light microscopy. OBM Genet. https://doi.org/10.21926/obm.genet.1901064 (2019).
Krufczik, M. et al. Combining low temperature fluorescence DNA-hybridization, immunostaining, and super-resolution localization microscopy for nano-structure analysis of ALU elements and their influence on chromatin structure. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18051005 (2017).
Deng, W., Shi, X., Tjian, R., Lionnet, T. & Singer, R. H. CASFISH: CRISPR/Cas9-mediated in situ labeling of genomic loci in fixed cells. Proc. Natl Acad. Sci. USA 112, 11870–11875 (2015).
Wang, Y. et al. Genome oligopaint via local denaturation fluorescence in situ hybridization. Mol. Cell 81, 1566–1577 e1568 (2021).
Bailey, S. M., Goodwin, E. H. & Cornforth, M. N. Strand-specific fluorescence in situ hybridization: the CO-FISH family. Cytogenet. Genome Res. 107, 14–17 (2004).
Goodwin, E. & Meyne, J. Strand-specific FISH reveals orientation of chromosome 18 alphoid DNA. Cytogenet. Cell Genet. 63, 126–127 (1993).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Schoenfelder, S. & Fraser, P. Long-range enhancer–promoter contacts in gene expression control. Nat. Rev. Genet. 20, 437–455 (2019).
van Steensel, B. & Furlong, E. E. M. The role of transcription in sha** the spatial organization of the genome. Nat. Rev. Mol. Cell Biol. 20, 327–337 (2019).
Hua, P. et al. Defining genome architecture at base-pair resolution. Nature 595, 125–129 (2021).
Osterwalder, M. et al. Enhancer redundancy provides phenotypic robustness in mammalian development. Nature 554, 239–243 (2018).
Oudelaar, A. M. et al. Dynamics of the 4D genome during in vivo lineage specification and differentiation. Nat. Commun. 11, 2722 (2020).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Dekker, J. Map** the 3D genome: aiming for consilience. Nat. Rev. Mol. Cell Biol. 17, 741–742 (2016).
Falk, M. et al. Heterochromatin drives compartmentalization of inverted and conventional nuclei. Nature 570, 395–399 (2019).
Feodorova, Y., Falk, M., Mirny, L. A. & Solovei, I. Viewing nuclear architecture through the eyes of nocturnal mammals. Trends Cell Biol. 30, 276–289 (2020).
Finn, E. H. et al. Extensive heterogeneity and intrinsic variation in spatial genome organization. Cell 176, 1502–1515 e1510 (2019).
Nir, G. et al. Walking along chromosomes with super-resolution imaging, contact maps, and integrative modeling. PLoS Genet. 14, e1007872 (2018).
Szabo, Q. et al. TADs are 3D structural units of higher-order chromosome organization in Drosophila. Sci. Adv. 4, eaar8082 (2018).
Brown, J. M. et al. A tissue-specific self-interacting chromatin domain forms independently of enhancer–promoter interactions. Nat. Commun. 9, 3849 (2018).
Miron, E. et al. Chromatin arranges in chains of mesoscale domains with nanoscale functional topography independent of cohesin. Sci. Adv. https://doi.org/10.1126/sciadv.aba8811 (2020).
Ochs, F. et al. Stabilization of chromatin topology safeguards genome integrity. Nature 574, 571–574 (2019).
Rhodes, J. D. P. et al. Cohesin disrupts polycomb-dependent chromosome interactions in embryonic stem cells. Cell Rep. 30, 820–835 e810 (2020).
Beckwith, K. et al. Visualization of loop extrusion by DNA nanoscale tracing in single human cells. Preprint at bioRxiv https://doi.org/10.1101/2021.04.12.439407 (2021).
Weiland, Y., Lemmer, P. & Cremer, C. Combining FISH with localisation microscopy: super-resolution imaging of nuclear genome nanostructures. Chromosome Res. 19, 5–23 (2011).
Kapuscinski, J. & Szer, W. Interactions of 4′, 6-diamidine-2-phenylindole with synthetic polynucleotides. Nucleic Acids Res. 6, 3519–3534 (1979).
Krasin, F. & Hutchinson, F. Double-strand breaks from single photochemical events in DNA containing 5-bromouracil. Biophys. J. 24, 645–656 (1978).
Limoli, C. L. & Ward, J. F. A new method for introducing double-strand breaks into cellular DNA. Radiat. Res. 134, 160–169 (1993).
Weghorst, C. M., Henneman, J. R. & Ward, J. M. Dose response of hepatic and renal DNA synthetic rates to continuous exposure of bromodeoxyuridine (BrdU) via slow-release pellets or osmotic minipumps in male B6C3F1 mice. J. Histochem. Cytochem. 39, 177–184 (1991).
Hutchinson, F. The lesions produced by ultraviolet light in DNA containing 5-bromouracil. Q. Rev. Biophys. 6, 201–246 (1973).
Kraus, F. et al. Quantitative 3D structured illumination microscopy of nuclear structures. Nat. Protoc. 12, 1011–1028 (2017).
Schmidt, T. L. et al. Scalable amplification of strand subsets from chip-synthesized oligonucleotide libraries. Nat. Commun. 6, 8634 (2015).
Beliveau, B. J. et al. Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc. Natl Acad. Sci. USA 109, 21301–21306 (2012).
Boettiger, A. & Murphy, S. Advances in chromatin imaging at kilobase-scale resolution. Trends Genet. 36, 273–287 (2020).
Jez, M. et al. The hazards of DAPI photoconversion: effects of dye, mounting media and fixative, and how to minimize the problem. Histochem. Cell Biol. 139, 195–204 (2013).
Brown, J. M. & Buckle, V. J. Detection of nascent RNA transcripts by fluorescence in situ hybridization. Methods Mol. Biol. 659, 33–50 (2010).
Boyle, S., Rodesch, M. J., Halvensleben, H. A., Jeddeloh, J. A. & Bickmore, W. A. Fluorescence in situ hybridization with high-complexity repeat-free oligonucleotide probes generated by massively parallel synthesis. Chromosome Res. 19, 901–909 (2011).
Brown, J. M. et al. Association between active genes occurs at nuclear speckles and is modulated by chromatin environment. J. Cell Biol. 182, 1083–1097 (2008).
Acknowledgements
We thank C. Lagerholm for extensive imaging support and D. Higgs for long-term support during the development of this technique. We thank R. Klose, J. and C. Lukas, F. Ochs, D. Higgs and J. Hughes for cells and images prepared during collaborations with them. We thank T. Brown and A. El-Sagheer for development and oversight of the oligonucleotide probe generation, and E. Heard for BAC RP24-217l10. Work in the Buckle laboratory was supported by MRC grants MC_UU_00016/1 and MR/N00969X/1, the latter in collaboration with J. Hughes, and by BBSRC grant BB/L01811X held in collaboration with T. Brown (Department of Physical Chemistry, Oxford University) and further supported by the Wolfson Imaging Centre Oxford funded by the Wolfson Foundation 18272, joint MRC/BBSRC/EPSRC MR/K015777X/1, Wellcome Trust Multi-User Equipment 104924/Z/14/Z. 3D-SIM imaging was performed at the Micron Oxford Advanced Bioimaging Unit funded by a Wellcome Trust Strategic Award 091911 and 107457/Z/15/Z. L.S. further acknowledges support by the EU Horizon 2020 Research and Innovation Program under the Marie Sklodowska-Curie grant agreement no. 766181. E.P. was sponsored by the International Internship Program (IIP) at Princeton University.
Author information
Authors and Affiliations
Contributions
J.M.B. and V.J.B. developed the protocol and hybridized, imaged and analyzed RASER-FISH preparations, S.D.O. synthesized, hybridized and analyzed the oligonucleotide probe preparations, L.S. and E.P. hybridized, imaged and analyzed preparations using structured illumination miroscopy, and J.M.B., L.S. and V.J.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Peer review
Peer review information
Nature Protocols thanks Marion Cremer, Michael Hausmann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Brown, J. et al. Nat. Commun. 9, 3849 (2018): https://doi.org/10.1038/s41467-018-06248-4
Ochs, F. et al. Nature 574, 571–574 (2019): https://doi.org/10.1038/s41586-019-1659-4
Miron, E. et al. Sci. Adv. 6, eaba8811 (2020): https://doi.org/10.1126/sciadv.aba8811
Supplementary information
Supplementary Information
Supplementary Methods 1 and 2 and Supplementary References.
Source data
Source Data Fig. 2
Statistical source data.
Rights and permissions
About this article
Cite this article
Brown, J.M., De Ornellas, S., Parisi, E. et al. RASER-FISH: non-denaturing fluorescence in situ hybridization for preservation of three-dimensional interphase chromatin structure. Nat Protoc 17, 1306–1331 (2022). https://doi.org/10.1038/s41596-022-00685-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00685-8
- Springer Nature Limited
This article is cited by
-
Enhancer selectivity in space and time: from enhancer–promoter interactions to promoter activation
Nature Reviews Molecular Cell Biology (2024)
-
From feulgen to modern methods: marking a century of DNA imaging advances
Histochemistry and Cell Biology (2024)