Abstract
The targeted insertion and stable expression of a large genetic payload in primary human cells demands methods that are robust, efficient and easy to implement. Large payload insertion via retroviruses is typically semi-random and hindered by transgene silencing. Leveraging homology-directed repair to place payloads under the control of endogenous essential genes can overcome silencing but often results in low knock-in efficiencies and cytotoxicity. Here we report a method for the knock-in and stable expression of a large payload and for the simultaneous knock-in of two genes at two endogenous loci. The method, which we named CLIP (for ‘CRISPR for long-fragment integration via pseudovirus’), leverages an integrase-deficient lentivirus encoding a payload flanked by homology arms and ‘cut sites’ to insert the payload upstream and in-frame of an endogenous essential gene, followed by the delivery of a CRISPR-associated ribonucleoprotein complex via electroporation. We show that CLIP enables the efficient insertion and stable expression of large payloads and of two difficult-to-express viral antigens in primary T cells at low cytotoxicity. CLIP offers a scalable and efficient method for manufacturing engineered primary cells.
Similar content being viewed by others
Main
Maximizing the therapeutic potential and improving the efficacy and safety of engineered cells often require the expression of large genes or of complex gene circuits1. For example, chimeric antigen receptor (CAR) T cells leverage a single engineered receptor to rewire its cytotoxicity towards cancers2,3. Efforts to improve their functionality, reduce toxicity or move beyond blood cancer into solid tumours, ageing, autoimmunity and viral clearance have leveraged combinations of tools that increase the ability of T cells to sense, process and respond to the disease. Biomolecular tools such as next-generation receptors4,5,6, clustered regularly interspaced short palindromic repeats (CRISPR) activation (CRISPRa) and CRISPR interference (CRISPRi)7,8,9,10, and logic gatesReporting summary Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the results in this study are available within the paper and its Supplementary Information. The raw and analysed immunopeptidome datasets generated during the study are too large to be publicly shared, yet they are available for research purposes from the corresponding author on reasonable request. Key constructs and plasmids will be available on Addgene (https://www.addgene.org/Stanley_Qi/). Source data are provided with this paper.
References
Cubillos-Ruiz, A. et al. Engineering living therapeutics with synthetic biology. Nat. Rev. Drug Discov. 20, 941–960 (2021).
Lim, W. A. & June, C. H. The principles of engineering immune cells to treat cancer. Cell 168, 724–740 (2017).
Labanieh, L., Majzner, R. G. & Mackall, C. L. Programming CAR-T cells to kill cancer. Nat. Biomed. Eng. 2, 377–391 (2018).
Kipniss, N. H. et al. Engineering cell sensing and responses using a GPCR-coupled CRISPR–Cas system. Nat. Commun. 8, 2212 (2017).
Roybal, K. T. et al. Engineering T cells with customized therapeutic response programs using synthetic Notch receptors. Cell 167, 419–432.e416 (2016).
Scheller, L., Strittmatter, T., Fuchs, D., Bojar, D. & Fussenegger, M. Generalized extracellular molecule sensor platform for programming cellular behavior. Nat. Chem. Biol. 14, 723–729 (2018).
Nakamura, M., Gao, Y., Dominguez, A. A. & Qi, L. S. CRISPR technologies for precise epigenome editing. Nat. Cell Biol. 23, 11–22 (2021).
Xu, X. & Qi, L. S. A CRISPR–dCas toolbox for genetic engineering and synthetic biology. J. Mol. Biol. 431, 34–47 (2019).
Dominguez, A. A., Lim, W. A. & Qi, L. S. Beyond editing: repurposing CRISPR–Cas9 for precision genome regulation and interrogation. Nat. Rev. Mol. Cell Biol. 17, 5–15 (2016).
Schmidt, R. et al. CRISPR activation and interference screens in primary human T cells decode cytokine regulation. Science https://doi.org/10.1126/science.abj4008 (2021).
Gao **ao**g, J., Chong Lucy, S., Kim Matthew, S. & Elowitz Michael, B. Programmable protein circuits in living cells. Science 361, 1252–1258 (2018).
Kempton, H. R., Goudy, L. E., Love, K. S. & Qi, L. S. Multiple input sensing and signal integration using a split Cas12a system. Mol. Cell 78, 184–191.e183 (2020).
Nissim, L. et al. Synthetic RNA-based immunomodulatory gene circuits for cancer immunotherapy. Cell 171, 1138–1150.e1115 (2017).
Herbst, F. et al. Extensive methylation of promoter sequences silences lentiviral transgene expression during stem cell differentiation in vivo. Mol. Ther. 20, 1014–1021 (2012).
Jones, S. et al. Lentiviral vector design for optimal T cell receptor gene expression in the transduction of peripheral blood lymphocytes and tumor-infiltrating lymphocytes. Hum. Gene Ther. 20, 630–640 (2009).
Ellis, J. Silencing and variegation of gammaretrovirus and lentivirus vectors. Hum. Gene Ther. 16, 1241–1246 (2005).
Klug, C. A., Cheshier, S. & Weissman, I. L. Inactivation of a GFP retrovirus occurs at multiple levels in long-term repopulating stem cells and their differentiated progeny. Blood 96, 894–901 (2000).
Golding, M. C., Zhang, L. & Mann, M. R. W. Multiple epigenetic modifiers induce aggressive viral extinction in extraembryonic endoderm stem cells. Cell Stem Cell 6, 457–467 (2010).
Müller-Kuller, U. et al. A minimal ubiquitous chromatin opening element (UCOE) effectively prevents silencing of juxtaposed heterologous promoters by epigenetic remodeling in multipotent and pluripotent stem cells. Nucleic Acids Res. 43, 1577–1592 (2015).
Wiebking, V. et al. Metabolic engineering generates a transgene-free safety switch for cell therapy. Nat. Biotechnol. 38, 1441–1450 (2020).
Martin, R. M. et al. Highly efficient and marker-free genome editing of human pluripotent stem cells by CRISPR–Cas9 RNP and AAV6 donor-mediated homologous recombination. Cell Stem Cell 24, 821–828.e825 (2019).
Bak, R. O., Dever, D. P. & Porteus, M. H. CRISPR/Cas9 genome editing in human hematopoietic stem cells. Nat. Protoc. 13, 358–376 (2018).
Roth, T. L. et al. Reprogramming human T cell function and specificity with non-viral genome targeting. Nature 559, 405–409 (2018).
Nguyen, D. N. et al. Polymer-stabilized Cas9 nanoparticles and modified repair templates increase genome editing efficiency. Nat. Biotechnol. 38, 44–49 (2020).
Banasik, M. B. & McCray, P. B. Integrase-defective lentiviral vectors: progress and applications. Gene Ther. 17, 150–157 (2010).
Wanisch, K. & Yáñez-Muñoz, R. J. Integration-deficient lentiviral vectors: a slow coming of age. Mol. Ther. 17, 1316–1332 (2009).
Roberts, B. et al. Systematic gene tagging using CRISPR/Cas9 in human stem cells to illuminate cell organization. Mol. Biol. Cell 28, 2854–2874 (2017).
Jiang, F. & Doudna, J. A. CRISPR–Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Song, F. & Stieger, K. Optimizing the DNA donor template for homology-directed repair of double-strand breaks. Mol. Ther. Nucleic Acids 7, 53–60 (2017).
Suzuki, K. & Izpisua Belmonte, J. C. In vivo genome editing via the HITI method as a tool for gene therapy. J. Hum. Genet. 63, 157–164 (2018).
Kantor, B. et al. Notable reduction in illegitimate integration mediated by a PPT-deleted, nonintegrating lentiviral vector. Mol. Ther. 19, 547–556 (2011).
Pavel-Dinu, M. et al. Gene correction for SCID-X1 in long-term hematopoietic stem cells. Nat. Commun. 10, 1634 (2019).
Zhao, Y. et al. High-efficiency transfection of primary human and mouse T lymphocytes using RNA electroporation. Mol. Ther. 13, 151–159 (2006).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676.e614 (2018).
Wessels, H.-H. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat. Biotechnol. 38, 722–727 (2020).
Gerby, B. et al. Optimized gene transfer into human primary leukemic T cell with NOD-SCID/leukemia-initiating cell activity. Leukemia 24, 646–649 (2010).
Guo, L. Y. et al. Multiplexed genome regulation in vivo with hyper-efficient Cas12a. Nat. Cell Biol. 24, 590–600 (2022).
Abbott, T. R. et al. Development of CRISPR as an antiviral strategy to combat SARS-CoV-2 and influenza. Cell 181, 865–876.e812 (2020).
Acknowledgements
We thank V. Wiebking and M. Proteus for providing IL2RG knock-in plasmids. We thank Stanford Blood Center for providing peripheral blood mononuclear cells for isolating primary T cells. We thank Cayman Chemicals for performing MHC-I immunopeptidomics. We thank V. Ramani for help designing off-target experiments. M.C. acknowledges support from NSF GRFP, ARCS Foundation, NIH F31 fellowships and Siebel Scholarship. L.S.Q. acknowledges support from Li Ka Shing Foundation, National Science Foundation CAREER award (award number 2046650), Stanford Maternal & Child Health Research Institute (MCHRI) and California Institute for Regenerative Medicine (CIRM, DISC2-12669). The work is supported by a Li Ka Shing Foundation gift fund, National Science Foundation CAREER award (award number 2046650) and Stanford Maternal & Child Health Research Institute (MCHRI). L.S.Q. is a Chan Zuckerberg Biohub – San Francisco Investigator.
Author information
Authors and Affiliations
Contributions
M.C. and L.S.Q. conceived of the idea and designed the experiments. M.C., D.A.R. and X.C. performed the experiments. M.C. and L.S.Q. analysed the experimental data. M.C. wrote the manuscript with input from L.S.Q. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a related patent via Stanford University PCT/US2021/072335. L.S.Q. is a founder of Epic Bio and a scientific advisor of Kytopen, and Laboratory of Genomics Research. M.C. is a co-founder of Enoda Cellworks Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
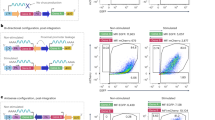
Extended Data Fig. 1 CLIP’s cut sites enable more efficient HDR than IDLV.
CLIP strategy compared to IDLV (same as CLIP but without inclusion of cut sites) with and without RNP 3, 5, and 7 days after electroporation in K562s. The high expression peak is indicative of a targeted knock-in while the low expression peak is indicative of transient episomal expression of the reverse transcribed viral genome. Data from two biological replicates each with technical duplicates (replicate 1 shown in Fig. 2c).
Extended Data Fig. 2 CLIP is robust to protocol perturbation in primary T cells.
a, CLIP pseudovirus transduced into primary T cells 48 hours or 24 hours before electroporation of RNP showed minimal effect on transduction efficiency. b, 2.5×105, 5×105, or 1×106 cells were aliquoted into a cuvette for electroporation of Cas9-RNP, which had minimal effect on the knock-in efficiency. Data from one donor with technical duplicate.
Extended Data Fig. 3 Pseudovirus delivers genetic payloads with low toxicity to primary T cells.
Extended Data Fig. 4 CLIP integrates Cas13d into both ACTB and IL2RG, but is more highly expressed in ACTB.
a, Expression of Cas13d in either the ACTB or IL2RG locus in K562s (2 biological replicates with technical duplicates). b, Apparent knock-in efficiency is lower in IL2RG than ACTB (left) and the magnitude of expression of Cas13 is also lower in IL2RG than ACTB (right) (****: p < 0.0001) c, gDNA from K562s that underwent Cas13d CLIP in both loci was isolated. PCR was performed using a forward primer targeting Cas13d and reverse targeting IL2RG downstream of homology arm (left, expected band: 1101 bp) or using a forward primer targeting Cas13d and reverse targeting ACTB downstream of homology arm (right, expected band: 1059 bp). Data is from two biological replicates each with two technical replicates.
Extended Data Fig. 5 CLIP enables Cas13d stabilization in primary T cells.
a, Magnitude of expression at Day 7 when GFP was knocked into IL2RG or ACTB in primary T cells (****: p < 0.0001). Histogram data shown in Figs. 4c,d and supplemental Fig. 6. b, Stability of mCherry-P2A-Cas13d in primary T cells compared between CLIP in the ACTB locus and randomly integrating lentiviruses driven by an EF1α or SFFV promoter (error bars represent SEM).
Extended Data Fig. 6 hyperdCas12a-miniVPR expression when transfected but not transduced into HEK293T cells.
a, A representative histogram of the hyperdCas12-miniVPR-T2A-GFP expression in HEK293T cells 2 days after transfection. b, The MFI of the hyperdCas12a-miniVPR+ cells. c, When the same vector is packaged into lentivirus and transduced into K562 cells, no expression is seen 2 days after transduction.
Extended Data Fig. 7 CLIP enables the expression and presentation of SARS-CoV-2 antigens.
a, Stability of S1-RdRP+ for both CLIP and EF1α driven lentivirus expression (error bars represent SEM). b, Histogram of sorted S1-RdRP+ Jurkat cells immediately before cryopreservation for immunopeptidomics.
Supplementary information
Supplementary Information
Supplementary figures, and nucleic acid and protein sequences.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chavez, M., Rane, D.A., Chen, X. et al. Stable expression of large transgenes via the knock-in of an integrase-deficient lentivirus. Nat. Biomed. Eng 7, 661–671 (2023). https://doi.org/10.1038/s41551-023-01037-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-023-01037-x
- Springer Nature Limited