Abstract
Unlike for DNA and RNA, accurate and high-throughput sequencing methods for proteins are lacking, hindering the utility of proteomics in applications where the sequences are unknown including variant calling, neoepitope identification, and metaproteomics. We introduce Spectralis, a de novo peptide sequencing method for tandem mass spectrometry. Spectralis leverages several innovations including a convolutional neural network layer connecting peaks in spectra spaced by amino acid masses, proposing fragment ion series classification as a pivotal task for de novo peptide sequencing, and a peptide-spectrum confidence score. On spectra for which database search provided a ground truth, Spectralis surpassed 40% sensitivity at 90% precision, nearly doubling state-of-the-art sensitivity. Application to unidentified spectra confirmed its superiority and showcased its applicability to variant calling. Altogether, these algorithmic innovations and the substantial sensitivity increase in the high-precision range constitute an important step toward broadly applicable peptide sequencing.
Similar content being viewed by others
Introduction
Liquid chromatography tandem mass spectrometry is the method of choice for identifying proteins at high throughput1. To this end, proteins are first digested into peptides whose mass-to-charge (m/z) ratios are determined in a first mass spectrum. Next, selected peptides are fragmented along their backbone bonds to generate series of peptide fragments whose m/z ratios can be identified in a second mass spectrum2. In principle, this spectrum allows the reconstruction of the peptide sequence by reading out the m/z differences between consecutive peaks of the same ion series3,4. In practice, this task is very hard due to missing peaks, contamination peaks, and because the ion series of the peaks are not known a priori. Peptide identification is greatly facilitated when the experimental spectrum is compared to expected spectra from a limited set of possible peptides, typically the in-silico digested proteome of an organism under study5. This strategy, which requires a precomputed database of possible peptides, is called database search6,7,8. The vast majority of proteomics studies rely on database search, even though, by design, database search does not allow the identification of novel or unexpected peptides. This prevents proteomics from being efficiently used in applications where the peptide sequences are not known a priori. This concerns neoepitope identification9, antibody sequencing10, pathogen surveillance11, microbial community studies12, and paleontology13. Therefore, efficient de novo peptide sequencing algorithms, which aim to identify peptides directly from spectra without relying on any database, are highly needed.
Most de novo peptide sequencing algorithms implement a combinatorial optimization approach in which the peptide that best fits the spectra is searched for. Various peptide-spectrum match (PSM) scores, i.e. scores that assess how well a candidate peptide corresponds to a given spectrum, combined with combinatorial optimization techniques including dynamic programming14,15,16,17,18 and genetic algorithms19,20 have been used to identify best-fitting peptides. Nevertheless, missing and contamination peaks have strongly limited the accuracy of those algorithms. Parallel to this work, we and others have leveraged deep learning to make major progress on the forward problem, i.e., predicting a spectrum given a peptide sequence21,22,23. While these algorithms do not predict contamination peaks, they can predict the peak intensities and missing peaks of a given peptide. Hence, their predictions can be leveraged to develop more discriminative PSM scoring functions for de novo peptide sequencing algorithms as in the algorithm pNovo324. Complementary to these algorithms based on combinatorial optimization, neural networks that directly predict the sequence of a peptide from the spectrum have recently been proposed. This includes DeepNovo33, of a candidate peptide to the correct peptide of a given spectrum. Here and elsewhere, we considered a candidate peptide to be correct if it exactly matched the peptide identified by MaxQuant at 1% FDR up to isoleucine-leucine substitution. We reasoned that the Levenshtein estimates would yield a quantitative notion of how far a peptide sequence is from the correct one in sequence space, making it a valuable loss function for combinatorial optimization algorithms.
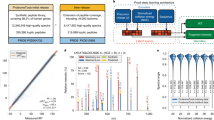
We trained a random forest to predict the Levenshtein distance of a candidate peptide to the correct peptide given 114 features including the number of predicted bin class changes by the bin reclassification model and the similarity between experimental and Prosit-predicted spectra (Methods, Supplementary Table 1). Overall, the method provided good estimates of the Levenshtein distance for peptides predicted by both Casanovo and Novor (Fig. 4a). No improved performance was achieved when fitting an XGBoost34 model instead (Supplementary Fig. 6). We, therefore, continued with the random forest. A notable deviation was seen for sequences with a Levenshtein distance of 1, which typically had a mass differing from the mass of the correct peptide by more than 20 ppm, possibly because our training dataset did not include these instances and because the computed spectral angles, Pearson and Spearman correlation coefficients were generally lower than the ones for sequences with close Levenshtein distances (Supplementary Fig. 7). Importantly, the predicted distance for the correct peptides was smaller than 1 for the vast majority of correct peptides, indicating that our scoring function is able to separate correct from incorrect peptide sequences. Consistent with this observation, the estimated Levenshtein distance was smaller for the correct peptide than for corresponding incorrect candidates on around 93% of the evaluated spectra for both Novor and Casanovo (Fig. 4b). The good discriminability indicated that our Levenshtein distance prediction could be used as a cost function in a combinatorial optimization-based de novo peptide sequencing algorithm.
a Estimated against actual Levenshtein distances of incorrect peptide identifications by Novor and Casanovo to the correct peptide sequence by MaxQuant across all 30 human samples. b Estimated Levenshtein distances of incorrect peptides against the estimated Levenshtein distances of the corresponding correct peptide sequences for a given spectrum across all 30 samples. The percentage of points above or on the diagonal line and below the diagonal line is labeled. c Precision-recall curves at peptide level before and after rescoring peptide identifications by Novor and Casanovo on the heart sample with Spectralis-score, our Levenshtein distance estimator, including the precision and recall for peptides from Casanovo with Novor substitutes for peptides with wrong mass (Casanovo-Novor). d Recall at 90% precision before and after rescoring peptides from Novor, Casanovo, as well as the combination of Casanovo and Novor sequences (Casanovo-Novor) across all n = 30 samples (different tissues). Statistical significance from a two-sided paired Wilcoxon test. The data in (a) and (d) are represented as boxplots in which the middle line indicates the median, the bounds of the box indicate the first and third quartiles and the whiskers indicate ±1.5 × IQR (interquartile range) from the third and first quartile, respectively. Source data are provided as a Source Data file.
We named our Levenshtein distance estimator Spectralis-score. Rescoring PSMs using Spectralis-score instead of original scores by Novor and Casanovo consistently improved recall at all precisions, notably at high-precision ranges. For instance, the peptide recall at 90% precision increased by 76% (from 0.25 to 0.44) when compared to the initial ranking by Casanovo on the heart sample (Fig. 4c). Notably, the relative performance improvements in recall at 90% precision were larger for longer peptides, reflecting the increase difficulties of de novo tools to perfectly sequence long peptides (Supplementary Fig. 8). Moreover, Spectralis-score outperformed the ranking given by the spectral angles between experimental and Prosit-predicted spectra, which is a feature of the model, or using PredFull-predicted35 spectra (Supplementary Fig. 9). Further integrating the spectral angles with PredFull-predicted spectra into a combined score did not lead to any improvement (Supplementary Fig. 9, Methods). This shows that the bin reclassification model provides complementary information to mere spectrum predictions. Since the mass of the peptides proposed by Casanovo did not match the precursor mass in approximately half of the spectra (44% in the heart sample), we replaced these peptides with Novor peptides, similarly to the suggestion by the authors of Casanovo, and denoted this combination of peptides Casanovo-Novor. We applied Spectralis-score to this combination and achieved higher recall at all precision ranges (Fig. 4c). A similar performance increase was observed for all other human samples (Fig. 4d, Supplementary Fig. 10).
The performance improvement of Spectralis-score held consistently after stratifying by precursor charge state and peptide length (Supplementary Figs. 11, 12). Furthermore, applying Spectralis-score to peptides proposed by the de novo sequencing tools DeepNovo50 allowed optimizing the number of filters per inner layer, the number of layers, the dropout rate, and the learning rate. The best model consisted of 16 AA-gapped convolution layers with 20 filters in each layer, a dropout rate of 0.3, and was trained with a learning rate of 4 × 10−5. It was trained using PyTorch (v1.8.1)47 on four A40 GPUs for 30 epochs, which resulted in a total training time of ~1.5 days.
Deep learning-based guided mutations
We implemented a graph-based algorithm that generates additional peptide sequences for a given input sequence using the predicted bin probabilities for b-ions and y-ions by the bin reclassification model. We constructed a graph by introducing a node for every m/z bin with a predicted probability larger than 0.35.
In order to deal with prefix fragments (b-ions) and suffix fragments (y-ions) in a unified fashion, we transformed m/z values of predicted b-ions to their complement to the precursor m/z. We defined the maximum over the two predicted probabilities for the b-ions and y-ions as node weights.
In addition, we added a source node with m/z value equal to 19 (i.e., the mass of one water molecule and one proton) and a target node with m/z bin equal the discretized experimental peptide mass derived from the precursor m/z and a proton. Source and target nodes received a node weight of one. Moreover, nodes for the m/z bins corresponding to y-ions of the input sequence were introduced to the graph with a node weight of 0.01.
We allowed an edge between two nodes if the difference between the m/z bins of the nodes corresponded to the discretized molecular mass of any amino acid. We labeled the edges with all amino acids that fulfilled the constraint.
To create additional peptide sequences, we performed weighted random walks starting from the source node until the target node was reached. To ensure that all random walks starting from the source node eventually led to the target node, we removed all nodes and edges that were not contained in any path from source to target. Edge probabilities for transition were defined based on the node weights. For any edge e = (v, w) with node weights pv and pw for the nodes v and w, we computed its edge probability pe as follows:
The peptide sequence can be recovered by concatenating all edge labels in the reversed path, thus starting from the target node. If more than one amino acid was labeled in an edge, one of them is selected at random.
Scoring procedure
Spectralis-score of a PSM was estimated as the Levenshtein distance33 of an input peptide sequence to the correct peptide sequence. The Levenshtein distance was computed with equal weights for insertions, deletions, and substitutions using the Python package editdistance (https://github.com/roy-ht/editdistance, v0.5.3). A random forest regressor served to predict the Levenshtein distance of a peptide sequence to its correct sequence.
We defined 114 features as input for the model derived from the comparison between Prosit-predicted and experimental spectra. To this end, we first applied base peak normalization to each experimental spectrum denoted as (Mexp, Iexp) consisting of m/z value and intensity pairs and to each Prosit-predicted spectrum (Mtheo, Itheo) consisting of k ≤ m/z value and intensity pairs. Intensity values below 0.02 were set to zero. Next, we defined experimental peaks, if any, corresponding theoretical peaks. The corresponding intensity \({\hat{I}}_{k}^{\exp }\) to a theoretical intensity \( {I}_{k}^{{{\mbox{theo}}}}\) of a peak k in the theoretical spectrum was defined as the intensity \({I}_{j}^{\exp }\) of its closest peak j within a mass tolerance δ = 20 ppm as follows:
We labeled these corresponding experimental peaks as b and y fragment ions according to the Prosit-predicted annotation.
We provided the model with three complementary feature types: similarity features, counting features, and features derived from the bin reclassification model.
The similarity features capture the quantitative agreement between Prosit-predicted and experimental peak intensities. They consist of the normalized spectral angle as defined earlier22, the Pearson correlation coefficient, cosine similarity, as well as the mean, standard deviation, quantiles, maximum, and minimum of the absolute differences.
The counting features capture the qualitative agreement between Prosit-predicted and experimental m/z ratios of peaks. They consist of the number (absolute and relative to the number of predicted peaks) of corresponding peaks for all four combinations of zero and nonzero experimental and theoretical intensities.
The similarity features and the counting features were generated for all peaks jointly, as well as for the b fragment ions and for the y fragment ions separately.
The features derived from the bin reclassification model were the amount of predicted bin class changes at various bin probability thresholds (0.25, 0.3, 0.35, 0.4, 0.45, and 0.5).
A list containing all features and computed feature importances obtained from the mean of the computed absolute SHAP values51 is provided in Supplementary Table 1.
The random forest predictor was trained to predict log2 (d + 1), where d is the Levenshtein distance of the peptide, minimizing the sum of squared errors using scikit-learn (v0.24.2)52. The final model, selected after hyper-parameter search with optuna50, contained 86 individual trees, a maximum tree depth of 175, a maximum number of 36 features for each node split, and a minimum amount of 112 samples per leaf node.
For comparison, an XGBoost (v1.6.2)34 model was fitted with the same target variable and same features as the random forest using scikit-learn. The final XGBoost model, selected after hyper-parameter search with optuna, consisted of 410 gradient-boosted trees, a maximum depth of 10, a ratio of 0.17 for subsampling features when constructing each tree, and a ratio of 0.92 for subsampling the training dataset. All other hyper-parameters were set to the default values.
The score integrating Spectralis-score and the spectral angle between the spectrum predicted by PredFull35 and the experimental spectrum was derived by fitting a logistic regression on these two scalars as features without any interaction term on the training dataset of the heart sample.
An alternative score was trained taking the defined 114 features as input, as well as the original scores provided by Casanovo v3.2.0 employing the leave-one-species-out cross-validation proposed by Tran et al.25. This score was obtained by fitting an XGBoost model to predict the Levenshtein distance of a candidate peptide to the correct peptide.
Evolutionary algorithm
For each experimental spectrum, candidate peptides provided by any de novo peptide sequencing tool served as input for the evolutionary algorithm. Here, for each experimental spectrum, we considered the candidate peptide provided by Casanovo as the initial sequence only if the difference between the computed peptide mass and the experimental mass derived from the precursor m/z was not larger than 1 Da. Otherwise, we started the optimization procedure with the candidate peptide generated by Novor. We denote this combination of sequences by Casanovo and Novor as Casanovo-Novor.
An initial population of candidate peptides was constructed by random isobaric substitutions and permutations of certain residues of the initial sequence: at most 3 consecutive residues were replaced by a combination of amino acids so that the total mass difference to the initial peptide sequence was not larger than 20 ppm.
At each generation, a set of n peptides is selected for the next generation based on the Spectralis-score s1,...,sn of candidate peptides in a current generation. The ne highest-scored peptides were directly inherited to the next generation. To maintain a fixed number of individuals in each generation, j: = n − ne candidate peptides were selected for mutation in the next generation. For this, we assigned a weight for selection wi to each peptide with index i and score si as follows:
where T denotes the temperature constant of the optimization procedure and s* the score of the fittest element in the current generation. After defining all selection weights, the selection procedure chose j peptides for mutation according to these weights. On each of those j peptides, we applied the guided mutation procedure.
Both selection and mutation procedures were repeated for m generations, each of them with the same size of n individuals and elite size of ne, before the most highly scored candidate peptide according to the selected fitness function was returned as the final peptide sequence of the given spectrum. Hyper-parameter grid search on a random subset of the validation set identified m = 5, n = 1,024 and ne = 103 as optimal hyper-parameters.
For initial peptide sequences of length larger than 30 or precursor charge larger than 6, we returned the initial sequence and the lowest possible score. Initial peptide sequences with an estimated Levenshtein distance smaller than 1 and larger than 7 were not optimized but returned unmodified as the final sequence of the evolutionary algorithm.
Peptide alignment and variant calling
Peptide alignments were obtained by running blastp (version 2.12.0+)53 against all translations from Ensembl genes and ab initio gene predictions provided by Ensembl human proteome database46,54 genome build GRCh38, release 83. As a scoring matrix, we used the identity matrix, modified such that leucine and isoleucine were considered equivalent. All other blastp settings were set to their default values, including the value of 10 for the e-value. We restricted the output of blastp to at most one hit per queried peptide sequence. If multiple hits were returned by the search, we selected the hit with the lowest e-value. We defined a query peptide to be a perfect alignment if the peptide is identical to the target peptide except for differences between leucine and isoleucine.
For each method, we computed the score cutoff for three precision values (80, 90, and 95%) as the median across the 30 samples of the score cutoffs yielding these precision values on spectra identified by MaxQuant.
To call missense variants from the selected RNA-seq sample (RNA-seq ID: SAMEA2154361, corresponding proteomics ID: heart_5a), we first aligned the RNA-seq reads using STAR (v2.7.10a)55 as part of the nf-core rnaseq56 module to the hg38 genome assembly using default parameters. We used GATK haplotypecaller through the RNA-seq variant calling module of the Detection of RNA Outliers Pipeline (DROP v1.2.2)57 to call variants. Variants that were missense were identified using VEP v.10654.
To map peptides to RNA-seq based missense variants, we ran a BLAST53,58 search using tblastn which provided nucleotide coordinates for each peptide. The tblastn results were processed in a similar matter described for the blastp results above. We overlapped the nucleotide coordinates with the obtained missense variants from VEP using GenomicRanges59.
Evaluation metrics
On top of precision-recall at bin level, we further evaluated the model for bin reclassification with change precision and change recall curves, which use change probabilities instead of the original probabilities predicted by the model. For every bin, the change probability was defined as the predicted probability p, for bins with an initial label equal to 1, and 1 − p for bins with an initial label equal to 0.
We evaluated the de novo peptide sequencing methods with precision-recall curves at peptide level computed on the set of spectra identified with MaxQuant at 1% FDR. Peptide-level recall was defined as the fraction of correct peptide sequences over the total number of peptide sequences identified with MaxQuant at 1% FDR. Note that unlike recall defined for binary classifiers, peptide-level recall is not guaranteed to reach one at the most lenient score cutoff.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometric raw data from the human dataset by Wang et al. including the MaxQuant Spectronaut search data is available via the PRIDE database with the dataset identifier PXD010154. RNA-Seq data is available in the following database: ArrayExpress E-MTAB-2836. The human proteome database (genome build GRCh38, release 83) was downloaded from Ensembl (https://ftp.ensembl.org/pub/release-83/fasta/homo_sapiens/pep). The raw mass spectrometric data for the nine-species dataset by Tran et al. is available via the PRIDE database with identifiers: PXD005025, PXD004948, PXD004325, PXD004565, PXD004536,PXD004947, PXD003868, PXD004467, and PXD004424. The correct peptide identifications, as well as predictions by DeepNovo, can be downloaded from the MassIVE repository with identifier MSV000081382. Model weights for running Casanovo were downloaded from Zenodo with DOI zenodo.679126360. The trained bin reclassification model and random forest, as well as Novor, Casanovo, DeepNovo, PointNovo and Spectralis predicted peptides with respective scores are deposited on Zenodo with DOI zenodo.839384661. The data to reproduce the main figures in this study have been deposited in the Figshare repository with DOI figshare.2353679462. Source data are provided with this paper as a Source Data file. Source data are provided with this paper.
Code availability
Source code and scripts are available on GitHub at https://github.com/gagneurlab/spectralis63.
References
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Zhang, Y., Fonslow, B. R., Shan, B., Baek, M.-C. & Yates, J. R. Protein analysis by shotgun/bottom-up proteomics. Chem. Rev. 113, 2343–2394 (2013).
Dančík, V., Addona, T. A., Clauser, K. R., Vath, J. E. & Pevzner, P. A. De novo peptide sequencing via tandem mass spectrometry. J. Comput. Biol. 6, 327–342 (1999).
Taylor, J. A. & Johnson, R. S. Sequence database searches via de novo peptide sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom 11, 1067–1075 (1997).
Muth, T. & Renard, B. Y. Evaluating de novo sequencing in proteomics: already an accurate alternative to database-driven peptide identification? Brief. Bioinform. 19, 954–970 (2018).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Sadygov, R. G., Cociorva, D. & Yates, J. R. Large-scale database searching using tandem mass spectra: Looking up the answer in the back of the book. Nat. Methods 1, 195–202 (2004).
Steen, H. & Mann, M. The abc’s (and xyz’s) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Karunratanakul, K., Tang, H.-Y., Speicher, D. W., Chuangsuwanich, E. & Sriswasdi, S. Uncovering thousands of new peptides with sequence-mask-search hybrid de novo peptide sequencing framework. Mol. Cell. Proteomics 18, 2478–2491 (2019).
Peng, W., Pronker, M. F. & Snijder, J. Mass spectrometry-based de novo sequencing of monoclonal antibodies using multiple proteases and a dual fragmentation scheme. J. Proteome Res. 20, 3559–3566 (2021).
Svetličić, E. et al. Direct identification of urinary tract pathogens by MALDI-TOF/TOF analysis and de novo peptide sequencing. Molecules 27, 5461 (2022).
Kleikamp, H. B. C. et al. Database-independent de novo metaproteomics of complex microbial communities. Cell Syst 12, 375–383.e5 (2021).
Cappellini, E. et al. Ancient Biomolecules and Evolutionary Inference. Annu. Rev. Biochem. 87, 1029–1060 (2018).
Chi, H. et al. pNovo: de novo peptide sequencing and identification using HCD spectra. J. Proteome Res. 9, 2713–2724 (2010).
Frank, A. & Pevzner, P. PepNovo: de novo peptide sequencing via probabilistic network modeling. Anal. Chem. 77, 964–973 (2005).
Ma, B. Novor: real-time peptide de novo sequencing software. J. Am. Soc. Mass Spectrom. 26, 1885–1894 (2015).
Ma, B. et al. PEAKS: powerful software for peptidede novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom 17, 2337–2342 (2003).
Fischer, B. et al. NovoHMM: A Hidden Markov Model for de Novo Peptide Sequencing. Anal. Chem. 77, 7265–7273 (2005).
Azari, S., Xue, B., Zhang, M. & Peng, L. GA-Novo: De Novo Peptide Sequencing via Tandem Mass Spectrometry Using Genetic Algorithm. in Applications of Evolutionary Computation (eds. Kaufmann, P. & Castillo, P. A.) vol. 11454, 72–89 (Springer International Publishing, 2019).
Heredia-Langner, A., Cannon, W. R., Jarman, K. D. & Jarman, K. H. Sequence optimization as an alternative to de novo analysis of tandem mass spectrometry data. Bioinformatics 20, 2296–2304 (2004).
Degroeve, S., Maddelein, D. & Martens, L. MS 2 PIP prediction server: compute and visualize MS2 peak intensity predictions for CID and HCD fragmentation. Nucleic Acids Res. 43, W326–W330 (2015).
Gessulat, S. et al. Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning. Nat. Methods 16, 509–518 (2019).
Zhou, X.-X. et al. pDeep: predicting MS/MS spectra of peptides with deep learning. Anal. Chem. 89, 12690–12697 (2017).
Yang, H., Chi, H., Zeng, W.-F., Zhou, W.-J. & He, S.-M. pNovo 3: precise de novo peptide sequencing using a learning-to-rank framework. Bioinforma. Oxf. Engl. 35, i183–i190 (2019).
Tran, N. H., Zhang, X., **n, L., Shan, B. & Li, M. De novo peptide sequencing by deep learning. Proc. Natl. Acad. Sci. USA 114, 8247–8252 (2017).
Qiao, R. et al. Computationally instrument-resolution-independent de novo peptide sequencing for high-resolution devices. Nat. Mach. Intell. 3, 420–425 (2021).
Yilmaz, M. et al. De novo mass spectrometry peptide sequencing with a transformer model. in Proc. 39th International Conference on Machine Learning (eds. Chaudhuri, K. et al.) vol. 162, 25514–25522 (PMLR, 2022).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Wang, D. et al. A deep proteome and transcriptome abundance atlas of 29 healthy human tissues. Mol. Syst. Biol. 15, e8503 (2019).
Cormican, J. A., Horokhovskyi, Y., Soh, W. T., Mishto, M. & Liepe, J. inSPIRE: an open-source tool for increased mass spectrometry identification rates using prosit spectral prediction. Mol. Cell. Proteomics 21, 100432 (2022).
Wilhelm, M. et al. Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics. Nat. Commun. 12, 3346 (2021).
Zolg, D. P. et al. INFERYS rescoring: Boosting peptide identifications and scoring confidence of database search results. Rapid Commun. Mass Spectrom. https://doi.org/10.1002/rcm.9128 (2021).
Levenshtein, V. I. Binary codes capable of correcting deletions, insertions and reversals. Sov. Phys. Dokl. 10, 707–710 (1966).
Chen, T. & Guestrin, C. XGBoost: A Scalable Tree Boosting System. in Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining https://doi.org/10.1145/2939672.2939785 (2016).
Liu, K., Li, S., Wang, L., Ye, Y. & Tang, H. Full-Spectrum Prediction of Peptides Tandem Mass Spectra using Deep Neural Network. Anal. Chem. 92, 4275–4283 (2020).
Gholamizoj, S. & Ma, B. SPEQ: quality assessment of peptide tandem mass spectra with deep learning. Bioinformatics 38, 1568–1574 (2022).
Ning, K., Fermin, D. & Nesvizhskii, A. I. Computational analysis of unassigned high-quality MS/MS spectra in proteomic data sets. Proteomics 10, 2712–2718 (2010).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Boonen, K. et al. Beyond genes: re-identifiability of proteomic data and its implications for personalized medicine. Genes 10, 682 (2019).
Mann, S. P., Treit, P. V., Geyer, P. E., Omenn, G. S. & Mann, M. Ethical principles, constraints, and opportunities in clinical proteomics. Mol. Cell. Proteomics 20, 100046 (2021).
Bandeira, N., Deutsch, E. W., Kohlbacher, O., Martens, L. & Vizcaíno, J. A. Data management of sensitive human proteomics data: current practices, recommendations, and perspectives for the future. Mol. Cell. Proteomics 20, 100071 (2021).
Yilmaz, M. et al. Sequence-to-sequence translation from mass spectra to peptides with a transformer model. https://doi.org/10.1101/2023.01.03.522621 (2023).
Dorfer, V., Maltsev, S., Winkler, S. & Mechtler, K. CharmeRT: boosting peptide identifications by chimeric spectra identification and retention time prediction. J. Proteome Res. 17, 2581–2589 (2018).
Driver, T. et al. Chimera spectrum diagnostics for peptides using two-dimensional partial covariance mass spectrometry. Molecules 26, 3728 (2021).
Houel, S. et al. Quantifying the impact of chimera MS/MS spectra on peptide identification in large-scale proteomics studies. J. Proteome Res. 9, 4152–4160 (2010).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res 50, D988–D995 (2022).
Paszke, A. et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. https://doi.org/10.48550/ARXIV.1912.01703 (2019).
Kingma, D. P. & Ba, J. Adam: A Method for Stochastic Optimization. https://doi.org/10.48550/ARXIV.1412.6980 (2014).
Lin, T.-Y., Goyal, P., Girshick, R., He, K. & Dollar, P. Focal Loss for Dense Object Detection. in 2017 IEEE International Conference on Computer Vision (ICCV) 2999–3007 (IEEE, 2017). https://doi.org/10.1109/ICCV.2017.324.
Akiba, T., Sano, S., Yanase, T., Ohta, T. & Koyama, M. Optuna: A Next-generation Hyperparameter Optimization Framework. in Proc. 25rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (2019).
Lundberg, S. M. & Lee, S.-I. A Unified Approach to Interpreting Model Predictions. in Advances in Neural Information Processing Systems (eds. Guyon, I. et al.) vol. 30 (Curran Associates, Inc., 2017).
Pedregosa, F. et al. Scikit-learn: Machine Learning. Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol 17, 122 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Yépez, V. A. et al. Clinical implementation of RNA sequencing for Mendelian disease diagnostics. Genome Med. 14, 38 (2022).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Lawrence, M. et al. Software for Computing and Annotating Genomic Ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Yilmaz, M. Casanovo data set and model weights. https://doi.org/10.5281/ZENODO.6791263 (2022).
Klaproth-Andrade, D. et al. Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing. https://doi.org/10.5281/ZENODO.8393846 (2022).
Klaproth-Andrade, D. et al. Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing. figshare. Dataset. https://doi.org/10.6084/m9.figshare.23536794.
andradesalazar. gagneurlab/spectralis: Spectralis v1.0.0. https://doi.org/10.5281/ZENODO.10204089 (2023).
Acknowledgements
We thank Ogüz Gültepe for his contributions on the initial work of this project, as well as Vicente Yépez for comments on the manuscript. We thank Felix Brechtmann for suggesting modeling the Levenshtein distance and for many fruitful discussions, and Stefan Dvoretskii for using an evolutionary algorithm. We thank Alexander Karollus for the helpful discussions and for coming up with the name of the method. We thank Florian Hölzlwimmer for his considerate and talented support with the GPU infrastructure. Furthermore, we thank Wassim Gabriel and Ludwig Lautenbacher for their assistance with the client to obtain Prosit predictions and scoring features. The IBM infrastructure hosting Prosit is operated and maintained by the UCC at the TUM. This work is supported by the Bundesministerium für Bildung und Forschung (BMBF) through the project CLINSPECT-M (FKZ031L0214A to D.K., J.H., and J.G.), the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) via the project NFDI 1/1 “GHGA - German Human Genome-Phenome Archive” (#441914366 to N.H.S.), and the European Union through the Horizon 2020 Program under Grant Agreement 823839 (H2020-INFRAIA-2018-1; EPIC-XS to M.W.). D.K., M.W. and J.G. were supported by a TUM Munich Data Science Institute (MDSI) seed fund.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
M.W. and J.G. jointly supervised the research. M.W. and J.G. conceived the method with the help of D.K. and J.H. D.K. and J.H. implemented the methods and performed the analysis on spectra identified by MaxQuant, with the help of Y.B. N.H.S. performed the peptide alignment and variant calling analysis with the help of Y.B. J.T. contributed to the method development. D.K., J.H., M.W., and J.G. wrote the manuscript with the help of N.H.S. and Y.B. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.W. is founder and shareholder of OmicScouts GmbH and MSAID GmbH, with no operational role in either company. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ekapol Chuangsuwanich, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Klaproth-Andrade, D., Hingerl, J., Bruns, Y. et al. Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing. Nat Commun 15, 151 (2024). https://doi.org/10.1038/s41467-023-44323-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-44323-7
- Springer Nature Limited