Abstract
Defects in insulin processing and granule maturation are linked to pancreatic beta-cell failure during type 2 diabetes (T2D). Phosphatidylinositol transfer protein alpha (PITPNA) stimulates activity of phosphatidylinositol (PtdIns) 4-OH kinase to produce sufficient PtdIns-4-phosphate (PtdIns-4-P) in the trans-Golgi network to promote insulin granule maturation. PITPNA in beta-cells of T2D human subjects is markedly reduced suggesting its depletion accompanies beta-cell dysfunction. Conditional deletion of Pitpna in the beta-cells of Ins-Cre, Pitpnaflox/flox mice leads to hyperglycemia resulting from decreasing glucose-stimulated insulin secretion (GSIS) and reducing pancreatic beta-cell mass. Furthermore, PITPNA silencing in human islets confirms its role in PtdIns-4-P synthesis and leads to impaired insulin granule maturation and docking, GSIS, and proinsulin processing with evidence of ER stress. Restoration of PITPNA in islets of T2D human subjects reverses these beta-cell defects and identify PITPNA as a critical target linked to beta-cell failure in T2D.
Similar content being viewed by others
Introduction
Type 2 diabetes (T2D) is a non-autoimmune disease of impaired insulin signaling that afflicts ~10% of the population in the United State alone1,2. Both impaired insulin release and reduced beta-cell mass contribute to the beta-cell failure that occurs during T2D3,4,5. Beta-cell failure is calculated to associate with a 24–65% loss of measurable beta-cell mass and a 50–97% loss of secretory capacity after disease onset6,7. Prior to the events that cause this decline, beta-cells functionally accommodate peripheral insulin resistance for a limited time in two ways. First, beta-cells increase insulin production2,4. Second, beta-cells increase their proliferation to expand the pool of insulin-producing cells in order to compensate for increased metabolic demand8,9,10. However, beta-cells in T2D patients ultimately succumb to multiple complications that include endoplasmic reticulum (ER) stress, glucotoxicity, and dedifferentiation11,12,13,14. The precise mechanisms underlying the decline of both beta-cell secretion and mass remain unclear. As a result, worldwide efforts continue to focus on identifying the molecular bases of these defects15,16,17.
Phosphoinositides define a set of chemically distinct phosphorylated derivatives of the glycerophospholipid phosphatidylinositol18. The central importance of phosphoinositide signaling in regulating cellular homeostasis in eukaryotes is demonstrated in two ways. First, the diversity of cellular activities regulated by phosphoinositide metabolism is striking. Phosphoinositide signaling controls cellular functions that range from membrane trafficking to receptor signaling at the plasma membrane, autophagy, transcription, mRNA transport, cytoskeleton dynamics, and numerous other activities19,20,21,22. Second, even subtle derangements in phosphoinositide metabolism contribute instrumentally to many diseases—including diabetes23,24,25. Phosphatidylinositol transfer proteins (PITPs) are highly conserved molecules that regulate the interface between lipid metabolism and cellular functions26,27. PITPs promote the activity of phosphatidylinositol (PtdIns) 4-OH kinases and PtdIns-4 phosphate (PtdIns-4-P) synthesis in eukaryotic cells28,29,37,38,39,40.
Here we first demonstrate that functional ablation of Pitpna in murine beta-cells results in random-fed hyperglycemia due to both impaired glucose-stimulated insulin secretion (GSIS) and reduced beta-cell number. These defects are accompanied by induction of ER stress and deranged mitochondrial dynamics and performance. Consistent with the murine studies, we further show that expression of PITPNA (referred to as human PITPNA and mouse Pitpna) is markedly diminished in pancreatic islets of T2D human subjects compared to non-diabetic donors. Such a downregulation is of functional consequence as reduced PITPNA levels in isolated human islets compromised PtdIns-4-P synthesis in the Golgi system, impaired insulin granule maturation and docking, and induced both ER and mitochondrial stress. Finally, we demonstrate that restoration of PITPNA expression in isolated pancreatic islets from T2D human subjects rescued insulin secretory capacity and granule biogenesis and alleviated ER stress. Taken together, these results establish that diminished PITPNA function is a major cell-autonomous contributor to reduced beta-cell mass and insulin output and, ultimately, to the beta-cell failure that represents a cardinal feature of T2D pathogenesis.
Results
Pitpna is a direct target of miR-375 in the pancreatic beta-cell
The microRNA miR-375 is a potent regulator of insulin secretion that directly targets expression of several genes including Myotrophin, Cadm1, Gephyrin (Gphn), and Elavl4/HuD41,42,43,44. An extended analysis using the TargetScan algorithm identified a candidate binding site for miR-375 in the 3′UTR of the gene Pitpna45. This gene encodes a phosphatidylinositol transfer protein and is expressed in throughout the pancreatic islet in humans and mice and is not restricted to beta-cells (Fig. 1a and Supplementary Fig. 1a). Immunostaining of human pancreas for PITPNA reveals its co-localization with markers of multiple organelles including the Golgi (Giantin), the ER (Calreticulin, CALR), and intracellular vesicles (CD63) (Fig. 1a). Similarly, in mouse pancreas, Pitpna co-localized with KDEL (ER) and GM130 (Golgi) (Supplementary Fig. 1a). To determine whether Pitpna is a genuine miR-375 target, the full-length mouse Pitpna 3′UTR (2709-nt, Pitpna WT) was subcloned into a luciferase reporter construct and the effects of modulating miR-375 levels on reporter expression were assessed. As expected, luciferase expression was inhibited in the presence of the miR-375 mimic (375-mimic) in comparison to cells transfected with a pool of scrambled control mimics (Ctrl-mimic) (Supplementary Fig. 1b). Moreover, direct targeting of this specific site binding site was supported by our result showing that site-directed mutagenesis of the putative binding site in the 3′UTR (Pitpna MUT) abolished the inhibitory effect of the miR-375 mimic (Supplementary Fig. 1b). To test whether endogenous Pitpna expression is subject to regulation by miR-375 in vivo, murine insulinoma MIN6 cells were transfected with an inhibitory antisense RNA oligonucleotide directed against miR-375 (Antg-375) to reduce expression of this miRNA (Supplementary Fig. 1c). The Antg-375-mediated silencing of miR-375 resulted in increased Pitpna mRNA levels when compared to cells transfected with a control pool of scrambled antisense oligonucleotides (Antg-Ctrl) (Supplementary Fig. 1c). Immunoblot analyses confirmed that inhibition of miR-375 resulted in elevated steady-state levels of Pitpna as well as other miR-375 targets (i.e., Cadm1, Gphn) (Supplementary Fig. 1d). Conversely, transfection with the 375-mimic reduced the steady-state levels of all three of these proteins in a dose-dependent manner (Supplementary Fig. 1e). Direct binding of miR-375 with its target genes is mediated by the RNA-binding protein Argonaute2 (Ago2)42. Consistent with the blocking miR-375 action, conditional deletion of Ago2 in pancreatic beta-cells (Ins-Cre, Ago2flox/flox) de-repressed Pitpna, Cadm1, and Gphn expression (Supplementary Fig. 1f).
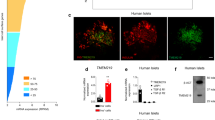
a Immunostaining of endogenous INSULIN (cyan), PITPNA (green), and protein markers (magenta) for the Golgi (GIANTIN), ER (CALR), and intracellular granules (CD63) in pancreatic islets isolated from a non-diabetic human subject. White arrows denote the colocalization of protein markers with PITPNA. b UMAP projection and graph-based clustering of scRNA-Seq analysis performed on isolated human pancreatic islet cell types. c Relative abundance of PITPNA in islet cell clusters from human donors. d Comparison of PITPNA expression in islet endocrine cell types from T2D (green) and non-diabetic donors (red). e Normalized PITPNA expression from bulk RNA sequencing of isolated human islets across non-diabetic (n = 51), pre-diabetic (n = 27), and diabetic human subjects (n = 11). Normalized expression values are shown in reads per million (RPM). PNon vs Pre-diabetes = 0.7248, PNon vs Diabetes = 0.025, PDiabetes vs Pre-diabetes = 0.1278, f Correlation analysis between normalized islet PITPNA expression and HbA1c of human subjects (n = 77). The R2 value indicates the correlation coefficient. g Correlation analysis between normalized islet PITPNA expression and body mass index (BMI) of human subjects (n = 89). The R2 value indicates the correlation coefficient. h qRT-PCR analysis of PITPNA mRNA expression in pancreatic islets isolated from non-diabetic (n = 15) and T2D (n = 5) human donors. P = 0.0006. Error bar: ND = 1.00 ± 0.02, T2D = 0.43 ± 0.04. i Western blot analysis of PITPNA and AGO2 expression in isolated islets of non-diabetic human donors (ND) and T2D donors (T2D). Data are presented as mean values ± SEM for (e), (h). *P < 0.05, ***P < 0.001. Ordinary one-way ANOVA with Turkey’s multiple comparisons test was used for (e). Linear regression was used for (f), (g). Two-tailed unpaired Student t-test were used for (h). All primary source data are reported in the Source data file.
Pitpna activity represents an interesting target for miR-375 control as it is an established mediator of PtdIns-4-P synthesis within the mammalian TGN51. Floxed Ago2 mice were generated and crossed with Ins-Cre mice from P. Herrera as described42. Numbers of animals are reported in each figure legends, and experiments were conducted in a blinded manner where the genotype is unknown during actual testing.
Gene expression analysis in mouse and human islets
Total RNA was extracted using the TRIzol reagent (Invitrogen). Quantitative real-time PCR (qRT-PCR) for miR-375 was quantified by TaqMan Assays using the TaqMan MicroRNA Reverse Transcription Kit and hsa-miR-375 primer sets (Thermo Fisher Scientific, 000564). MiR-375 levels were normalized to miR-U6 expression. For the expression of gene mRNAs, cDNA was synthesized using RevertAid First Strand cDNA synthesis kit (Fermentas), and qRT-PCR was measured using gene-specific primers with FastStart SYBR Green PCR Master Mix (Roche) on a StepOne Real-Time PCR System (ThermoFisher) (Supplementary Tables 2 and 5). Human islet expression data and accompanying donor information were previously published47 and are publicly accessible at Gene Expression Omnibus (GEO accession number GSE50398). Briefly, RNA-seq datasets were downloaded, trimmed (TrimGalore), and mapped to GRCh38 (HISAT2 mapper)90. Read counts for each sample were generated in SeqMonk software and normalized. The expression levels for PITPNA and INSULIN were correlated to the published clinical data included with the GEO submission. Single-cell RNA-seq data (GEO accession GSE85241)48 was downloaded from https://hemberg-lab.github.io/scRNA.seq.datasets/human/pancreas/ as a log normalized single cell experiment R object and processed using the R package Seurat v3.2.391.
Analytic procedures
Insulin measurements from plasma and pancreatic extracts were measured by ELISA (Crystal Chem), blood glucose and luciferase assays were measured as described44. Islet morphometric analysis was performed on 8 μm sections of paraffin-embedded pancreas approximately 150–200 μm apart. Sections were dewaxed, washed, and stained for insulin (Dako A0564, Agilent IR00261-2), glucagon (Millipore MABN238), Ki-67 (NovaCastra), Pitpna (Proteintech, 16613-1-AP), GM130 (BD, 610822), KDEL (Novus, NBP1-97469), Calreticulin (NOVUS, NBP1-47518), Giantin (NOVUS, AF8159), CD63 (Bio-Rad, MCA4754), or TUNEL (Roche cat. no. 11684795910). Cell numbers from all islets in 3–7 sections were counted with ImageJ software from 20X images obtained using a Nikon A1RSI Spectral Confocal Microscope. In vivo insulin release and glucose (GTT) tolerance tests were performed following a 16-h fast and intraperitoneally injection of glucose (2 g/kg BW). Insulin secretion from isolated islets was performed as described44. All antibodies and key reagents used in this study are listed in Supplementary Tables 4 and 5, respectively.
Cell culture, immunoprecipitation, and western blotting antibodies
Insulin secretion assay
MIN6 cells were cultured in DMEM (Invitrogen) containing 4.5 g/L glucose supplemented with 15% v/v heat-inactivated FCS, 50 μM β-mercaptoethanol, and 50 mg/mL penicillin and 100 mg/ml streptomycin and were a kind gift from Paolo Meda (University of Geneva, Switzerland). Cells were preincubated for 30 min in HEPES-balanced Krebs-Ringer bicarbonate buffer (KRH: 119 mM NaCl, 4.74 mM KCl, 2.54 mM CaCl2, 1.19 mM MgCl2, 1.19 mM KH2PO4, 25 mM NaHCO3, and 10 mM HEPES, pH 7.4) containing 0.5% BSA with 5 mM glucose and then were incubated for 2 h with various concentrations of glucose. Insulin release was measured by ELISA (Crystal Chem, cat. no 90080)41.
Western blotting
The following primary antibodies were used for Western blots at 1:1000 dilution: PITPNA (Abcam, ab180234), Cadm1 (MBL, CM004-3), Gephyrin (BD Biosciences, 610585), CHOP (Cell Signaling, 2895S), BiP/GRP78 (Cell Signaling, 3177S), DRP1 (Proteintech, 12957-1-AP), β-Actin (Cell Signaling, 3700S), and γ-Tubulin (Sigma, T6557). The following primary antibodies were used for immunofluorescence: PITPNA (1:200, Sigma, SAB1400211). Antibodies were used on paraffin-embedded pancreata fixed in 4% paraformaldehyde for 3 h. Image densitometry of 16-bit TIF images for all western blots was performed using ImageJ. All original uncropped western blot images are presented in Supplementary Figs. 7–13.
Transfection of microRNA mimics and related studies
MicroRNA mimics and siRNA pools were purchased from Qiagen GmbH (Germany) and scrambled pool controls are defined as an equimolar stock solution of either 48 random siRNA sequences, or 12 unique mimics of miRNAs not expressed in the beta-cell (i.e., miR-122, miR-1) and not predicted by the TargetScan algorithm to bind the 3′UTR of Pitpna. For biochemical fractionation, an eight-step sucrose gradient was performed on MIN6 cells as described previously43. Briefly, MIN6 cells were washed, pelleted, and resuspended in buffer containing 5 mM HEPES, 0.5 mM EGTA, and 1X Complete Protease inhibitors (Roche Applied Science) at pH 7.4 and homogenized. Homogenate was spun at 3000 × g for 10 min at 4 °C, and the post-nuclear supernatant was loaded onto an 8-step discontinuous sucrose density gradient (HEPES-buffered 0.2–2 M sucrose) and centrifuged at 242,000 × g for 2 h at 4 °C using an MLS50 rotor (Beckman Coulter). Extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) were measured in MIN6 cells using an XF24 Analyzer (Seahorse Bioscience, MA, USA).
Lentiviral-mediated over-expression and knockdown in isolated human islets
Lentiviruses were generated after subcloning the PITPNA cDNA sequence into the expression vector pCCL-cPPT-PGK-IRES-WPRE (Addgene). The resulting construct was transfected along with packaging plasmids pMD2.G and pSPAX2 (Addgene) into HEK293T cells (ATCC, CRL-3216). Cell culture media containing the virus was collected 48 and 72 h after transfection, concentrated, and stored at −80 °C. Knockdown of PITPNA by MISSION shRNA vectors (Sigma-Aldrich) was confirmed in human pancreatic 1.1B4 cells and isolated islets. Human islets were treated with non-overlap** shRNAs against the human PITPNA mRNA (accession number NM_006224), and TRCN00000299703 (SHCLNV 06302009MN) was used for all studies. TRC2 pLKO.5 Lentiviral Transduction Particles (pLKO.5-puro non-Mammalian shRNA Control Plasmid DNA; SHC00204V) were used to treat control human islets. Polybrene (Santa Cruz Biotechnology, Cat# sc-134220, Texas, USA) was added to the media with the final concentration of 10 μg/ml before infection. In brief, 250 islet equivalents (IEQ) seeded in each 12-well plate were infected with each lentivirus at an M.O.I of 20 for 48–72 h to ensure complete infection. All plasmids and shRNAs related to these procedures are listed in Supplementary Table 3.
Total internal reflection fluorescence (TIRF) microscopy
For TIRF microscopy experiments, human islets were obtained from the Nordic Network for Clinical Islet Transplantation, Uppsala Sweden, with ethical clearance (Uppsala Regional Ethics Board 2006/348) and the donor families’ written informed consent. Islets (donor IDs R442, 2583, 2585) were dispersed into single cells in cell dissociation buffer (Thermo Fisher Scientific) supplemented with trypsin (0.005%, Life Technologies), washed and plated in serum-containing medium on 22-mm polylysine-coated coverslips, allowed to settle overnight, and then transduced with adenovirus coding for the granule marker NPY-tdmOrange2. Lentiviral vectors expressed EGFP which was used to select cells for imaging as described previously68 using a lens-type total internal reflection (TIRF) microscope, based on an AxioObserver Z1 with a 100x/1.46 objective (Carl Zeiss). TIRF illumination with a decay constant of ~100 nm (calculated based on exit angle) was created using two DPSS lasers at 491 and 561 nm (Cobolt, Stockholm, Sweden) that passed through a cleanup filter (zet405/488/561/640x, Chroma) and was controlled with an acousto-optical tunable filter (AA-Opto, France). Excitation and emission light were separated using a beamsplitter (ZT405/488/561/640rpc, Chroma) and the emission light chromatically separated (QuadView, Roper) onto separate areas of an EMCCD camera (QuantEM 512SC, Roper) with a cutoff at 565 nm (565dcxr, Chroma) and emission filters (ET525/50 m and 600/50 m, Chroma). Scaling was 160 nm per pixel. Cells were imaged in (mM) 138 NaCl, 5.6 KCl, 1.2 MgCl2, 2.6 CaCl2, 0.2 diazoxide (to prevent spontaneous depolarizations), 10 D-glucose, 5 HEPES (pH 7.4 with NaOH) at ~35 °C. Exocytosis was evoked with high 75 mM K+ (equimolarly replacing Na+), applied by computer-timed local pressure ejection through a pulled glass capillary. Exocytosis events were identified manually based on the characteristic rapid loss of the granule marker fluorescence (1–2 frames).
Analysis of intracellular calcium
Intracellular calcium [iCa+2] assay was performed (Fluo-4NW Invitrogen, F36206, excitation 494 nm, emission 516 nm) according to the manufacturer’s instructions. Briefly, [iCa+2] was recorded for 60–90 s after addition of the KCl. Human islets were fixed in black 96-well optical bottom plates with poly-D-lysine coating. After the dye loading for an hour, the recording was done under confocal microscope (×40 objective) at room temp using an excitation filter of 488 nm. Fold change [iCa+2] was calculated from the baseline fluorescence recorded during the first 5 s before the addition of KCl. Images were captured at 1 s intervals for up to 60 s and the intracellular free calcium concentration is represented by mean fluorescence intensity.
Immunostaining and confocal microscopy
The following primary antibodies were used for immunofluorescence: anti-GOLPH3 (1:1000, Abcam, ab98023), anti-Golgin97 (1:100, Invitrogen, A-21270), anti-PtdIns-4-P (1:500, Echelon Biosciences cat. no. Z-P004), and anti-insulin (1:1000, Dako cat. no A0564). For immunostaining, both primary and secondary antibodies were diluted in 1x PBS containing 2.5% bovine serum albumin and 0.2% Triton-X-100. Antibody incubation steps (primary antibody: 3–4 h; secondary antibody: 1 h) were performed in a humidified chamber protected from direct light. Alexa Fluor 488, 594, 647 anti-rabbit, anti-mouse, or anti-guinea pig secondary antibodies used in this study are listed in Supplementary Table 4. Cell nuclei were stained with DAPI and mounted with Fluorsave reagent (MilliporeSigma, 345789) for fluorescence microscopy. Confocal images were acquired on a Nikon TiE confocal microscope using the NIS-Elements software with ×60 oil immersion objective. Images were imported into the Fiji version (http://fiji.sc) of the ImageJ software and the colocalization analyses were performed using the Coloc2 plugin (https://imagej.net/plugins/coloc-2)—an automated system that evaluates the fluorescent intensities of every pixel within an area of interest. Quantification of colocalization was performed using Pearson’s correlation coefficient. The Pearson’s correlation coefficient reflects the degree of linear relationship between two variables; in this case, the fluorescence intensities of two fluorescently tagged proteins GOLPH3 and Golgin97.
Transmission electron microscopy (TEM)
Isolated mouse islets and MIN6 cells were fixed in 2% paraformaldehyde/2.5% glutaraldehyde in 0.1 M Sodium Cacodylate buffer (cat. no. 15960-01 Electron Microscopy Sciences) for 2 h at 4 °C and then stained in 1.0% osmium tetroxide (cat. no. 19100 Electron Microscopy Sciences) for 1 h. After dehydrated in ethanol, cells were embedded with Spurr’s Low Viscosity Embedding Kit (cat. no. EM0300-1KT, Electron Microscopy Sciences), sectioned (70–90 nm thick), placed on Formvar (200 mesh) copper grids and contrasted with uranyl acetate (cat. no. 22409 Electron Microscopy Sciences) and lead citrate (cat. no. 22410 Electron Microscopy Sciences). Imaging was performed on a Philips Morgagni transmission electron microscope and acquired images were analyzed with respect to insulin granule and mitochondrial morphology.
Mass spectrometric analysis
Analysis of MIN6 cells
Cells were harvested by trypsinization and washed twice with ice-cold PBS. A modified protocol of Bligh & Dyer was used to extract lipids from cells92. Briefly, 900 μL of chloroform:methanol (1:2, v-v) (Thermo Fisher) was added to 2 × 106 cells. After vortexing for 1 minute and incubating for 15 minutes on ice, 300 μL of chloroform was added to the mixture, followed by mild vortexing and addition of 300 μL distilled water. The mixture was vortexed for 2 min and centrifuged at 20,000 × g for 2 min at 4 °C. The lipids were isolated from the lower organic phase. The sample was vacuum dried (Thermo Savant SPD SpeedVac) and the dried extract resuspended in 200 μL of chloroform:methanol (1:2, v/v) containing standards: PC 28:0, PE 28:0, PI 25:0, PG 28:0, PA 28:0, PS 28:0, LPC 17:0, LPE 14:0, d6-CE 18:0, and d5-TAG 48:0 (Avanti Polar Lipids). Phospholipids and neutral lipids were analyzed on an Agilent 1290 HPLC system coupled with an Agilent Triple Quadrupole mass spectrometer 6460, using Zorbax Eclipse Plus C18 column, 2.1 × 50 mm, 1.8 μm. The mobile phases were: A (acetonitrile:10 mM ammonium formate, 40:60) and B (acetonitrile:10 mM ammonium formate, 90:10). For phospholipids separation the gradient was as follows: start at 20% B, to 60% B in 2 min, to 100% B in 5 min, hold at 100% B for 2 min, back to 20% be in 0.01 min, hold 20% B 1.79 min (total runtime 10.8 min), the flow rate was 0.4 mL/min and the column temperature 30 °C. For neutral lipids separation the gradient was as follows: start at 20% B, to 75% B in 2 min, to 100% B in 4 min, hold at 100% B for 3 min, back to 20% be in 0.01 min, hold 20% B 1.79 min (total runtime 10.8 min), the flow rate was 0.4 mL/min and the column temperature 40 °C. Positive and negative electrospray ionization (ESI) was undertaken using the following parameters: gas temperature, 300 °C; gas flow, 5 l/min; nebulizer, 45 psi; sheath gas temperature, 250 °C; and sheath gas flow, 11 l/min; capillary voltage, 3.5 kV. Phospholipids and neutral lipids were measured using multiple reaction monitoring (MRM). Each biological replicate was measured twice and the average measurement used for analysis. MRM transitions areas were normalized to the areas of the internal standard from the same lipid class. Identification of peaks were based on retention time (RT) and specific MRM transitions for each lipid. Lipid species with CoV >25% were removed. Raw peak areas were integrated using Agilent MassHunter Quantitative Analysis software (version B.06.00), with further analysis performed in Microsoft Excel (v 16.0). Individual lipid species were quantified by comparison with spiked internal standards and lipid species with Signal-to-Noise <3 were removed. The molar fractions of individual lipid species and each lipid class were normalized to total lipids as follows: individual lipid intensities were divided by the relevant internal standard’s intensity and multiplied by the standard’s concentration; the obtained concentration value was divided by the sum of all lipids concentrations to yield molar fractions (mol%). Data were plotted using GraphPad Prism version 9.1.0. and R 4.1.0 (package ggplot2 and tidyr). Each biological replicate was measured twice and the average measurement was used for analysis. Comparisons between two groups were evaluated using an unpaired Student’s t test with Benjamini-Hochberg correction for mulitiple testing. Data summary for mass spectrometric analysis of MIN6 cells is present in the Source data file.
Analysis of human islets
Human pancreatic islets were washed with PBS and collected from 6 well plates then transfer into Eppendorf Lo-Bind polypropylene centrifuge tubes followed by centrifugation at 30,000 × g for 1 min at 4 °C. After removing the PBS, 0.5 M TCA was added to the pellet, vortexed, and incubated on ice for 10 min. The cooled mixture (TCA and islets) was centrifuged at 30,000 × g for 3 min at 4 °C and the supernatant discarded. Finally, 5% (w/v) TCA containing 10 mM EDTA was added to the pellet and vortexed, and then stored at −80 °C93,94. Internal standards of 20 ng PtdIns(4,5)P2, 20 ng PtdIns(4)P, and 100 ng PtdIns were added to the precipitates prepared from samples as described above. The lipid analytical internal standards were ammonium salts of, 1-heptadecanoyl-2-(5Z,8Z,11Z,14Z-eicosatetraenoyl)-sn-glycero-3-phospho-(1′-myo-inositol-4′,5′-bisphosphate) [17:0, 20:4 PtdIns(4,5)P2], 1-heptadecanoyl-2-(5Z,8Z,11Z,14Z-eicosatetraenoyl)-sn-glycero-3-phospho-(1′-myo-inositol-4′-phosphate) [17:0, 20:4 PtdIns(4)P], and 1-heptadecanoyl-2-(5Z,8Z,11Z,14Z-eicosatetraenoyl)-sn-glycero-3-phospho-(1′-myo-inositol) [17:0, 20:4 PI] from Avanti Polar Lipids (LIPID MAPS MS Standards)93. Following the addition of internal standards, 670 μl of ice-cold chloroform–methanol–12.1 N HCl (40:80:1) was added to each sample, after which the samples were vortexed for 2 min and allowed to sit on ice for another 10 min. Then, 650 μl of chloroform and 300 μl of 1 N HCl were added to generate two phases. Vortexed samples were centrifuged at 10,000 × g for 2 min to generate 2 separate phases. The lower phases were collected in a fresh 2 ml tube. An additional 950 μl of a mixture of chloroform, methanol, 1.74 M HCl (v:v:v) was added to the upper phase followed by vortexing and centrifugation at 10,000 × g for an additional 2 min. The resultant lower phase was combined with the previously collected lower phase and the combination dried under a nitrogen stream using a BiotageTM evaporator. The dried extracts were derivatized (methylated) with TMS-DM and resuspended in methanol for analysis94. Samples were resuspended in 25–100 µl 100% methanol (LC-MS Optima grade, Fisher) prior to chromatographic separation at ambient temperature using a C4 column (Waters Acquity UPLC Protein BEH C4, 1.7 µm 1.1 × 100; 300 A). A Waters Acquity FTN autosampler set at 4 °C injected 5 µl of sample via Waters Acquity UPLC. For chromatography of phosphoinositides the mobile phase was delivered over an 18.5 min runtime at a flow rate of 0.1 ml/min by a Waters Acquity UPLC. The gradient was initiated with 10 mM formic acid in water (A)/10 mM formic acid in acetonitrile (37:63 v/v) (B), held for 2 min, then increased to 15:85, v/v in 10 min, then increased to 100% B and held for 2.8 min followed by 3 min re-equilibration to starting conditions. The effluent was monitored by a Waters XEVO TQ-S MS/MS in multiple reaction monitoring mode (MRM) using electrospray and positive ion mode with post column infusion of 50 μM Na formate at 5 μL/min. to encourage the formation of sodiated adducts. Derivatized phosphoinositide species were quantified by targeted analysis. Specifically, peak areas for lipid species and standards were quantified by integrating curves using Waters’ MassLynxs software 4.2 employing TargetLynxs for peak integration and outputs. (Waters Corporation, Milford, MA, USA). For absolute calibrations and comparisons of different samples, peak areas of endogenous species were normalized to those of the corresponding internal standards. Two approximate assumptions were made: (1) the extraction and detection efficiency of phosphoinositides with each of the different fatty acyl combinations was the same as for the 37:4 standard, and (2) peak areas were proportional to lipid concentration. Data summary of mass spectrometric analysis of human islets is included in the Source data file.
Statistics and reproducibility
All results are expressed as mean ± standard error (SEM) and statistical analysis including all n numbers is summarized alongside primary data in the Source data file that accompanies this paper Results with human and mouse pancreatic islets and cell lines were all achieved with test groups of biologically independent samples of n ≥ 3 to determine reproducibility. Experiments with transgenic mice implemented test groups of each genotype of n ≥ 4 to determine reproducibility. All animal experiments implemented cohorts derived from multiple breeding pairs and were repeated at least once to determine reproducibility. A P-value of less than or equal to 0.05 was considered statistically significant. *P < 0.05, **P < 0.01, and ***P < 0.001. All graphical and statistical analyses were performed using the Prism8 software (Graphpad Software, USA) and Microsoft Excel. All measurements are from distinct samples and none of the results are derived from repeated measurement of the same sample. Comparisons between datasets with two groups were evaluated using an unpaired Student’s t test. ANOVA analysis was performed for comparisons of three or more groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All primary data including original uncropped western blots and raw mass spectrometry data generated in this study are provided in the Supplementary Information/Source data file provided with this paper. The following datasets containing expression analysis from human islet cells were re-analyzed for this study: GEO accession number GSE5039847 and GEO accession GSE8524148. Source data are provided with this paper.
References
Eizirik, D. L., Pasquali, L. & Cnop, M. Pancreatic beta-cells in type 1 and type 2 diabetes mellitus: different pathways to failure. Nat. Rev. Endocrinol. 16, 349–362 (2020).
Nolan, C. J. & Prentki, M. Insulin resistance and insulin hypersecretion in the metabolic syndrome and type 2 diabetes: Time for a conceptual framework shift. Diab. Vasc. Dis. Res. 16, 118–127 (2019).
Porksen, N. et al. Pulsatile insulin secretion: detection, regulation, and role in diabetes. Diabetes 51, S245–S254 (2002).
Campbell, J. E. & Newgard, C. B. Mechanisms controlling pancreatic islet cell function in insulin secretion. Nat. Rev. Mol. Cell Biol. 22, 142–158 (2021).
Rhodes, C. J. Type 2 diabetes-a matter of beta-cell life and death? Science 307, 380–384 (2005).
Chen, C., Cohrs, C. M., Stertmann, J., Bozsak, R. & Speier, S. Human beta cell mass and function in diabetes: Recent advances in knowledge and technologies to understand disease pathogenesis. Mol. Metab. 6, 943–957 (2017).
Liu, M. et al. Normal and defective pathways in biogenesis and maintenance of the insulin storage pool. J. Clin. Invest. 131, e142240 (2021).
Alejandro, E. U., Gregg, B., Blandino-Rosano, M., Cras-Meneur, C. & Bernal-Mizrachi, E. Natural history of beta-cell adaptation and failure in type 2 diabetes. Mol. Asp. Med. 42, 19–41 (2015).
Muoio, D. M. & Newgard, C. B. Mechanisms of disease: molecular and metabolic mechanisms of insulin resistance and beta-cell failure in type 2 diabetes. Nat. Rev. Mol. Cell Biol. 9, 193–205 (2008).
Ferrannini, E. The stunned beta cell: a brief history. Cell Metab. 11, 349–352 (2010).
Fonseca, S. G., Gromada, J. & Urano, F. Endoplasmic reticulum stress and pancreatic beta-cell death. Trends Endocrinol. Metab. 22, 266–274 (2011).
Shrestha, N., De Franco, E., Arvan, P. & Cnop, M. Pathological beta-cell endoplasmic reticulum stress in type 2 diabetes: current evidence. Front, Endocrinol. 12, 650158 (2021).
Back, S. H. & Kaufman, R. J. Endoplasmic reticulum stress and type 2 diabetes. Annu. Rev. Biochem. 81, 767–793 (2012).
Talchai, C., Lin, H. V., Kitamura, T. & Accili, D. Genetic and biochemical pathways of beta-cell failure in type 2 diabetes. Diabetes Obes. Metab. 11, 38–45 (2009).
van Raalte, D. H. & Verchere, C. B. Improving glycaemic control in type 2 diabetes: stimulate insulin secretion or provide beta-cell rest? Diabetes Obes. Metab. 19, 1205–1213 (2017).
Yong, J., Johnson, J. D., Arvan, P., Han, J. & Kaufman, R. J. Therapeutic opportunities for pancreatic beta-cell ER stress in diabetes mellitus. Nat. Rev. Endocrinol. 17, 455–467 (2021).
Rohm, T. V., Meier, D. T., Olefsky, J. M. & Donath, M. Y. Inflammation in obesity, diabetes, and related disorders. Immunity 55, 31–55 (2022).
Balla, T. Phosphoinositides: tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, 1019–1137 (2013).
Martin, T. F. Phosphoinositide lipids as signaling molecules: common themes for signal transduction, cytoskeletal regulation, and membrane trafficking. Annu. Rev. Cell Dev. Biol. 14, 231–264 (1998).
De Camilli, P., Emr, S. D., McPherson, P. S. & Novick, P. Phosphoinositides as regulators in membrane traffic. Science 271, 1533–1539 (1996).
Hokin, M. R. & Hokin, L. E. Enzyme secretion and the incorporation of P32 into phospholipides of pancreas slices. J. Biol. Chem. 203, 967–977 (1953).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Bridges, D. & Saltiel, A. R. Phosphoinositides in insulin action and diabetes. Curr. Top. Microbiol. Immunol. 362, 61–85 (2012).
Rameh, L. E. & Deeney, J. T. Phosphoinositide signalling in type 2 diabetes: a beta-cell perspective. Biochem. Soc. Trans. 44, 293–298 (2016).
Wuttke, A. Lipid signalling dynamics at the beta-cell plasma membrane. Basic Clin. Pharm. Toxicol. 116, 281–290 (2015).
Cleves, A. E. et al. Mutations in the CDP-choline pathway for phospholipid biosynthesis bypass the requirement for an essential phospholipid transfer protein. Cell 64, 789–800 (1991).
Bankaitis, V. A., Mousley, C. J. & Schaaf, G. The Sec14 superfamily and mechanisms for crosstalk between lipid metabolism and lipid signaling. Trends Biochem. Sci. 35, 150–160 (2010).
**e, Z., Hur, S. K., Zhao, L., Abrams, C. S. & Bankaitis, V. A. A Golgi lipid signaling pathway controls apical Golgi distribution and cell polarity during neurogenesis. Dev. Cell 44, 725–740.e724 (2018).
Lete, M. G., Tripathi, A., Chandran, V., Bankaitis, V. A. & McDermott, M. I. Lipid transfer proteins and instructive regulation of lipid kinase activities: Implications for inositol lipid signaling and disease. Adv. Biol. Regul. 78, 100740 (2020).
Ashlin, T. G., Blunsom, N. J. & Cockcroft, S. Courier service for phosphatidylinositol: PITPs deliver on demand. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1866, 158985 (2021).
Dickeson, S. K. et al. Isolation and sequence of cDNA clones encoding rat phosphatidylinositol transfer protein. J. Biol. Chem. 264, 16557–16564 (1989).
Tanaka, S. & Hosaka, K. Cloning of a cDNA encoding a second phosphatidylinositol transfer protein of rat brain by complementation of the yeast sec14 mutation. J. Biochem. 115, 981–984 (1994).
Fullwood, Y., dos Santos, M. & Hsuan, J. J. Cloning and characterization of a novel human phosphatidylinositol transfer protein, rdgBbeta. J. Biol. Chem. 274, 31553–31558 (1999).
Helmkamp, G. M. Jr., Harvey, M. S., Wirtz, K. W. & Van Deenen, L. L. Phospholipid exchange between membranes. Purification of bovine brain proteins that preferentially catalyze the transfer of phosphatidylinositol. J. Biol. Chem. 249, 6382–6389 (1974).
Wirtz, K. W. Phospholipid transfer proteins. Annu. Rev. Biochem. 60, 73–99 (1991).
Schaaf, G. et al. Functional anatomy of phospholipid binding and regulation of phosphoinositide homeostasis by proteins of the sec14 superfamily. Mol. Cell 29, 191–206 (2008).
Bankaitis, V. A., Aitken, J. R., Cleves, A. E. & Dowhan, W. An essential role for a phospholipid transfer protein in yeast Golgi function. Nature 347, 561–562 (1990).
Ohashi, M. et al. A role for phosphatidylinositol transfer protein in secretory vesicle formation. Nature 377, 544–547 (1995).
Hay, J. C. & Martin, T. F. Phosphatidylinositol transfer protein required for ATP-dependent priming of Ca(2+)-activated secretion. Nature 366, 572–575 (1993).
**e, Z. & Bankaitis, V. A. Phosphatidylinositol transfer protein/planar cell polarity axis regulates neocortical morphogenesis by supporting interkinetic nuclear migration. Cell Rep. 39, 110869 (2022).
Poy, M. N. et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432, 226–230 (2004).
Tattikota, S. G. et al. Argonaute2 mediates compensatory expansion of the pancreatic beta cell. Cell Metab. 19, 122–134 (2014).
Tattikota, S. G. et al. Argonaute2 regulates the pancreatic beta-cell secretome. Mol. Cell Proteom. 12, 1214–1225 (2013).
Poy, M. N. et al. miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proc. Natl Acad. Sci. USA 106, 5813–5818 (2009).
Agarwal, V., Subtelny, A. O., Thiru, P., Ulitsky, I. & Bartel, D. P. Predicting microRNA targeting efficacy in Drosophila. Genome Biol. 19, 152 (2018).
Cruz-Garcia, D. et al. Recruitment of arfaptins to the trans-Golgi network by PI(4)P and their involvement in cargo export. EMBO J. 32, 1717–1729 (2013).
Fadista, J. et al. Global genomic and transcriptomic analysis of human pancreatic islets reveals novel genes influencing glucose metabolism. Proc. Natl Acad. Sci. USA 111, 13924–13929 (2014).
Muraro, M. J. et al. A single-cell transcriptome atlas of the human pancreas. Cell Syst. 3, 385–394.e383 (2016).
**n, Y. et al. RNA sequencing of single human islet cells reveals Type 2 diabetes genes. Cell Metab. 24, 608–615 (2016).
Alb, J. G. Jr. et al. Mice lacking phosphatidylinositol transfer protein-alpha exhibit spinocerebellar degeneration, intestinal and hepatic steatosis, and hypoglycemia. J. Biol. Chem. 278, 33501–33518 (2003).
Thorens, B. et al. Ins1(Cre) knock-in mice for beta cell-specific gene recombination. Diabetologia 58, 558–565 (2015).
Harding, H. P. & Ron, D. Endoplasmic reticulum stress and the development of diabetes: a review. Diabetes 51, S455–S461 (2002).
Wright, J. et al. Endoplasmic reticulum oxidoreductin-1alpha (Ero1alpha) improves folding and secretion of mutant proinsulin and limits mutant proinsulin-induced endoplasmic reticulum stress. J. Biol. Chem. 288, 31010–31018 (2013).
Tabara, L. C., Morris, J. L. & Prudent, J. The complex dance of organelles during mitochondrial division. Trends Cell Biol. 31, 241–253 (2021).
Haythorne, E. et al. Diabetes causes marked inhibition of mitochondrial metabolism in pancreatic beta-cells. Nat. Commun. 10, 2474 (2019).
Smirnova, E., Griparic, L., Shurland, D. L. & van der Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Friedman, J. R. et al. ER tubules mark sites of mitochondrial division. Science 334, 358–362 (2011).
Hennings, T. G. et al. In vivo deletion of beta-cell Drp1 impairs insulin secretion without affecting islet oxygen consumption. Endocrinology 159, 3245–3256 (2018).
Arunagiri, A. et al. Misfolded proinsulin in the endoplasmic reticulum during development of beta cell failure in diabetes. Ann. N. Y Acad. Sci. 1418, 5–19 (2018).
Song, B., Scheuner, D., Ron, D., Pennathur, S. & Kaufman, R. J. Chop deletion reduces oxidative stress, improves beta cell function, and promotes cell survival in multiple mouse models of diabetes. J. Clin. Invest. 118, 3378–3389 (2008).
Marciniak, S. J. et al. CHOP induces death by promoting protein synthesis and oxidation in the stressed endoplasmic reticulum. Genes Dev. 18, 3066–3077 (2004).
Zinszner, H. et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev. 12, 982–995 (1998).
Grabon, A., Bankaitis, V. A. & McDermott, M. I. The interface between phosphatidylinositol transfer protein function and phosphoinositide signaling in higher eukaryotes. J. Lipid Res. 60, 242–268 (2019).
Bankaitis, V. A., Garcia-Mata, R. & Mousley, C. J. Golgi membrane dynamics and lipid metabolism. Curr. Biol. 22, R414–R424 (2012).
Kuna, R. S. & Field, S. J. GOLPH3: a Golgi phosphatidylinositol(4)phosphate effector that directs vesicle trafficking and drives cancer. J. Lipid Res. 60, 269–275 (2019).
Hammond, G. R. et al. PI4P and PI(4,5)P2 are essential but independent lipid determinants of membrane identity. Science 337, 727–730 (2012).
Stephens, L. R., Jackson, T. R. & Hawkins, P. T. Agonist-stimulated synthesis of phosphatidylinositol(3,4,5)-trisphosphate: a new intracellular signalling system? Biochim. Biophys. Acta 1179, 27–75 (1993).
Gandasi, N. R. et al. Glucose-dependent granule docking limits insulin secretion and is decreased in human Type 2 diabetes. Cell Metab. 27, 470–478.e474 (2018).
Alb, J. G. Jr., Gedvilaite, A., Cartee, R. T., Skinner, H. B. & Bankaitis, V. A. Mutant rat phosphatidylinositol/phosphatidylcholine transfer proteins specifically defective in phosphatidylinositol transfer: implications for the regulation of phospholipid transfer activity. Proc. Natl Acad. Sci. USA 92, 8826–8830 (1995).
Alb, J. G. Jr., Phillips, S. E., Wilfley, L. R., Philpot, B. D. & Bankaitis, V. A. The pathologies associated with functional titration of phosphatidylinositol transfer protein alpha activity in mice. J. Lipid Res. 48, 1857–1872 (2007).
Behnia, R. & Munro, S. Organelle identity and the signposts for membrane traffic. Nature 438, 597–604 (2005).
Nguyen, P. M. et al. The PI(4)P phosphatase Sac2 controls insulin granule docking and release. J. Cell Biol. 218, 3714–3729 (2019).
MacDonald, M. J., Ade, L., Ntambi, J. M., Ansari, I. U. & Stoker, S. W. Characterization of phospholipids in insulin secretory granules and mitochondria in pancreatic beta cells and their changes with glucose stimulation. J. Biol. Chem. 290, 11075–11092 (2015).
Omar-Hmeadi, M. & Idevall-Hagren, O. Insulin granule biogenesis and exocytosis. Cell Mol. Life Sci. 78, 1957–1970 (2021).
Arunagiri, A. et al. Proinsulin misfolding is an early event in the progression to type 2 diabetes. Elife 8, e44532 (2019).
Sun, J. et al. Proinsulin misfolding and endoplasmic reticulum stress during the development and progression of diabetes. Mol. Asp. Med. 42, 105–118 (2015).
Mousley, C. J. et al. Trans-Golgi network and endosome dynamics connect ceramide homeostasis with regulation of the unfolded protein response and TOR signaling in yeast. Mol. Biol. Cell 19, 4785–4803 (2008).
Nagashima, S. et al. Golgi-derived PI(4)P-containing vesicles drive late steps of mitochondrial division. Science 367, 1366–1371 (2020).
Youle, R. J. & van der Bliek, A. M. Mitochondrial fission, fusion, and stress. Science 337, 1062–1065 (2012).
Mishra, P. & Chan, D. C. Mitochondrial dynamics and inheritance during cell division, development and disease. Nat. Rev. Mol. Cell Biol. 15, 634–646 (2014).
Liesa, M. & Shirihai, O. S. Mitochondrial dynamics in the regulation of nutrient utilization and energy expenditure. Cell Metab. 17, 491–506 (2013).
Bock, F. J. & Tait, S. W. G. Mitochondria as multifaceted regulators of cell death. Nat. Rev. Mol. Cell Biol. 21, 85–100 (2020).
Sidarala, V. et al. Mitophagy protects beta cells from inflammatory damage in diabetes. JCI Insight 5, e141138 (2020).
Poy, M. N. MicroRNAs: an adaptive mechanism in the pancreatic beta-cell…and beyond? Best. Pr. Res Clin. Endocrinol. Metab. 30, 621–628 (2016).
LaPierre, M. P. & Stoffel, M. MicroRNAs as stress regulators in pancreatic beta cells and diabetes. Mol. Metab. 6, 1010–1023 (2017).
Kameswaran, V. et al. Epigenetic regulation of the DLK1-MEG3 microRNA cluster in human type 2 diabetic islets. Cell Metab. 19, 135–145 (2014).
Bolmeson, C. et al. Differences in islet-enriched miRNAs in healthy and glucose intolerant human subjects. Biochem. Biophys. Res Commun. 404, 16–22 (2011).
Locke, J. M., da Silva Xavier, G., Dawe, H. R., Rutter, G. A. & Harries, L. W. Increased expression of miR-187 in human islets from individuals with type 2 diabetes is associated with reduced glucose-stimulated insulin secretion. Diabetologia 57, 122–128 (2014).
Henquin, J. C., Ibrahim, M. M. & Rahier, J. Insulin, glucagon and somatostatin stores in the pancreas of subjects with type-2 diabetes and their lean and obese non-diabetic controls. Sci. Rep. 7, 11015 (2017).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
de la Cruz, L., Traynor-Kaplan, A., Vivas, O., Hille, B. & Jensen, J. B. Plasma membrane processes are differentially regulated by type I phosphatidylinositol phosphate 5-kinases and RASSF4. J. Cell Sci. 133, jcs233254 (2020).
Traynor-Kaplan, A. et al. Fatty-acyl chain profiles of cellular phosphoinositides. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1862, 513–522 (2017).
Acknowledgements
The authors thank David Castle, T. Osborne, L. Nagy, and M. Elena-Arango for assistance in the conduct of this work. This work was funded by Johns Hopkins All Children’s Hospital (M.N.P.), NIH grants R01 DK135688 (M.N.P.) and R35 GM131804 (V.A.B.), the Helmholtz Gemeinschaft (M.N.P.), two European Foundation for the Study of Diabetes EFSD/Lilly Programme Grants (M.N.P. and S.B.), Swedish Science Council (S.B.), NovoNordisk Foundation (S.B.), and the Deutsche Forschungsgemeinschaft (YA 721/3-1 to X.Y., LU 1455/6-1 to Y.L.). Human pancreatic islets and/or other resources were provided by the NIDDK-funded Integrated Islet Distribution Program (IIDP) (RRID:SCR _014387) at City of Hope, NIH Grant # U24DK098085 and the JDRF-funded IIDP Islet Award Initiative BS522P (to M.N.P.).
Author information
Authors and Affiliations
Contributions
Y.T.Y. and C.S. performed the primary expression analysis, animal husbandry, electron microscopy image analysis, immunohistochemical, and morphometric analysis and edited the manuscript. X.Y., Y.W., S.J., S.K., S.N., and A.A., performed expression analysis. L.L. and S.B. performed TIRF microscopy and edited the manuscript. Y.L., A.P., and M.M. performed immunohistochemical analysis. A.G. and F.v.M. reanalyzed public expression datasets. Y.W., A.C.-G., and M.W. performed and analyzed the lipidomic analysis. A.T.K. quantified phosphoinositides. P.A. edited the manuscript. Z.X. and V.A.B. developed and provided the Pitpna mutant animal lines and edited the manuscript. M.N.P. conceived and designed the study, wrote the manuscript, and is the guarantor of this work and takes responsibility for the integrity of the data and the accuracy of the data analysis. All authors contributed to interpretation of the data and approved the final version of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yeh, YT., Sona, C., Yan, X. et al. Restoration of PITPNA in Type 2 diabetic human islets reverses pancreatic beta-cell dysfunction. Nat Commun 14, 4250 (2023). https://doi.org/10.1038/s41467-023-39978-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-39978-1
- Springer Nature Limited