Abstract
Acute myeloid leukemia (AML) harboring inv(16)(p13q22) expresses high levels of miR-126. Here we show that the CBFB-MYH11 (CM) fusion gene upregulates miR-126 expression through aberrant miR-126 transcription and perturbed miR-126 biogenesis via the HDAC8/RAN-XPO5-RCC1 axis. Aberrant miR-126 upregulation promotes survival of leukemia-initiating progenitors and is critical for initiating and maintaining CM-driven AML. We show that miR-126 enhances MYC activity through the SPRED1/PLK2-ERK-MYC axis. Notably, genetic deletion of miR-126 significantly reduces AML rate and extends survival in CM knock-in mice. Therapeutic depletion of miR-126 with an anti-miR-126 (miRisten) inhibits AML cell survival, reduces leukemia burden and leukemia stem cell (LSC) activity in inv(16) AML murine and xenograft models. The combination of miRisten with chemotherapy further enhances the anti-leukemia and anti-LSC activity. Overall, this study provides molecular insights for the mechanism and impact of miR-126 dysregulation in leukemogenesis and highlights the potential of miR-126 depletion as a therapeutic approach for inv(16) AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is an aggressive hematopoietic malignancy characterized by excessive proliferation of immature leukemic blasts. Approximately 21,000 patients are diagnosed with AML each year in the United States, and the latest 5-year overall survival rate remains at only 28% (https://seer.cancer.gov)1,2. AML is maintained and propagated by a population of leukemia-initiating cells or leukemia stem cells (LSCs), which feature quiescence and therapy resistance, and contribute to subsequent clonal evolution of the disease and relapse3,4.
AML comprises multiple entities characterized by specific gene mutations and chromosomal abnormalities that drive leukemogenesis and can be used as prognosticators5,6. One of the most frequent chromosomal translocations detected in 5–12% AML patients is chromosome 16 inversion, inv(16)(p13q22) or t(16;16)(p13.1;q22) [henceforth inv(16)], which is associated with the FAB M4Eo AML subtype7,8. At the molecular level, inv(16) disrupts the CBFB gene encoding the CBFβ subunit of the core-binding factor (CBF) transcription factor complex that acts as a master regulator of hematopoietic development and lineage specification9,10. Specifically, inv(16) breaks and joins CBFB and the smooth-muscle myosin heavy chain (MYH11) gene, creating a fusion gene CBFB-MYH11 (CM) which encodes a fusion protein CBFβ-SMMHC11. Although inv(16) AML patients have a relatively favorable prognosis, only approximately 50–60% eventually achieve long-term survival with standard chemotherapy12,13. Therefore, the development of therapies capable of targeting LSCs is necessary to achieve a cure in the vast majority of the patients.
MicroRNAs (miRNAs) are small non-coding RNA molecules that modulate multiple targets by promoting mRNA degradation or repressing translation14,15. MiRNAs are transcribed as primary (pri)-miRNAs, processed into precursor (pre)-miRNAs by DROSHA, exported into the cytoplasm via the RAN (a small GTP-binding RAS-related nuclear protein)-Exportin-5 (XPO5) complex16,17, and processed by Dicer into mature miRNAs18. In hematopoiesis, miRNAs play critical roles in coordinating context-dependent differentiation programs and cell fate decisions19. MiR-126 is highly expressed in endothelial cells and plays a key role in angiogenesis, vascular development and homeostasis20,21. Within the hematopoietic compartment, miR-126 expression is enriched in long-term hematopoietic stem cells (HSCs) and plays a pivotal role in restraining cell cycle progression and maintaining HSC quiescence22. Aberrant miRNA expression profiles have been associated with malignant transformation and prognosis in leukemia and other hematologic malignancies23,24,25,26,27. Notably, miR-126 is aberrantly expressed in CBF leukemias including t(8;21) and inv(16)25,26. We and others have reported that miR-126 contributes to LSC activity, maintenance, and drug resistance in myeloid leukemias28,29,30. Furthermore, we recently reported that activating tyrosine kinase mutations (e.g., BCR-ABL, FLT3-ITD) deregulate intronic miRNA (e.g., miR-126) biogenesis by interfering with RAN-XPO5 mediated pre-miRNA processing via phosphorylation of SPRED1, a member of the Sprouty family of proteins30,31.
Here, we investigated the mechanism and functional impact of miR-126 dysregulation in inv(16) AML. We utilized the conditional Cbfb-MYH11 knock-in mouse model32,33 combined with a miR-126 floxed allele34 to determine the function and regulatory mechanism of miR-126 during inv(16)-induced AML development and evaluated the efficacy of targeting LSCs using miRisten, an anti-miR-126 oligonucleotide therapeutic.
Results
Cbfb-MYH11 (CM) upregulates miR-126 expression in hematopoietic stem and progenitor cells (HSPCs)
Previous studies have shown that miR-126 is highly expressed and contributes to the relative quiescence of normal CD34+ HSCs22,28,29,30. Herein we show that CD34+ cells from inv(16) AML patients have even higher miR-126 levels than CD34+ cells from normal healthy donors (Fig. 1a). MiR-126 is located within intron 7 of the EGFL7 gene35, which was highly co-expressed with miR-126 (Fig. 1b).
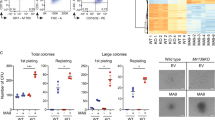
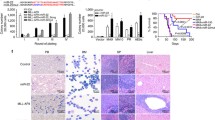
a Relative levels of miR-126 in CD34+ and CD34− cells from healthy (HL; black circle; n = 6; HL CD34+ vs. HL CD34− p = 0.0036) donors and inv(16) AML patients (red square; n = 10; inv(16) CD34+ vs. HL CD34+ p = 0.024; inv(16) CD34− vs. HL CD34− p = 0.0053; inv(16) CD34+ vs. inv(16) CD34− p = 0.0002). b Relative levels of EGFL7 in CD34+ and CD34− cells from HL donors (black circle; n = 6; HL CD34+ vs. HL CD34− p = 0.016) and inv(16) AML patients (red square; n = 10; inv(16) CD34+ vs. HL CD34+ p = 0.006; inv(16) CD34+ vs. inv(16) CD34− p < 0.0001). c Counts per million (CPM) reads for miR-126-3p in PB based on miRNA-seq of control (Ctrl; black line; n = 7) and CM (red line; n = 6) mice over time until moribund with leukemia (two-way ANOVA analysis showed CM vs. Ctrl p < 0.0001). d Normalized reads per kilobase per million of transcript (RPKM) for Egfl7 in PB based on RNA-seq of Ctrl (black line; n = 7) and CM (red line; n = 6) mice over time (two-way ANOVA analysis showed Ctrl vs. CM p = 0.0003). e Relative levels of miR-126 in CM (red square) preleukemic (6 weeks after induction; left; n = 6) or leukemic (right; n = 6) vs. Ctrl (black circle; n = 6) HSPC populations, including LSK (preleukemic p = 0.0074; leukemic p = 0.049), GMP (preleukemic p = 0.0064; leukemic p = 0.043), Pre-Meg/E (preleukemic p = 0.00015; leukemic p = 0.0012), Pre-GM (preleukemic p < 0.0001; leukemic p = 0.0002), and EP (leukemic p < 0.0001). f Western blot for CBFβ and CM using anti-CBFβ in 32D cells transduced with CBFB vs. CM (left). Relative levels of primary pri-miR-126, precursor pre-miR-126, mature miR-126, and Egfl7 in 32D cells expressing CM (red dots/bars) vs. CBFβ (black dots, gray bars). g Western blot analysis for CM in 32D-CM non-silencing (NS) control vs. CM shRNA (A3, D4) cells (left). Relative levels of mature miR-126, pri-miR-126, pre-miR-126 and Egfl7 expression in NS (red), CM shRNA-A3 (blue) or CM shRNA-D4 (gray) cells (right). h Relative Gata2 expression in BM samples of control (black/gray; n = 5) vs. CM leukemia (red; n = 4; p = 0.0002) mice. i Relative level of Gata2 in leukemia BM cells transduced with NS control (red), Gata2 shRNA #2 (dark blue), Gata2 shRNA #4 (light blue). j Relative expression of miR-126 in leukemia BM cells transduced with NS control (red), Gata2 shRNA #2 (dark blue), Gata2 shRNA #4 (light blue). k EGFL7 promoter-firefly luciferase reporter activity normalized with renilla luciferase as internal control in 293T cells co-transfected with or without GATA2, RUNX1, CBFβ or CM expression vectors as indicated (gray: none; green: GATA2; light blue: RUNX1 + CBFβ; dark blue: RUNX1 + CBFβ + GATA2; orange: RUNX1 + CM; Red: RUNX1 + CM + GATA2). Each dot represents results from an individual sample and data are presented as the mean ± SEM in a, b, e, h; each line represents the trajectory of an individual mouse in c, d. Representative data of at least three independent experiments are shown in f, g, i, j and data are presented as the mean ± SD. Q-PCR data in a, b, e, f for pri-miR-126, pre-miR-126, Egfl7, and Gata2 were normalized to Beta-2-microglobulin (B2M) and miR-126 data were normalized to RNU44 (human) or snoRNA234 (mouse). The statistical significance for all comparisons shown was determined using two-tailed Student’s T tests (*p < 0.05; **p < 0.01; ***p < 0.001).
We previously generated a conditional CM knock-in (Cbfb56M/+/Mx1-Cre) mouse model32,33, which recapitulates human inv(16) AML development in an average of 3–4 months after induction of CM expression by poly(I:C) treatment. In a time-sequential analysis of miR-126 levels in mononuclear cells from peripheral blood (PB) of CM knock-in mice, we observed a progressive increase of both miR-126 and Egfl7 levels over time (Fig. 1c, d) and in multiple CM HSPC populations33,36,37 (Fig. 1e). Significant upregulation of miR-126 was detected as early as 1 month after CM induction (Supplementary Fig. 1a).
CM upregulates Egfl7/miR-126 transcription in concert with Gata2
To determine whether CM directly dysregulates miR-126 expression, we forced CM or wild-type CBFβ expression in 32D myeloblast cells via MSCV-IRES-GFP (MIG) retroviral transduction. We observed upregulated miR-126, pri-miR-126 and pre-miR-126 as well as Egfl7 in CM-transduced GFP+ cells compared to CBFB-transduced GFP+ cells (Fig. 1f). Conversely, shRNA-mediated CM knockdown resulted in significant downregulation of miR-126, pri-miR126, pre-miR-126 and Egfl7 in 32D-CM cells (Fig. 1g). We and others have shown that CM induces upregulation of the Gata2 transcription factor33,38, which reportedly transactivates the EGFL7/miR-126 promoter39,40. Accordingly, we observed increased Gata2 levels upon CM transduction in 32D cells (Supplementary Fig. 1b), as well as in CM knock-in PB (Supplementary Fig. 1c) along with miR-126/Egfl7 levels (Fig. 1c, d), and AML cells from CM knock-in mice (Fig. 1h). Conversely, Gata2 knockdown in CM knock-in AML cells resulted in downregulation of miR-126 (Fig. 1I, j) and Egfl7 (Supplementary Fig. 1d). Analysis of the promoter regions of both murine and human EGFL7 identified several putative CBF binding sites (TRANSFAC database41) nearby the GATA-binding sites (Supplementary Fig. 1e). Chromatin immunoprecipitation (ChIP)-qPCR showed that both CBFβ and CM protein indeed could occupy these CBF sites within the Egfl7 promoter region and that some of the nearby GATA sites were also occupied by GATA2 (Supplementary Fig. 1e). In addition, we confirmed RUNX1, CBFβ/CM occupancy on the reported RUNX1 site42,43 as well as GATA2 binding to the reported GATA2 site42,43 in the human EGFL7 promoter by ChIP-qPCR using primary inv(16) AML CD34+ cells (Supplementary Fig. 1f). We then performed luciferase reporter assays to evaluate transactivation of the human EGFL7 promoter44 by GATA2 and the CBF complex, which is consisted of RUNX1/CBFβ or RUNX1/CM. A significant increase of EGFL7-luciferase transactivation was observed with forced expression of GATA2, RUNX1/CBFβ and RUNX1/CM (Fig. 1k). Co-transfection of RUNX1/CBFβ or RUNX1/CM with GATA2 further increased EGFL7 promoter transactivation, suggesting a co-regulatory activity of GATA2 and RUNX1/CBF on EGFL7/miR-126 transcription.
CM increases biogenesis of mature miR-126 through the HDAC8/RAN–XPO5–RCC1 axis
Next, we tested whether additional mechanisms other than transcriptional regulation of EGFL7/miR-126 were active in inducing high levels of mature miR-126 in CM cells. To this end, we previously reported and confirmed that CM interacts and enhances the activity of a class I histone deacetylase (HDAC), HDAC845 (Supplementary Fig. 2a). Notably, HDAC8 overexpression (OE) in 32D cells significantly increased mature miR-126 levels while reducing the pre-miR-126 precursor levels compared to vector alone (Supplementary Fig. 2b). Conversely, Hdac8 knockdown or treatment with an HDAC8-selective inhibitor (HDAC8i; 22d) significantly downregulated the levels of mature miR-126 and increased those of pre-miR-126 (Fig. 2a, b). These results led us to postulate a possible regulatory role of CM in enhancing the miR-126 biogenesis via HDAC8.
a Western blot analysis of HDAC8 in 32D-CM cells with NS or Hdac8 shRNA (#1, #2) (left). Relative expression of pre-miR-126 and mature miR-126 in 32D-CM cells with NS (red) vs. Hdac8 shRNA#1 (light blue), Hdac8 shRNA#2 (dark blue) (right). b Relative expression in 32D-CM cells treated with vehicle (red) vs. HDAC8i (22d 2.5 μM or 5 μM; purple) for pre-miR-126 (left; 2.5 μM p < 0.0001; 5 μM p < 0.0001) and miR-126 (right; 2.5 μM p = 0.012; 5 μM p = 0.0055). c Representative image of IF co-staining of RAN (green) and HDAC8 (red) in 32D cells (left; scale bar 10 μm). IP with anti-IgG control or anti-HDAC8 followed by immunoblotting (IB) with anti-RAN antibodies (right). Representative results of two independent experiments with similar results are shown. d IP with anti-RAN and IB with antibodies for Ac-Lys, RCC1, XPO5, or RAN in 32D-CM cells not-treated (NT) or treated with DMSO vehicle or 22d (2 µM) (left). The levels of RAN-GTP determined by RAN activation assay (right). The densitometry of RAN-GTP levels measured from three assays are shown on the bottom. e IP with anti-RAN and IB with antibodies for Ac-Lys, RCC1, XPO5, or RAN (left), and RAN activation assay (right) in 32D-CM cells not-treated (NT) or transduced with shCtrl control or shCM for 24 h. f IP with anti-RAN and IB with antibodies for Ac-Lys, RCC1, XPO5, or RAN (left), and RAN activation assay (right) of BM cells isolated from Ctrl or CM mice. g Schematic model of the mechanism by which CM regulates miR-126/EGFL7 transcription as well as miR-126 biogenesis through enhancing HDAC8 activity which in turn promotes RAN-XPO5 mediated transportation. Pre-miR-126 was normalized to B2m and mature miR-126 was normalized to snoRNA234. Data are presented as the mean ± SD and statistical significance shown was determined using two-tailed Student’s T tests (*p < 0.05; **p < 0.01; ***p < 0.001).
First, we tested the impact of CM on HDAC8 protein stability. We found that CM expression enhanced the half-life of HDAC8 protein following cycloheximide treatment (Supplementary Fig. 2c). HDAC8 protein is subjected to MDM2-mediated ubiquitination (Supplementary Fig. 2d), which was reduced by CM expression (Supplementary Fig. 2e, left) and increased by CM knockdown (Supplementary Fig. 2e, right). ErbB3-binding protein 1 (EBP1) reportedly interacts with another HDAC family member HDAC2Small RNA library preparation and next-generation sequencing (NGS) All libraries were prepared using the Illumina TruSeq Small RNA protocol with minor modification following the manufacturer’s instructions. Briefly, 280 ng of pooled total RNA was ligated to the sRNA 3′ adaptor (TCT GGA ATT CTC GGG TGC CAA GGA ACT CC) with T4 RNA Ligase 2, truncated (New England BioLabs) for 1 h at 22 °C, and subsequently ligated to a 5′ adaptor (0.5 μL of 5 μM per reaction): GUUCAGAGUUCUACAGUCCGACGAUCNNN with T4 RNA ligase1 (New England BioLabs) for 1 h at 20 °C. The constructed smRNA library was first reverse-transcribed using GX1 (5′-GGAGTTCCTTGGCACCCGAGA) as the RT primer and then subjected to PCR amplification for 13 cycles, using primers GX1 (5′-CAAGCAGAAGACGGCATACGAGAT[NNNNNN]GTGACTGGAGTTCCTTGGCACCCGAGAATTCCA) and GX2 (5′-AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCT CTTCCGATCT) then followed by 6% TBE PAGE gel purification with size selection (for targeted small RNAs of 17–35 nt). The purified library was quantified using qPCR with a forward primer (5′-CAAGCAGAAGACGGCATACG) and a reverse primer (5′-AATGATACGGCGACCACCGA). Individual libraries were prepared using a unique index primer in order to allow for pooling of multiple samples prior to sequencing. The library was quantified using qPCR. Sequencing of 50 cycles was performed on a HiSeq 2500 (Illumina Inc., San Diego, CA), and image processing and base calling were conducted using Illumina’s pipeline. Raw small RNA sequences were trimmed to remove the 3′-adapter (TCTGGAATTCTCGGGTGCCAAGGAACTCC) using cutadapt v0.9.3. Reads longer than 16 bp after trimming were aligned to mouse genome assembly mm9 using Bowtie v 0.12.7 with default settings. The expression level of mouse mature miRNAs from Sanger mirBase v18 were counted as previously described68. The counts of miRNAs were then normalized by the trimmed mean of M-values (TMM) method and counts per million (CPM) values were calculated by Bioconductor package “edgeR” v3.4.2. Total RNA was extracted using the AllPrep DNA/RNA/miRNA Universal Kit (Qiagen, 80224); quality and quantity were estimated using BioAnalyser Systems (Agilent Technologies). Samples with a RIN >8.0 were included. External RNA Controls Consortium (ERCC) Spike-In Control Mix (Thermo Fisher Scientific, 4456740) was added to all samples per the manufacturer’s recommendations, although these were not used for downstream analyses. Sequencing libraries were constructed using the KAPA RNA HyperPrep Kit with RiboErase (HMR) (Roche, KK8560), loaded on to a cBot system for cluster generation, and sequenced on a Hiseq 2500 System (Illumina) with single end 51-bp for mRNA-seq to a nominal depth of 50 million reads. Raw RNA-seq sequences were subjected to adapter trimming using Trimmomatic v0.38 and poly(A) tails were removed using FASTP v0.19.4. The trimmed reads were aligned to mouse genome mm10 using Tophat v2.0.8 with default settings. Expression level of RefSeq gene (downloaded on 02/13/2020) were counted using HTSeq count v0.6.1. The raw count data were normalized by the TMM method using Bioconductor package “edgeR” v3.20.9. Differential expression analysis was carried out using the quasi-likelihood (QL) F-test implemented in “edgeR” to determine the differentially expressed genes, with the cutoff of average RPKM in one group ≥1, p values ≤0.01 and fold change ≥1.5. The gene set enrichment analysis (GSEA69) was performed by ClusterProfiler v3.10.1 to identify the affected GO, hallmark and KEGG pathways from MSigDB v770, using the pre-ranked gene list sorted by the −log10(p value) with a sign determined by the fold change direction. Comparison between groups was performed by a two-tailed, paired or unpaired Student’s T-test, one-way or two-way ANOVA for normal distributions. The log-rank test was used to assess significant differences between survival curves. All statistical analyses were performed using Prism version 8.0 software (GraphPad Software). Sample sizes chosen are indicated in the individual figure legends and were not based on formal power calculations to detect prespecified effect sizes. A p value less than 0.05 was considered statistically significant. All experiments are repeated independently with similar results at least two times. Further information on research design is available in the Nature Research Reporting Summary linked to this article.miRNA-seq data analysis
RNA-seq and bioinformatics
Statistics and reproducibility
Reporting summary
Data availability
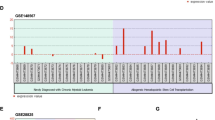
The raw data and processed RNA-seq data for LSK cells generated in this study have been deposited in the GEO repository under accession number GSE184015, which can be assessed with no restriction. The raw data and processed RNA-seq data associated with Fig. 1D, S1C was deposited under GEO accession number GSE133642, which can be assessed with no restriction. Raw miRNA-seq data associated with Fig. 1C were deposited to GEO under accession number GSE173785. The uncropped blots generated in this study are provided in the Supplementary Information/Source data file. Source data are provided with this paper.
References
Bloomfield, C. D., Marcucci, G., Dohner, K. & Dohner, H. Acute myeloid leukemia. Introduction. Semin. Oncol. 35, 324–325 (2008).
Hourigan, C. S. Acute myeloid leukemia: Introduction. Semin. Hematol. 52, 149 (2015).
Thomas, D. & Majeti, R. Biology and relevance of human acute myeloid leukemia stem cells. Blood 129, 1577–1585 (2017).
Pollyea, D. A. & Jordan, C. T. Therapeutic targeting of acute myeloid leukemia stem cells. Blood 129, 1627–1635 (2017).
Dohner, H. et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood 129, 424–447 (2017).
Khaled, S., Al Malki, M. & Marcucci, G. Acute myeloid leukemia: biologic, prognostic, and therapeutic insights. Oncology (Williston Park) 30, 318–329 (2016).
Liu, P. P. et al. Identification of the chimeric protein product of the CBFB-MYH11 fusion gene in inv(16) leukemia cells. Genes Chromosomes Cancer 16, 77–87 (1996).
Mrózek, K., Heerema, N. A. & Bloomfield, C. D. Cytogenetics in acute leukemia. Blood Rev. 18, 115–136 (2004).
de Bruijn, M. F. & Speck, N. A. Core-binding factors in hematopoiesis and immune function. Oncogene 23, 4238–4248 (2004).
Speck, N. A. & Gilliland, D. G. Core-binding factors in haematopoiesis and leukaemia. Nat. Rev. Cancer 2, 502–513 (2002).
Liu, P. et al. Fusion between transcription factor CBF beta/PEBP2 beta and a myosin heavy chain in acute myeloid leukemia. Science 261, 1041–1044 (1993).
Prebet, T. et al. Acute myeloid leukemia with translocation (8;21) or inversion (16) in elderly patients treated with conventional chemotherapy: a collaborative study of the French CBF-AML intergroup. J. Clin. Oncol. 27, 4747–4753 (2009).
Ustun, C. et al. Core-binding factor acute myeloid leukemia with inv(16): older age and high white blood cell count are risk factors for treatment failure. Int. J. Lab. Hematol. 43 e19–e25 (2020).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Bohnsack, M. T., Czaplinski, K. & Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J. E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Chen, C. Z., Li, L., Lodish, H. F. & Bartel, D. P. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004).
Wang, S. et al. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev. Cell 15, 261–271 (2008).
Fish, J. E. et al. miR-126 regulates angiogenic signaling and vascular integrity. Dev. Cell 15, 272–284 (2008).
Lechman, E. R. et al. Attenuation of miR-126 activity expands HSC in vivo without exhaustion. Cell Stem Cell 11, 799–811 (2012).
Li, Z. et al. Distinct microRNA expression profiles in acute myeloid leukemia with common translocations. Proc. Natl Acad. Sci. USA 105, 15535–15540 (2008).
Jongen-Lavrencic, M., Sun, S. M., Dijkstra, M. K., Valk, P. J. & Lowenberg, B. MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia. Blood 111, 5078–5085 (2008).
Garzon, R. et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 111, 3183–3189 (2008).
Cammarata, G. et al. Differential expression of specific microRNA and their targets in acute myeloid leukemia. Am. J. Hematol. 85, 331–339 (2010).
Marcucci, G., Radmacher, M. D., Mrozek, K. & Bloomfield, C. D. MicroRNA expression in acute myeloid leukemia. Curr. Hematol. Malig. Rep. 4, 83–88 (2009).
de Leeuw, D. C. et al. Attenuation of microRNA-126 expression that drives CD34+38- stem/progenitor cells in acute myeloid leukemia leads to tumor eradication. Cancer Res 74, 2094–2105 (2014).
Lechman, E. R. et al. miR-126 regulates distinct self-renewal outcomes in normal and malignant hematopoietic stem cells. Cancer Cell 29, 602–606 (2016).
Zhang, B. et al. Bone marrow niche trafficking of miR-126 controls the self-renewal of leukemia stem cells in chronic myelogenous leukemia. Nat. Med. 24, 450–462 (2018).
Nguyen, L. X. T. et al. Cytoplasmic DROSHA and non-canonical mechanisms of miR-155 biogenesis in FLT3-ITD acute myeloid leukemia. Leukemia 35, 2285–2298 (2021).
Kuo, Y. H. et al. Cbf beta-SMMHC induces distinct abnormal myeloid progenitors able to develop acute myeloid leukemia. Cancer Cell 9, 57–68 (2006).
Cai, Q. et al. CBFbeta-SMMHC creates aberrant megakaryocyte-erythroid progenitors prone to leukemia initiation in mice. Blood 128, 1503–1515 (2016).
Kuhnert, F. et al. Attribution of vascular phenotypes of the murine Egfl7 locus to the microRNA miR-126. Development 135, 3989–3993 (2008).
Parker, L. H. et al. The endothelial-cell-derived secreted factor Egfl7 regulates vascular tube formation. Nature 428, 754–758 (2004).
Hua, W. K. et al. HDAC8 regulates long-term hematopoietic stem-cell maintenance under stress by modulating p53 activity. Blood 130, 2619–2630 (2017).
Jeannet, R., Cai, Q., Liu, H., Vu, H. & Kuo, Y.-H. Alcam Regulates long-term hematopoietic stem cell engraftment and self-renewal. Stem Cells 31, 560–571 (2013).
Saida, S. et al. Gata2 deficiency delays leukemogenesis while contributing to aggressive leukemia phenotype in Cbfb-MYH11 knockin mice. Leukemia 34, 759–770 (2020).
Hartmann, D. et al. MicroRNA-based therapy of GATA2-deficient vascular disease. Circulation 134, 1973–1990 (2016).
Le Bras, A. et al. VE-statin/egfl7 expression in endothelial cells is regulated by a distal enhancer and a proximal promoter under the direct control of Erg and GATA-2. PLoS ONE 5, e12156 (2010).
Wingender, E. The TRANSFAC project as an example of framework technology that supports the analysis of genomic regulation. Brief. Bioinformatics 9, 326–332 (2008).
Zhang, X. O., Gingeras, T. R. & Weng, Z. Genome-wide analysis of polymerase III-transcribed Alu elements suggests cell-type-specific enhancer function. Genome Res. 29, 1402–1414 (2019).
Zhang, J. et al. An integrative ENCODE resource for cancer genomics. Nat. Commun. 11, 3696 (2020).
Harris, T. A., Yamakuchi, M., Kondo, M., Oettgen, P. & Lowenstein, C. J. Ets-1 and Ets-2 regulate the expression of microRNA-126 in endothelial cells. Arterioscler. Thromb. Vasc. Biol. 30, 1990–1997 (2010).
Qi, J. et al. HDAC8 inhibition specifically targets Inv(16) acute myeloid leukemic stem cells by restoring p53 acetylation. Cell Stem Cell 17, 597–610 (2015).
Zhang, Y., Woodford, N., **a, X. & Hamburger, A. W. Repression of E2F1-mediated transcription by the ErbB3 binding protein Ebp1 involves histone deacetylases. Nucleic Acids Res. 31, 2168–2177 (2003).
Nguyen, D. Q. et al. The role of ErbB3 binding protein 1 in cancer: friend or foe? J. Cell Physiol. 233, 9110–9120 (2018).
Kim, V. N. MicroRNA precursors in motion: exportin-5 mediates their nuclear export. Trends Cell Biol. 14, 156–159 (2004).
de Boor, S. et al. Small GTP-binding protein Ran is regulated by posttranslational lysine acetylation. Proc. Natl Acad. Sci. USA 112, E3679–E3688 (2015).
Kuo, Y. H., Gerstein, R. M. & Castilla, L. H. Cbfbeta-SMMHC impairs differentiation of common lymphoid progenitors and reveals an essential role for RUNX in early B-cell development. Blood 111, 1543–1551 (2008).
Liu, L. Y. et al. Mir-126 inhibits growth of SGC-7901 cells by synergistically targeting the oncogenes PI3KR2 and Crk, and the tumor suppressor PLK2. Int J. Oncol. 45, 1257–1265 (2014).
Wakioka, T. et al. Spred is a Sprouty-related suppressor of Ras signalling. Nature 412, 647–651 (2001).
Lee, K. J. et al. Requirement for Plk2 in orchestrated ras and rap signaling, homeostatic structural plasticity, and memory. Neuron 69, 957–973 (2011).
Sears, R. et al. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 14, 2501–2514 (2000).
Coller, H. A., Forman, J. J. & Legesse-Miller, A. “Myc’ed messages”: myc induces transcription of E2F1 while inhibiting its translation via a microRNA polycistron. PLoS Genet. 3, e146 (2007).
Hermeking, H. et al. Identification of CDK4 as a target of c-MYC. Proc. Natl Acad. Sci. USA 97, 2229–2234 (2000).
Li, Z., Boone, D. & Hann, S. R. Nucleophosmin interacts directly with c-Myc and controls c-Myc-induced hyperproliferation and transformation. Proc. Natl Acad. Sci. USA 105, 18794–18799 (2008).
Fernandez, P. C. et al. Genomic targets of the human c-Myc protein. Genes Dev. 17, 1115–1129 (2003).
Pulikkan, J. A. et al. CBFbeta-SMMHC inhibition triggers apoptosis by disrupting MYC chromatin dynamics in acute myeloid leukemia. Cell 174, 1325 (2018).
Li, Z. et al. Overexpression and knockout of miR-126 both promote leukemogenesis. Blood 126, 2005–2015 (2015).
Mandoli, A. et al. CBFB-MYH11/RUNX1 together with a compendium of hematopoietic regulators, chromatin modifiers and basal transcription factors occupies self-renewal genes in inv(16) acute myeloid leukemia. Leukemia 28, 770–778 (2014).
Zhen, T. et al. RUNX1 and CBFbeta-SMMHC transactivate target genes together in abnormal myeloid progenitors for leukemia development. Blood 136, 2373–2385 (2020).
Nucera, S. et al. miRNA-126 orchestrates an oncogenic program in B cell precursor acute lymphoblastic leukemia. Cancer Cell 29, 905–921 (2016).
Castilla, L. H. et al. The fusion gene Cbfb-MYH11 blocks myeloid differentiation and predisposes mice to acute myelomonocytic leukaemia. Nat. Genet. 23, 144–146 (1999).
Zhao, L. et al. CBFB-MYH11 hinders early T-cell development and induces massive cell death in the thymus. Blood 109, 3432–3440 (2007).
Kuo, Y. H. et al. Runx2 induces acute myeloid leukemia in cooperation with Cbfbeta-SMMHC in mice. Blood 113, 3323–3332 (2009).
Kühn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Weng, L. et al. MicroRNA profiling of clear cell renal cell carcinoma by whole-genome small RNA deep sequencing of paired frozen and formalin-fixed, paraffin-embedded tissue specimens. J. Pathol. 222, 41–51 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. in Stem Cell Transcriptional Networks (ed. Kidder, B. L.) (Springer, 2014).
Acknowledgements
This work was supported in part by the National Institutes of Health under award number R01CA205247 (to Y.-H.K. and G.M.), U01CA250046 (to Y.-H.K., G.M. and R.C.R.), R01CA213131 (to M.K.), and the Gehr Family Center for Leukemia Research. Research reported in this publication included work performed in the Analytical Cytometry Core and Animal Resource Center supported by the National Cancer Institute of the National Institutes of Health under award number P30CA33572. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
L.Z. and L.X.T.N. designed and performed research, analyzed data, and wrote the manuscript; Y.-C.C., D.W., G.J.C., D.H.H, C.J.B., X.H., H.D., S.L., M.L., D.Z., J.Q.,W.-K.H., Q.C., E.C., W.C. and X.W. performed research, analyzed data, and reviewed manuscript; P.S. synthesized the CpG-anti-miR-126 and reviewed the manuscript; R.C.R., M.K., L.L, B.Z. and G.M. designed research and reviewed manuscript; Y.-H.K. designed research, analyzed and interpreted data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Joost Martens, Takahiro Ochiya and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, L., Nguyen, L.X.T., Chen, YC. et al. Targeting miR-126 in inv(16) acute myeloid leukemia inhibits leukemia development and leukemia stem cell maintenance. Nat Commun 12, 6154 (2021). https://doi.org/10.1038/s41467-021-26420-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-021-26420-7
- Springer Nature Limited
This article is cited by
-
OST-01, a natural product from Baccharis coridifolia, targets c-Myc-dependent ribogenesis in acute myeloid leukemia
Leukemia (2024)
-
Leukemogenesis occurs in a microenvironment enriched by extracellular microvesicles/exosomes: recent discoveries and questions to be answered
Leukemia (2024)
-
Metabolic dependencies of acute myeloid leukemia stem cells
International Journal of Hematology (2024)
-
Identifying key genes and functionally enriched pathways in acute myeloid leukemia by weighted gene co-expression network analysis
Journal of Applied Genetics (2024)
-
Targeting PRMT9-mediated arginine methylation suppresses cancer stem cell maintenance and elicits cGAS-mediated anticancer immunity
Nature Cancer (2024)