Abstract
Esophageal squamous cell carcinoma (ESCC) is prevalent in some geographical regions of the world. ESCC development presents a multistep pathogenic process from inflammation to invasive cancer; however, what is critical in these processes and how they evolve is largely unknown, obstructing early diagnosis and effective treatment. Here, we create a mouse model mimicking human ESCC development and construct a single-cell ESCC developmental atlas. We identify a set of key transitional signatures associated with oncogenic evolution of epithelial cells and depict the landmark dynamic tumorigenic trajectories. An early downregulation of CD8+ response against the initial tissue damage accompanied by the transition of immune response from type 1 to type 3 results in accumulation and activation of macrophages and neutrophils, which may create a chronic inflammatory environment that promotes carcinogen-transformed epithelial cell survival and proliferation. These findings shed light on how ESCC is initiated and developed.
Similar content being viewed by others
Introduction
Esophageal squamous cell carcinoma (ESCC) is a common gastrointestinal malignancy in some parts of the world like China, with a poor prognosis and high mortality mainly due to the lack of specific measures for early diagnosis and effective therapies. ESCC represents an extraordinary paradigm of carcinoma development, shaped in a sequential manner from inflammation (INF), hyperplasia (HYP), dysplasia (DYS), carcinoma in situ (CIS) to invasive carcinoma (ICA)1. However, up to date, how ESCC initiates and develops is largely unknown. This long-standing question is the major factor obstructing the early intervention and clinical care improvement of the disease. As such, exploring the mechanisms underlying ESCC formation and identifying biomarkers are the crucial tasks for early detection, diagnosis, and precision treatment of the cancer.
Recent genome studies, including The Cancer Genome Atlas (TCGA) project, have identified many genome variations in ESCC by using whole-exome or whole-genome sequencing on clinical tissue samples2,3. Although these studies have revealed an important role of the identified genome alterations in ESCC, it remains unresolved that how the normal epithelial cells may transit by the mutations through precancerous lesions to invasive carcinoma because all these previous studies were in cross-sectional design. Another important issue in this regard is that somatic mutations solely might not be sufficient for ESCC initiation and development because such mutations also occur in pathologically normal human esophageal tissues4. These findings imply that other mechanisms such as transcriptome aberrance in ESCC tumorigenesis may merit further investigation. In addition, it has been demonstrated that the complex context of tumor microenvironment (TME) also has important roles in tumor initiation and development. Therefore, elucidating dynamic transcriptomic changes of TME cellular compositions during tumorigenesis are significant and inevitable in discovering how ESCC develops. Recently established single-cell transcriptomic analysis is a promising approach, which allows to analyze the complex cellular compositions and to decipher cell state transition in tissue samples5.
Capturing continuous tumorigenic lesions from sole patient over time to perform such study is impossible and, therefore, to address this important issue would be highly dependent on well-established animal models. Fortunately, it has been shown that chemical carcinogen 4-nitroquinoline 1-oxide (4NQO) can induce mouse ESCC development in a manner that mimics the tumorigenic processes of ESCC in humans6,7. The distinct multiple stages of ESCC tumorigenesis induced by the carcinogen provide an excellent opportunity to interrogate cell state transition by single-cell RNA sequencing (scRNA-seq), which would elucidate the dynamic ESCC tumorigenesis.
Here we report a single cell-based transcriptomic profiling study on various types of cells across every pathogenic stage during ESCC initiation and development in a mouse model induced by 4NQO. We have built a complete atlas and characterized the transcriptomic profiles for the transition of esophageal epithelial cells under the attack of carcinogen and elucidated how they evolve over time in a holistic approach. We have also depicted the transition landscapes of non-epithelial cells, i.e., fibroblasts and immune cells, in the esophageal microenvironments of different stages of tumorigenesis. Furthermore, we have found that some key changes in mice also occur in human esophageal tissue samples. These results shed light on the phenotypes and transition fates of different cell types across the tumorigenic processes of ESCC in animal model and may be implicated in human ESCC.
Results
Bulk RNA-seq and scRNA-seq of mouse esophageal samples
To explore the transcriptomic alterations at various pathological stages during ESCC tumorigenesis, 8-week-old female C57BL/6 mice were treated with 4NQO for 16 weeks, which resulted in five recognizable precancerous and cancerous lesions in the esophagus, i.e., INF, HYP, DYS, CIS, and ICA (Fig. 1a; Supplementary Fig. 1a). We examined mice receiving 4NQO and found they all developed the expected lesions in the esophagus at different time points of experiment. The ESCC number per animal (mean ± SD) at week 26 was 6.0 ± 3.6. We first performed conventional RNA-seq on mini-bulks of normal epithelial samples obtained by laser-capture microdissection (LCM) from control mice at different ages of 1, 2, 8, and 25 months and various precancerous or cancerous lesions from 4NQO-treated mice. Principal component analysis (PCA) of the differentially expressed genes showed that the expression programs in mice exposed to 4NQO were substantially different from that in control mice; however, there existed some overlaps in the expression profiles between 4NQO-exposed and non-exposed mice. For instance, the transcriptomic profile for stage INF in 4NQO-exposed mice was similar to that in control mice aged 25 months (Fig. 1b), indicating that conventional RNA-seq could not precisely clarify the path of malignant cell transition during the development and progression of ESCC due to high intra-tissue heterogeneity.
a Induction of esophageal precancerous and cancerous lesions in mice. Mice were treated with 4NQO in drinking water (100 μg/ml) for 16 weeks and then kept without 4NQO treatment for another 10 weeks (upper panel). Mice were killed before (week 0), during (week 12) and after treatment (weeks 20, 22, 24, or 26), respectively. Hematoxylin–eosin (H&E) staining and immunohistochemistry (IHC) analysis of Mki67 on esophageal epithelium slides clearly identified six different pathological lesions, i.e., normal (NOR), inflammation (INF), hyperplasia (HYP), dysplasia (DYS), carcinoma in situ (CIS), and invasive carcinoma (ICA) (lower panel). Similar staining results were observed in over three visual fields from each stage of esophageal lesions (more staining image in Supplementary Fig. 2d). Scale bar, 100 μm. b Plot of principal component analysis (PCA) of mini-bulk tissue RNA-seq on different pathological lesions indicated by different colors. M month of mouse age. c Overview of the experimental design of scRNA-seq. Pathological lesions of the esophagus were dissected and digested into single-cell suspensions for further separation using FITC-CD45 antibody via FACS (1–4). CD45+ and CD45− cells whose numbers in different lesions are shown on right panel were scRNA-sequenced, respectively.
We therefore conducted a time-ordered single-cell transcriptomic profiling on various esophageal lesions in 4NQO-exposed mice. For stage NOR or stage INF, we used the whole esophagus from 30 or 20 mice. For other stages, we used the lesion foci and the sample numbers were 32 HYP from 25 mice, 24 DYS from 17 mice, 24 CIS from 20 mice and 25 ICA from 23 mice, respectively (Fig. 1c). A total of 66,089 cells including 29,975 CD45+ immune cells and 36,114 CD45− non-immune cells were obtained across various pathological stages (Fig. 1c; Supplementary Fig. 1b). The median unique molecular identifier (UMI) per cell was 7748 in immune cells and 9370 in non-immune cells (Supplementary Fig. 1c); with a median signal detect ability of 1936 genes for immune cells and 2620 genes for non-immune cells (Supplementary Fig. 1c), respectively. Based on the expressions of canonical markers, we classified immune cells into T cells, B cells, myeloid cells and natural killer cells (Fig. 1c; Supplementary Fig. 1d, e) and identified four clusters of non-immune cells including epithelial cells, fibroblasts, endothelial cells and myocytes by using t-distributed stochastic neighbor embedding (tSNE) (Fig. 1c; Supplementary Fig. 1f, g).
Identifying epithelial cell types during ESCC tumorigenesis
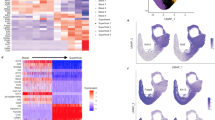
To discover how normal esophageal epithelium develops into invasive carcinoma, we next examined the expression alterations and functional changes in epithelial cells during the transition from normal to precancer or cancer. We identified 1756 epithelial cells across all six stages that were classified into six subtypes designated as EpiC 1 to EpiC 6 (Fig. 2a; Supplementary Fig. 2a and Supplementary Table 1). Through the analysis of pathway activities (Fig. 2b; Supplementary Fig. 2b), we found that EpiC 1 (n = 339) had higher expression of genes (e.g., Birc5, Mki67, Top2a, and Ube2c) in mitosis and proliferation8,9,10, ESCC tumor, dysplasia lesions and tumor-adjacent (>5 cm) normal tissues of the same patients (n = 4) used for LCM and esophageal lesions of different pathological stages (n = 45) used for bulk RNA sequencing were collected during surgery or endoscopy in Linzhou Esophageal Cancer Hospital (Henan Province, China) from 2018 to 2019. The various lesions were diagnosed independently by at least two pathologists according to the American Joint Committee on Cancer Eighth edition. No patient had received chemotherapy or radiotherapy before biopsy or surgery. This study was approved by the Institutional Review Boards of Cancer Hospital, Chinese Academy of Medical Sciences and informed consent was obtained from each patient. Clinical information was collected from patients’ medical records. Animal experiments in this study were conducted in compliance with approved protocols and guidelines from the Institutional Animal Care and Use Committee of the Chinese Academy of Medical Sciences. Eight-week-old female C57BL/6 mice, purchased from the Bei**g Huafukang bioscience company in China, were maintained in local housing facility of a controlled condition (23 ± 1 °C, 50 ± 10% humidity and 12–12 h light-dark cycle). Mice were treated with 4NQO (Sigma-Aldrich) in drinking water (100 μg/ml) for 16 weeks to induce multi-staged ESCC carcinogenesis. Drinking water containing the carcinogen was replaced once a week with freshly prepared one and mice were allowed to access drinking water ad libitum during treatment. After 16 weeks of carcinogen treatment, 4NQO drinking water was replaced by sterile water until the mice were killed. The esophageal lesions were identified by two independent pathologists based on the histopathological criteria described previously58 (Fig. 1a). Briefly, stage NOR was well-oriented stratified epithelium consists of basal zone and superficial zone. Stage INF was normal epithelium with focal aggregates of epithelial lymphocytes. Stage DYS was defined as loss of polarity in the epithelial cells, nuclear pleomorphism, hyperchromatic, and increased or abnormal mitoses. In stage HYP, these abnormalities were confined to the lower third of the epithelium while in DYS they present in lower two thirds of the epithelium. Lesions with such abnormal changes involving the entire thickness of epithelium were considered as carcinoma in situ (stage CIS). Stage ICA was defined as a lesion with invasion into the sub-epithelial tissues. Esophageal samples of 4NQO-induced mice were subjected to single-cell RNA sequencing at six different time points and indicated different pathological stages: stage NOR (at week 0), stage INF (at week 12), stage HYP at (week 20), stage DYS (at week 22), stage CIS (at week 24), and stage ICA (at week 26). A group of control mice treated without 4NQO were killed at month 1, 2, 8, and 25 (n = 2, respectively). The esophagus was removed immediately when the animal was killed. Cross-sections of the esophagus were cut and stored at –80 °C and sections of the frozen tissues were stained with hematoxylin–eosin (H&E) for histopathological examination and microdissection for bulk RNA sequencing. Tissue samples from human or mice were cut into 5–10 consecutive sections (8 μm) and the epithelial layer contained 30–50 cells on each section was micro-dissected with a Leica LMD7000 laser-capture microdissection system. RNA was isolated from the mini-bulk samples and cDNA was prepared for sequencing based on the Geo-seq protocol59. Sequencing libraries were built using the TruePrep DNA Library Prep Kit V2 for Illumina (Vazyme), and evaluated by Bioanalyzer (DNA HS kit, Agilent). RNA-seq data were mapped to GRCh38 human genome and GRCm38 murine genome by HISAT2 (version 2.1.0)60 with default parameter for human and murine samples, respectively. The gene expression matrix of raw reads counts after annotation by HTSeq (version 0.6.1p1) was processed using the DESeq2 (version 1.22.2)61 and visualized by showing the first 3 dimensions calculated by plotPCA function. We used TRIZOL to extract bulk RNA from patient biopsy (n = 45), and constructed sequencing library using NEBNext Ultra II RNA Library Prep Kit for Illumina. Sequencing data were processed by HISAT2 and HTseq as described above. The normalized expression from bulk samples of human precancerous lesions was used to estimated neutrophil fraction with CIBERSORT (version 1.06)62. For mice at stage NOR and INF, the whole esophagi were taken and for mice at other stages, the dysplastic/malignant lesions were taken immediately after killed. The tissue samples of ESCC and various precursor lesions were gently minced into small pieces and digested for with in RPMI-1640 medium (Invitrogen) containing collagenase IV (Gibco) and hyaluronidase (Sigma-Aldrich). CD45-FITC (553080, BD Biosciences, dilution 1:20) antibody staining was performed for fluorescence activated cell sorting (FACS) on a FACSAria sorter (BD Biosciences). Single cells with or without GFP signal, representing immune or non-immune cells, were sorted and captured respectively in nanoliter droplets using Chromium (10× Genomics). scRNA-seq libraries were prepared using Chromium Single Cell 5′ Reagent Kits (10× Genomics) and sequencing was accomplished with an Illumina HiSeq x10 System. Raw gene expression matrices obtained per sample using CellRanger (version 2.1.0, 10× genomics) were combined using the Seurat R package (version 2.3.4)63. Genes detected in <0.1% of all cells were filtered. We further excluded cells with gene counts <500 and cells that had >10% of mitochondrial gene expressions. After quality control, 66,089 cells were further analyzed for their gene expression profiles. The genes with normalized expression between 0.0125 and 3, and dispersion >0.5 were selected as highly variable genes. The resultants were first summarized by principle component analysis (PCA) and then first several PCs were selected for tSNE dimensional reduction using the default settings of the RunTSNE function. The numbers of resulting highly variable genes and the select PCs are shown in Supplementary Table 2. Cell clusters in the resulting two-dimensional representation were annotated as known biological cell types using canonical marker genes. We reanalyzed epithelial cells and stromal cells separately to identify their sub-clusters by using highly variable gene identification and dimensional reduction as described above (Supplementary Table 2). Cells with mix features were removed from further analysis (e.g., cells with both Cd3d and Cd19 expression indicating T cell and B cell Multiplet). Clusters were identified using FindClusters function, and the specific gene markers for each cluster were determined using the FindAllMarkers function implanted in Seurat package. Pathway analyses were predominantly performed on the 50 hallmark pathways described in the molecular signature database, exported using the MSigDB database (version 6.2)64. We also assessed biological process activities using a described biological process of Gene Ontology (GO) dataset. To assign pathway activity estimates to individual cells, we applied the GSVA using standard settings, as implemented in the GSVA package (version 1.30.0)65. To assess differential activities of pathways between sub-cluster of cells, we contrasted the activity scores for each cell using Limma package (version 3.38.3)66. Differential activities of pathways were calculated for each identified cluster. T-values of the results of some significant differential pathways (P < 0.05) in top 10 were visualized using heatmaps with average pathway activity scores of each cluster. SCENIC (version 1.1.0)67 was used to assess the transcriptional activity of epithelial cells with high quality (UMI > 5500). The analysis used the motifs database for RcisTarget and GRNboost (corresponding to GENIE3 1.4.3, AUCell 1.4.1 and RcisTarget 1.2.1; with mm10__refseq-r80__10kb_up_and_down_tss.mc9nr). The input matrix was read counts. Monocle 2 (version 2.10.1)68 was used for the trajectory analysis on high quality epithelial cells (UMI > 5500). All the top 100 markers of each cluster were used for the cell ordering. Dimensionality reduction and trajectory construction were performed on the selected genes with default methods and parameters. We calculated diffusion components using the RunDiffusion function as implemented in Seurat package with default parameters. The first three dimensions were used to draw diffusion maps. Mean coordinates of all of each cluster’s cells were considered as the center of the cluster. The farthest cell of stage NOR from the total distance of other stages was the start point. We used CellPhoneDB (version 2.0.6)69 with default arguments to reveal interaction between cell types. Each cluster analyzed was downsampled to 100 cells since the low cell number of some epithelial cluster. Interactions of epithelial cells with immune cells and immune cells with epithelial cells were demonstrated respectively. Formalin-fixed paraffin-embedded (FFPE) sections of esophageal precursor lesions were collected from 30 patients between 2016 and 2018 in Linzhou Cancer Hospital, including INF (n = 5), HYP (n = 13), DYS (n = 8) and CIS (n = 3), to validate the results obtained in mice. The protein expression levels of the marker genes were detected by IHC staining for mice tissues and immunofluorescence for human specimens with antibodies (Abcam) shown in Supplementary Table 3. The samples were incubated with antibody against Ki67 (1:50 for IHC, ab16667), Top2a (1:8000 for IHC, 1:10,000 for IF, ab52934), Aldh3a1 (1:200 for IHC, 1:600 for IF, ab76976), Atf3 (1:200 for IHC, 1:600 for IF, ab216569), S100a8 (1:500 for IHC, 1:1500 for IF, ab92331), Mmp14 (1:2000 for IHC, 1:6000 for IF, ab51074), or Itga6 (1:250 for IHC, 1:750 for IF, ab181551). Opal multiplex staining was performed according to the Opal 5-Color Manual IHC Kit (Perkin Elmer). Opal DAPI, Opal 520, Opal 570, Opal 620, and Opal 690 were used to generate different signals. Slides were counterstained with DAPI (1:2000) for nuclei visualization, and subsequently coverslipped using a VectaShield Hardset mounting media. The slides were imaged using Vectra Polaris Automated Quantitative Pathology Imaging System (Perkin Elmer). We used inForm software (Perkin Elmer) to unmix and remove autofluorescence and to analyze the multispectral images. Statistical analyses were conducted by using R v3.5.170 and Prism 7 (Graphpad Software). Pearson’s correlation was calculated with the R function cor() and the significance was determined using two-sided unpaired Wilcoxon rank-sum test. P < 0.05 was considered statistically significant. Further information on research design is available in the Nature Research Reporting Summary linked to this article.Methods
Human biospecimen collection
Induction of multi-staged ESCC development and sample preparations
Bulk RNA sequencing and data analysis
Single-cell RNA sequencing (scRNA-seq) and data analysis
Major cell-type clustering and marker gene identification
Gene set variation analysis (GSVA)
Analysis of transcription factor expression
Cell transition trajectory and diffusion map analysis
Analysis of interaction between cell types
Immunohistochemistry and immunofluorescent detection
Statistical analysis
Reporting summary
Data availability
The raw sequencing data and processed gene expression matrix of mouse model have been deposited in GSA (Genome Sequence Archive in BIG Data Center, Bei**g Institute of Genomics, Chinese Academy of Sciences, http://gsa.big.ac.cn) under the accession number CRA002118. The raw sequencing data of human esophageal tissues has been deposited in GSA-Human (https://bigd.big.ac.cn/gsa-human) under the accession number HRA000093. All the other data supporting the findings of this study are available within the article and its supplementary information files and from the corresponding author upon reasonable request. A reporting summary for this article is available as a Supplementary Information file. Source data are provided with this paper.
Code availability
Example scripts to process and analyze data are available at https://github.com/ESCCemAll/scESCC_mice. Detailed information will be available from the corresponding authors upon reasonable request.
References
Chen, X. X. et al. Genomic comparison of esophageal squamous cell carcinoma and its precursor lesions by multi-region whole-exome sequencing. Nat. Commun. 8, 524 (2017).
Chang, J. et al. Genomic analysis of oesophageal squamous-cell carcinoma identifies alcohol drinking-related mutation signature and genomic alterations. Nat. Commun. 8, 15290 (2017).
Gao, Y. et al. Genetic landscape of esophageal squamous cell carcinoma. Nat. Genet. 46, 1097–1102 (2014).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Tang, X. et al. Oral cavity and esophageal carcinogenesis modeled in carcinogen-treated mice. Clin. Cancer Res. 10, 301–313 (2004).
Chu, J. et al. Metabolic remodeling by TIGAR overexpression is a therapeutic target in esophageal squamous-cell carcinoma. Theranostics 10, 3488–3502 (2020).
Kelly, R. J. et al. Impacting tumor cell-fate by targeting the inhibitor of apoptosis protein survivin. Mol. Cancer 10, 35 (2011).
Nitiss, J. L. DNA topoisomerase II and its growing repertoire of biological functions. Nat. Rev. Cancer 9, 327–337 (2009).
Sales Gil, R. & Vagnarelli, P. Ki-67: more hidden behind a ‘classic proliferation marker’. Trends Biochem. Sci. 43, 747–748 (2018).
**ong, Y. et al. UBE2C functions as a potential oncogene by enhancing cell proliferation, migration, invasion, and drug resistance in hepatocellular carcinoma cells. Biosci. Rep. 39, BSR20182384 (2019).
Bhattacharjee, P. et al. Functional compensation of glutathione S-transferase M1 (GSTM1) null by another GST superfamily member, GSTM2. Sci. Rep. 3, 2704 (2013).
Chen, C. H. et al. Pharmacological recruitment of aldehyde dehydrogenase 3A1 (ALDH3A1) to assist ALDH2 in acetaldehyde and ethanol metabolism in vivo. Proc. Natl Acad. Sci. USA 112, 3074–3079 (2015).
Osier, M. V. et al. Possible epistatic role of ADH7 in the protection against alcoholism. Am. J. Med. Genet. B Neuropsychiatr. Genet. 126B, 19–22 (2004).
Carr, T. M. et al. JunB promotes Th17 cell identity and restrains alternative CD4(+) T-cell programs during inflammation. Nat. Commun. 8, 301 (2017).
Inoue, M. et al. The stress response gene ATF3 is a direct target of the Wnt/beta-catenin pathway and inhibits the invasion and migration of HCT116 human colorectal cancer cells. PLoS ONE 13, e0194160 (2018).
Liu, C. et al. Folate receptor alpha is associated with cervical carcinogenesis and regulates cervical cancer cells growth by activating ERK1/2/c-Fos/c-Jun. Biochem. Biophys. Res. Commun. 491, 1083–1091 (2017).
Tiedje, C. et al. The RNA-binding protein TTP is a global post-transcriptional regulator of feedback control in inflammation. Nucleic Acids Res. 44, 7418–7440 (2016).
Nedjadi, T. et al. S100A8 and S100A9 proteins form part of a paracrine feedback loop between pancreatic cancer cells and monocytes. BMC Cancer 18, 1255 (2018).
Sakamoto, K. et al. Down-regulation of keratin 4 and keratin 13 expression in oral squamous cell carcinoma and epithelial dysplasia: a clue for histopathogenesis. Histopathology 58, 531–542 (2011).
Brooks, D. L. et al. ITGA6 is directly regulated by hypoxia-inducible factors and enriches for cancer stem cell activity and invasion in metastatic breast cancer models. Mol. Cancer 15, 26 (2016).
Zhang, R. et al. Golgi membrane protein 1 (GOLM1) promotes growth and metastasis of breast cancer cells via regulating matrix metalloproteinase-13 (MMP13). Med. Sci. Monit. 25, 847–855 (2019).
Leung, D. W. et al. Vascular endothelial growth factor is a secreted angiogenic mitogen. Science 246, 1306–1309 (1989).
Nguyen, A. T. et al. Organelle specific o-glycosylation drives MMP14 activation, tumor growth, and metastasis. Cancer Cell 32, 639–653 e636 (2017).
Wang, L. et al. Extracellular matrix protein 1 (ECM1) is over-expressed in malignant epithelial tumors. Cancer Lett. 200, 57–67 (2003).
Park, Y. R. et al. MicroRNA-30a-5p (miR-30a) regulates cell motility and EMT by directly targeting oncogenic TM4SF1 in colorectal cancer. J. Cancer Res. Clin. Oncol. 143, 1915–1927 (2017).
Satelli, A. et al. Vimentin in cancer and its potential as a molecular target for cancer therapy. Cell Mol. Life Sci. 68, 3033–3046 (2011).
Nakabayashi, M. et al. PITX1 is a reliable biomarker for predicting prognosis in patients with oral epithelial dysplasia. Oncol. Lett. 7, 750–754 (2014).
Kastenhuber, E. R. et al. Putting p53 in context. Cell 170, 1062–1078 (2017).
Lee, Y. et al. BCLAF1 is a radiation-induced H2AX-interacting partner involved in γH2AX-mediated regulation of apoptosis and DNA repair. Cell Death Dis. 3, e359 (2012).
Rodgers, J. J. et al. ETS1 induces transforming growth factor beta signaling and promotes epithelial-to-mesenchymal transition in prostate cancer cells. J. Cell Biochem. 120, 848–860 (2019).
Puisieux, A. et al. Oncogenic roles of EMT-inducing transcription factors. Nat. Cell Biol. 16, 488–494 (2014).
Li, J. et al. Co-inhibitory molecule B7 superfamily member 1 expressed by tumor-infiltrating myeloid cells induces dysfunction of anti-tumor CD8(+) T cells. Immunity 48, 773–786 (2018).
**ao, X., Li, B., Mitton, B., Ikeda, A. & Sakamoto, K. M. Targeting CREB for cancer therapy: friend or foe. Curr. Cancer Drug Targets 10, 384–391 (2010).
Lee, J. H. et al. ELK3 promotes the migration and invasion of liver cancer stem cells by targeting HIF-1alpha. Oncol. Rep. 37, 813–822 (2017).
Clausen, M. J. et al. RAB25 expression is epigenetically downregulated in oral and oropharyngeal squamous cell carcinoma with lymph node metastasis. Epigenetics 11, 653–663 (2016).
Ogunwobi, O. O. et al. Epigenetic upregulation of HGF and c-Met drives metastasis in hepatocellular carcinoma. PLoS ONE 8, e63765 (2013).
Katoh, M. et al. Precision medicine for human cancers with Notch signaling dysregulation. Int. J. Mol. Med. 45, 279–297 (2020).
Danilo, M. et al. Suppression of Tcf1 by inflammatory cytokines facilitates effector CD8 T cell differentiation. Cell Rep. 22, 2107–2117 (2018).
Forster, R. et al. CCR7 and its ligands: balancing immunity and tolerance. Nat. Rev. Immunol. 8, 362–371 (2008).
Kaech, S. M. et al. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat. Immunol. 4, 1191–1198 (2003).
Barber, D. L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687 (2006).
Blackburn, S. D. et al. Coregulation of CD8+ T cell exhaustion by multiple inhibitory receptors during chronic viral infection. Nat. Immunol. 10, 29–37 (2009).
Kim, T. D. et al. Human microRNA-27a* targets Prf1 and GzmB expression to regulate NK-cell cytotoxicity. Blood 118, 5476–5486 (2011).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Annunziato, F. et al. The 3 major types of innate and adaptive cell-mediated effector immunity. J. Allergy Clin. Immunol. 135, 626–635 (2015).
Narayana, S. K. et al. The interferon-induced transmembrane proteins, IFITM1, IFITM2, and IFITM3 inhibit hepatitis C virus entry. J. Biol. Chem. 290, 25946–25959 (2015).
Etesam, Z. et al. Altered expression of specific transcription factors of Th17 (RORγt, RORα) and Treg lymphocytes (FOXP3) by peripheral blood mononuclear cells from patients with multiple sclerosis. J. Mol. Neurosci. 60, 94–101 (2016).
Carroll, M. C. The complement system in regulation of adaptive immunity. Nat. Immunol. 5, 981–986 (2004).
Stables, M. J. et al. Transcriptomic analyses of murine resolution-phase macrophages. Blood 118, e192–e208 (2011).
Arlauckas, S. P. et al. Arg1 expression defines immunosuppressive subsets of tumor-associated macrophages. Theranostics 8, 5842–5854 (2018).
Iannone, R. et al. Blockade of A2b adenosine receptor reduces tumor growth and immune suppression mediated by myeloid-derived suppressor cells in a mouse model of melanoma. Neoplasia 15, 1400–1409 (2013).
Mittal, S. K. et al. Suppression of antigen presentation by IL-10. Curr. Opin. Immunol. 34, 22–27 (2015).
Pelletier, M. et al. Evidence for a cross-talk between human neutrophils and Th17 cells. Blood 115, 335–343 (2010).
Abnet, C. C. et al. Epidemiology of esophageal squamous cell carcinoma. Gastroenterology 154, 360–373 (2018).
Zhang, P. et al. Dissecting the single-cell transcriptome network underlying gastric premalignant lesions and early gastric cancer. Cell Rep. 27, 1934–1947 (2019).
Bernard, V. et al. Single-cell transcriptomics of pancreatic cancer precursors demonstrates epithelial and microenvironmental heterogeneity as an early event in neoplastic progression. Clin. Cancer Res. 25, 2194–2205 (2019).
Dawsey, S. M. et al. Squamous esophageal histology and subsequent risk of squamous cell carcinoma of the esophagus. A prospective follow-up study from Linxian, China. Cancer 74, 1686–1692 (1994).
Chen, J. et al. Spatial transcriptomic analysis of cryosectioned tissue samples with Geo-seq. Nat. Protoc. 12, 566–580 (2017).
Kim, D. et al. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Love, M. I. et al. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Butler, A. et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Hanzelmann, S. et al. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2018).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (81725015 to C. Wu, 21675098 to J.W., 21927802 to J.W.), Medical and Health Technology Innovation Project of Chinese Academy of Medical Sciences (2016-I2M-3-019 to D.L., 2016-I2M-4-002 to C. Wu., 2019-I2M-2-001 to D.L. and C. Wu), Bei**g Outstanding Young Scientist Program (BJJWZYJH01201910023027 to C. Wu), 2018 Bei**g Brain Initiative (Z181100001518004 to J.W.), Bei**g Advanced Innovation Center for Genomics, and Bei**g Advanced Innovation Center for Structural Biology.
Author information
Authors and Affiliations
Contributions
J.W., C. Wu., and D.L. conceptualized and supervised this study. J.Y., Q.C., W.F., Y.C., and T.L. contributed to the study design and performed most experiments. W.G., Y.X., and L.L. were engaged in sample collection and preparation. Y.M., J.Y., X.Z., and Y. Lin contributed to bioinformatics and statistical analysis. C. Wang, L.P., Y. Luo, and A.L. were responsible for animal experiments. J.Y., Q.C., W.F., Y.M., and Y.C. drafted the manuscript. W.T., C. Wu., J.W., and D.L. reviewed and prepared the final manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Marnix Jansen, Shao Li and the other, anonymous, reviewer for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yao, J., Cui, Q., Fan, W. et al. Single-cell transcriptomic analysis in a mouse model deciphers cell transition states in the multistep development of esophageal cancer. Nat Commun 11, 3715 (2020). https://doi.org/10.1038/s41467-020-17492-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-17492-y
- Springer Nature Limited
This article is cited by
-
Single-cell profiling of response to neoadjuvant chemo-immunotherapy in surgically resectable esophageal squamous cell carcinoma
Genome Medicine (2024)
-
scRNA-seq revealed high stemness epithelial malignant cell clusters and prognostic models of lung adenocarcinoma
Scientific Reports (2024)
-
Decoding spatiotemporal transcriptional dynamics and epithelial fibroblast crosstalk during gastroesophageal junction development through single cell analysis
Nature Communications (2024)
-
Tumor cell enrichment by tissue suspension improves sensitivity to copy number variation in diffuse gastric cancer with low tumor content
Scientific Reports (2024)
-
Reciprocal inhibition between TP63 and STAT1 regulates anti-tumor immune response through interferon-γ signaling in squamous cancer
Nature Communications (2024)