Abstract
Metastasis is a bottleneck in cancer treatment. Studies have shown the pivotal roles of long noncoding RNAs (lncRNAs) in regulating cancer metastasis; however, our understanding of lncRNAs in gastric cancer (GC) remains limited. RNA-seq was performed on metastasis-inclined GC tissues to uncover metastasis-associated lncRNAs, revealing upregulated small nucleolar RNA host gene 26 (SNHG26) expression, which predicted poor GC patient prognosis. Functional experiments revealed that SNHG26 promoted cellular epithelial–mesenchymal transition and proliferation in vitro and in vivo. Mechanistically, SNHG26 was found to interact with nucleolin (NCL), thereby modulating c-Myc expression by increasing its translation, and in turn promoting energy metabolism via hexokinase 2 (HK2), which facilitates GC malignancy. The increase in energy metabolism supplies sufficient energy to promote c-Myc translation and expression, forming a positive feedback loop. In addition, metabolic and translation inhibitors can block this loop, thus inhibiting cell proliferation and mobility, indicating potential therapeutic prospects in GC.
Similar content being viewed by others
Introduction
Gastric cancer (GC) is the fifth most commonly diagnosed tumor in humans, with over 1 million estimated new cases diagnosed annually [1, 2]. GC is the fourth most common cause of cancer-related death due to its high metastatic frequency, with over 60% of patients presenting with local or distant metastasis at diagnosis [3, 4]. Despite substantial progress in the development of molecular targeted drugs and immunotherapy, the 5-year survival rate of GC patients remains unsatisfactory due to metastasis [5, 6]. Thus, investigations into the mechanisms underlying GC metastasis are urgently required for the development of new therapeutic approaches for GC.
Long noncoding RNAs (lncRNAs) exceed 200 nucleotides and have limited protein-coding capacities [7, 8]. LncRNAs are known to be widely expressed and play pivotal roles in gene regulation [9]. LncRNAs perform diverse functions through interactions with RNA, DNA, and proteins [10]. Numerous studies have demonstrated that aberrant expression of lncRNAs is associated with cancer cell behaviors [11]. For example, lncRNA C1RL-AS1 promotes GC proliferation and metastasis via the AKT/β-catenin pathway [12]. The lncRNA BDNF-AS/WDR5/FBXW7 axis mediates ferroptosis in GC peritoneal metastasis by regulating VDAC3 ubiquitination [3A). As shown in Fig. 3D, the in vitro RNA pull-down indicated that the 1–500 bp sequence of SNHG26 was crucial for its interaction with NCL. Regarding the protein domain (Fig. 3E, Supplementary Fig. 3B), SNHG26 interacted with the 382–710 amino acid (aa) region of NCL (Fig. 3F). Overall, these results suggest that SNHG26 likely exerts its function by binding with NCL.
A. RNA pull-down assay showing the proteins that bind with biotinylated SNHG26 in vitro by silver staining. B. Immunoblotting for specific associations of NCL with SNHG26 from RNA pull-down assays. C. RIP assays were performed using antibodies against NCL, and qPCR was used to detect SNHG26 enrichment. D. Immunoblotting for NCL in samples pulled down with full-length biotinylated-SNHG26 or truncated RNA motifs. E, F. Deletion map** to identify the domains of NCL that bind to SNHG26. RIP analysis for SNHG26 enrichment in cells transiently transfected with full-length or truncated FLAG-tagged constructs.
SNHG26 promotes mRNA translation in GC
NCL is a nucleocytoplasmic protein involved in many biological processes, including ribosomal assembly and rRNA processing, indicating that NCL might be linked with ribosomal biogenesis [24, 25]. Intriguingly, we found that NCL could modulate the expression of ribosomal proteins, such as RPS3 and RPL4 (Supplementary Fig. 4A). The qPCR results showed that ribosomal prerequisite RNA (47 S rRNA) expression was upregulated after SNHG26 overexpression and vice versa when silencing SNHG26 (Supplementary Fig. 4B). Additionally, NCL might interact with RPS3 and RPL4 (www.string .com) (Supplementary Fig. 4C). Co-IP revealed that NCL interacted with RPS3 and RPL4 in the SNHG26 vector and overexpressing MKN-28 cells (Supplementary Fig. 4D). Moreover, a recent study demonstrated that NCL binds to the promoter and coding regions of rDNA and stimulates rDNA transcription [26]. Given the above findings, we used ChIP analysis to verify the distribution of NCL on the rDNA of GC cells at specific rDNA promoters and coding regions. SNHG26 overexpression increased NCL occupancy in the rDNA coding region (H8, H13; Supplementary Fig. 4E). This finding suggests that SNHG26 promotes the binding of NCL to the rDNA promoter and coding regions to enhance transcription.
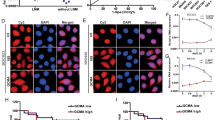
Dysregulated ribosomal biogenesis is usually associated with protein translation, indicating that SNHG26 might promote protein translation via NCL. We then performed an OPP assay to detect active protein synthesis in GC cells to test this hypothesis. Significantly increased protein synthesis was observed in SNHG26-overexpressing MKN-28 cells, whereas protein synthesis was reduced in HGC-27 and MKN-28 cells after SNHG26 knockdown (Fig. 4A–D). In addition, we performed SUnSET assays to detect the puromycin-labeled polypeptides and evaluate global protein synthesis. Consistent with the OPP assay results, global protein synthesis changes were detected after ectopic expression of SNHG26 (Fig. 4E). Overall, the above results suggest that SNHG26 promotes mRNA translation and protein synthesis in GC.
A–C Protein synthesis was measured in MKN-28 and HGC-27 GC cell lines with SNHG26 overexpressed or knocked down (OPP, green; DAPI, blue). Scale bars = 25 μM. D Quantification of Alexa 488 fluorescence intensity. E Western blotting was performed to assess protein synthesis in MKN-28 and HGC-27 cell lines overexpressing or knocking down SNHG26 treated with puromycin labeling, followed by detection with puromycin antibody. All data are from three independent experiments. The data are presented as the mean ± SD values (n = 3). *P <0.05; **P <0.01; ***P <0.001; ****P<0.0001.
LncRNA SNHG26 promotes the proliferation and migration of GC cells by stimulating c-Myc translation in an NCL-dependent manner
Interestingly, previous studies have shown that NCL can directly bind with the promoter region of c-Myc and transcriptionally regulate its expression [27]. We first performed RIP assays to investigate whether SNHG26 binding to NCL also transcriptionally promotes c-Myc expression. As shown in Fig. 5A, our experimental results revealed that NCL can bind to c-Myc mRNA. However, lncRNA SNHG26 significantly regulated c-Myc expression at the protein level but not at the mRNA level (Fig. 5B, C), suggesting that SNHG26 does not transcriptionally upregulate c-Myc expression, different from the reported mechanism by which NCL upregulates c-Myc expression. Moreover, after treatment with CHX over time, the half-life of c-Myc did not change after SNHG26 overexpression (Fig. 5D). The above results indicate that SNHG26 regulates c-Myc not through transcription but possibly through a translational mechanism.
A c-Myc mRNA was highly enriched in the RNAs enriched by anti-Flag NCL RIP by RT-qPCR in HGC-27 cells. B mRNA expression of c-Myc was detected by qPCR after modulating SNHG26 status in MKN-28 and HGC-27 GC cells. C Protein expression of c-Myc was examined by western blotting after modulating SNHG26 status in MKN-28 and HGC-27 GC cells. D A CHX (100 µg/ml) chase experiment showed no significant change in the half-life of c-Myc protein in MKN-28 cells overexpressing SNHG26. E Percentage of c-Myc mRNAs measured by qPCR in each fraction collected from the polysome profiling after overexpressing SNHG26 in MKN-28 cells. F Expression of pRF with pRmF of firefly or Renilla luciferase in 293 T cells after overexpression or knockdown of SNHG26. G Western blotting showing that the increases in the levels of NCL and c-Myc were significantly reversed after transfection of siNCL into SNHG26-overexpressed MKN-28 cells. H The expression of firefly versus Renilla luciferase was evaluated after transfection of cells stably expressing the empty vector, SNHG26 gene, with pRF, pRmF reporter plasmid, and NC, siNCL small interfering RNA. I A Co-IP assay verified the presence of interactions between NCL and EIF4AI/II. J MKN-28 cells overexpressing SNHG26 were transfected with shc-Myc or shNC for 48 h. Cell malignant metastasis was determined by Transwell assays. K MKN-28 cells overexpressing SNHG26 were transfected with shc-Myc or shNC for 48 h. Cell proliferation was determined by CCK8. L Western blot showing that the increases in N-cadherin, Vimentin, Snail, and c-Myc levels were significantly reversed after transfection of shc-Myc plasmid into SNHG26-overexpressing cells for 48 h. In contrast, the decrease in E-cadherin levels was significantly reversed. All data are from three independent experiments. The data are presented as the mean ± SD values (n≥ 3). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
To determine whether SNHG26 regulates c-Myc translation, we conducted polysome profiling in SNHG26-overexpressing cells. The qPCR results showed that upregulated SNHG26 significantly increased the translation of c-Myc mRNA in MKN-28 cells (Fig. 5E). To revalidate the experimental results of polysome profiling, we used bicistronic reporters to investigate the 5’ cap- and internal ribosomal entry site (IRES)-dependent translation efficiencies. The luciferase reporter results indicated that, compared to control cells, SNHG26-overexpressing cells demonstrated increased IRES-dependent translation activity, as indicated by increased firefly luciferase activity (Fig. 5F). It has been reported that NCL can self-regulate its expression by binding to the c-Myc promoter [28]. To explore whether NCL can increase its transcription by binding to the promoter of c-Myc in GC, we conducted a ChIP experiment; however, NCL did not bind to c-Myc (Supplementary Fig. 5A), consistent with the literature. It has been shown that typical posttranscriptional Cap-dependent translation is achieved by an increase in RNA binding to ribosomes [29]. In contrast, IRES-dependent translation does not result in an increase in RNA binding and may be achieved by increasing the rate of bound ribosome assembly. We first performed western blotting experiments to demonstrate whether SNHG26 promotes c-Myc protein translation dependent on NCL. NCL knockdown in SNHG26-overexpressing GC cells significantly downregulated c-Myc expression (Fig. 5G). Luciferase reporter gene experiments showed that NCL knockdown reduced the IRES-dependent translational activity of c-Myc caused by SHNG26 overexpression (Fig. 5H). Additionally, Co-IP showed that NCL interacted with the translation initiation factor EIF4AI/II (Fig. 5I). The above results suggest that SNHG26 recruits NCL to the IRES site of c-Myc mRNA and facilitates NCL recruitment of the transcription initiation factor EIF4AI/II with ribosomes to initiate IRES-dependent Myc translation. To determine whether function is exerted through c-Myc, we silenced c-Myc in GC cells overexpressing SNHG26 and controls. CCK8 and Transwell assays as well as western blotting showed that downregulation of c-Myc expression could reverse the alteration of proliferation, migration, and EMT marker expression in GC cells caused by SNHG26 overexpression (Fig. 5J–L, Supplementary Fig. 5B).
CX-5461 is a recently developed inhibitor of ribosomal RNA synthesis that selectively inhibits Pol I-driven transcription, DNA replication, and protein translation [30]. To determine whether CX-5461 inhibits the proliferation and migration ability of GC cells, we performed CCK-8 and Transwell assays in MKN-28 cells, revealing that CX-5461 treatment decreased the proliferation and migration abilities of GC cells compared to those observed in the control group (Supplementary Fig. 5C, D). The subsequent OPP and luciferase reporter gene assays showed that the increase in translation of SNHG26 action was similarly inhibited by treatment with CX-5461 (Supplementary Fig. 5E–G). Overall, CX-5461 inhibited SNHG26-mediated protein translation, thus suppressing the migration and proliferation capabilities of GC cells.
SNHG26 regulates energy metabolism through the c-Myc/HK2 pathway, facilitating GC proliferation and migration
Recently, several studies have indicated that dysregulated energy metabolism supports cancer cell growth and facilitates metastasis [31]. Previous studies have shown a link between c-Myc regulation and cellular metabolism, proliferation, and metastasis [32]. To explore whether SNHG26 demonstrates the above mechanism, we first performed targeted metabolomics in Metware Biotech Company (Wuhan, China). In this assay, 42 metabolites were tested for differentially abundant metabolite screening by fold change ≥1.5 or ≤0.67 as metabolic thresholds. Multiple differentially abundant metabolites were identified, thirteen upregulated and seven downregulated (Fig. 6A). There was an upregulation in the levels of metabolites of interest, such as glucose, lactate, and ATP (Fig. 6B). Given the targeted metabolomics results, we verified the changing levels of glucose, lactate, and ATP. Glucose uptake, lactate production, and ATP levels were dramatically increased after overexpressing SNHG26 in MKN-28 cells and decreased when SNHG26 was knocked down in HGC-27 cells (Fig. 6C–H). Next, we silenced c-Myc in GC cells overexpressing SNHG26, significantly decreasing glucose uptake, lactate production, and ATP levels (Fig. 6I).
A Differentially abundant metabolite bar chart. B Differentially abundant metabolite histograms with horizontal coordinates indicating group and vertical coordinates indicating expression. C–E Changes in glucose uptake, lactate production, and ATP levels in MKN-28 cells overexpressing SNHG26. F–H Changes in glucose uptake, lactate production, and ATP levels in MKN-28 and HGC-27 cells with SNHG26 knockdown. I Glucose uptake, lactate production, and ATP levels were enhanced after lncRNA SNHG26 overexpression, while changes in metabolic levels were reversed by silencing c-Myc. J–L. The increases in glucose uptake, lactate production, and ATP levels were significantly reversed after transfection of the shHK2 plasmid into SNHG26-overexpressing cells. M Western blotting showing that the increase in HK2 levels was significantly reversed after transfection of the shc-Myc plasmid into SNHG26-overexpressing cells for 48 h. All data are from three independent experiments. The data are presented as the mean ± SD values (n ≥ 3). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
To explore which glycolysis-related proteins can be regulated by SNHG26, we first measured the expression of glycolysis-related proteins after modulating SNHG26. As shown in Supplementary Fig. 6A, HK2 but not GLUT1, LDHA, LDHB, ALDOA, or ALDOB was significantly upregulated after overexpressing SNHG26. We knocked down HK2 in SNHG26-overexpressing MKN-28 cells, and as expected, the detected glucose uptake, lactate production, and ATP levels were inhibited (Fig. 6J–L). We silenced the expression of c-Myc, and the expression of HK2 was correspondingly downregulated (Fig. 6M). Then, we detected the proliferation and migration capacities and EMT-associated markers of MKN-28 SNHG26-overexpressing cells after HK2 knockdown. The Transwell and CCK8 assays as well as western blotting analyses showed that the migration and proliferation abilities and EMT marker expression patterns were reversed after silencing HK2 expression (Fig. 7A–C). Interestingly, c-Myc expression was also downregulated after HK2 knockdown (Fig. 7C).
A Transwell assays showed that the increased migration of SNHG26-overexpressing cells was significantly reversed after transfection with the shHK2 plasmid. Scale bars = 100 μM. B The CCK-8 assay showed that the increased proliferation of SNHG26-overexpressing cells was significantly reversed after transfection with the shHK2 plasmid. C Western blotting showing that the increases in HK2, N-cadherin, Vimentin, Snail, and c-Myc levels were significantly reversed after transfection of the shHK2 plasmid into SNHG26-overexpressing cells for 48 h. In contrast, the decrease in E-cadherin levels was significantly reversed. D–F Glucose uptake, lactate production, and ATP levels were significantly decreased after the addition of 2-DG (5 mM) in control cells and SNHG26-overexpressing cells. G MKN-28 cells overexpressing SNHG26 were treated with 2-DG (5 mM). Cell proliferation was determined by CCK8 assay. H MKN-28 cells overexpressing SNHG26 were treated with 2-DG (5 mM). Cell malignant metastasis was determined by Transwell assay. Scale bars = 100 μM. All data are from three independent experiments. The data are presented as the mean ± SD values (n ≥ 3). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
To investigate whether SNHG26 promotes GC cell proliferation and migration through the HK2-mediated glycolytic pathway, we added the HK2 inhibitor 2-DG to SNHG26-overexpressing cells to inhibit glycolysis, and treatment with 2-DG abolished the SNHG26-induced increase in glucose uptake, lactate production, and ATP production (Fig. 7D–F). We next assessed the effect of 2-DG on the proliferation and migratory capacity of GC cells. 2-DG significantly reduced the proliferation and migration capabilities of SNHG26-OE cells (Fig. 7G, H). We also performed ATP compensation experiments, revealing that the proliferation and metastatic capacities of the cells were significantly improved by back-supplementing ATP in the presence of inhibited ATP levels (Supplementary Fig. 7A, B). This finding suggests that SNHG26 regulates energy metabolism through the c-Myc/HK2 pathway and promotes GC proliferation and migration.
SNHG26 promotes energy production through regulation of the c-Myc/HK2 pathway, and positive feedback promotes the translation and expression of c-Myc
As we had previously found that c-Myc expression was downregulated after HK2 knockdown, we hypothesized that SNHG26 promotes energy production by regulating the c-Myc/HK2 pathway, which positively feeds back to promote c-Myc translation and expression. To test this hypothesis, we applied an OPP assay and luciferase reporter gene assay, and the results suggested that the overall protein translation and translation abilities of the c-Myc IRES were inhibited after the addition of 2-DG in SNHG26-overexpressing MKN-28 GC and control cells, respectively (Supplementary Fig. 7C–E). Western blotting indicated that the addition of 2-DG inhibited c-Myc expression, while the addition of ATP partially restored its expression (Supplementary Fig. 7F). When we added oligomycin, an ATP inhibitor, overexpression of SNHG26 caused an increase in c-Myc expression that was suppressed by oligomycin (Supplementary Fig. 7G). The above results suggest that SNHG26 promotes energy production by regulating the c-Myc/HK2 pathway, and positive feedback promotes the translation and expression of c-Myc.
The combination of a translation inhibitor (CX-5461) and a metabolic inhibitor (2-DG) had significant therapeutic effects in vivo and vitro
We next demonstrated the effects of treatment with the metabolic inhibitor 2-DG alone, the ribosome inhibitor CX-5461 alone, and the combination of 2-DG and CX-5461 on translation and metabolic levels in HEK293T and MKN-28 cells and found that the combination treatment group significantly reduced translation levels, glucose uptake, and lactate production in GC cells (Fig. 8A–C, Supplementary Fig. 8A, B). Interestingly, the level of ATP was higher in the CX-5461 group than in the control group (Fig. 8D). We speculate that capacity consumption may decrease after translation inhibition. Although we found that treatment with each drug alone inhibited GC cell proliferation and migration compared to controls, 2-DG combined with CX-5461 was significantly more potent than single-drug treatment. The inhibitory effect of the combination of 2-DG and CX-5461 in the SNHG26 overexpression group was more pronounced than that in the control group (Fig. 8E, F, Supplementary Fig. 8C). Moreover, we generated a subcutaneous tumor model by injecting control cells and SNHG26-overexpressing MKN-28 cells subcutaneously into nude mice. After 2 weeks, the mice were randomly assigned into four groups, one receiving blank saline, one receiving the ribosomal inhibitor CX-5461, one receiving the metabolic inhibitor 2-DG, and the last receiving a ribosomal inhibitor in combination with a metabolic inhibitor. After 4 weeks, the mice were sacrificed, and the tumors were weighed and measured. As indicated, injection of MKN-28 cells overexpressing SNHG26 significantly promoted tumor progression. Mice in the groups receiving ribosomal or metabolic inhibitors had smaller tumors than those in the control group, the tumor volume in the combination treatment group was smaller than that in the other three groups (Fig. 8G, H, Supplementary Fig. 8D), suggesting combination of the two inhibitors could increase the antitumor effect. Overall, our results suggest that SNHG26/NCL/c-Myc could be a potential biomarker of GC and therapeutic target.
A 293 T cells stably expressing the empty vector and SNHG26 gene were transfected with pRF and pRmF reporter plasmids with or without 2-DG and CX-5461 and assessed for firefly and Renilla luciferase expression. B–D. Changes in glucose uptake, lactate production, and ATP levels in MKN-28 cells were examined after treatment with 2-DG alone, CX-5461 alone, and combined treatment with 2-DG and CX-5461. E, F The proliferation and migration abilities of MKN-28 cells were analyzed by CCK-8 and Transwell assays after treatment with 2-DG alone, CX-5461 alone, and the combination of 2-DG and CX-5461. G Nude mice were injected with control or SNHG26-overexpressing cells and treated with blank (PBS), 2-DG (500 mg/kg), CX-5461 (40 mg/kg), or both drugs in combination after 1 week. Images of xenograft tumors in nude mice sacrificed after 4 weeks (n = 6). H Tumor weights of nude mice treated as described above. All data are from three independent experiments. The data are presented as the mean ± SD values (n ≥ 3). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Discussion
In the present study, we identified a novel lncRNA termed SNHG26 that was upregulated in GC tissues and predicted poor prognosis in GC patients. The subsequent loss- and gain-of-function analyses revealed that SNHG26 promoted the proliferation and metastasis of GC. Mechanistically, SNHG26 could directly interact with NCL, which binds to c-Myc at the mRNA level. Importantly, SNHG26 could promote c-Myc translation in an IRES-dependent manner via NCL. Furthermore, SNHG26 mediated glycolysis and energy metabolism through the c-Myc/HK2 pathway, promoting the proliferation and migration of GC cells. The increased glycolysis metabolism supplies energy to facilitate mRNA translation, forming a positive feedback loop. In addition, metabolic and translation inhibitors blocked the action of this positive feedback loop, thus inhibiting cancer cell proliferation and mobility, and the combination of both inhibitors could increase the antitumor effect (Fig. 9), indicating the potential therapeutic prospects in GC.
The lncRNA SNHG26 interacts with NCL and modulates c-Myc expression by increasing its translation to promote energy metabolism via HK2, which in turn promotes GC metastasis. The increase in energy metabolism supplies sufficient energy to promote the translation and expression of c-Myc, forming a positive feedback loop that promotes GC cell migration and invasion.
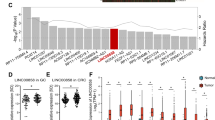
Numerous studies have demonstrated that alterations in lncRNA expression can promote tumorigenesis and metastasis in diverse malignant tumors and serve as biomarkers or therapeutic targets [9, 11]. SNHGs are a family of lncRNAs derived from snoRNAs, and SNHG1 [33, 34], SNHG5 [35], and SNHG20 [36, 37] have been reported to participate in tumor prognosis in various cancers. In the current study, according to the RNA-seq results obtained from six paired EGC tissues, we screened and identified a lncRNA termed SNHG26. However, the biological function of SNHG26 in cancer remains largely unknown. Only one study has been conducted by Jiang et al. and found that SNHG26 promoted cisplatin resistance in tongue squamous cell carcinoma through the PGK1/AKT/mTOR pathway [38]. Here, in vitro and in vivo experiments revealed that SNHG26 promoted proliferation and metastasis in GC, and high SNHG26 expression was associated with poor prognosis. To the best of our knowledge, this is the first study to demonstrate the biological function of SNHG26 in GC.
Many studies have demonstrated that lncRNAs can directly interact with DNA/RNA/protein and exert diverse functions. In the present study, combined with RNA pull-down and RIP assays, we found that SNHG26 could interact with NCL, a nucleocytoplasmic RNA-binding protein that plays multiple roles in cancer [39,40,41]. Several studies have demonstrated that NCL is mainly distributed in the nucleolus and can bind with ribonucleoprotein (RNP) or precursor ribosomal rRNA, thus affecting ribosomal biogenesis and enhancing the translation of target mRNAs [39]. Interestingly, the Co-IP results showed that NCL directly binds with the ribosomal proteins RPS3 and RPL4, suggesting that SNHG26 might impact mRNA translation via NCL. Indeed, our study revealed that SNHG26 could upregulate rRNA and the ribosomal proteins RPS3 and RPL4, providing a material basis for promoting translation. Subsequent OPP and SUnSET assays demonstrated that SNHG26 promoted mRNA translation in GC.
Interestingly, previous studies have shown that NCL could bind to the c-Myc promoter, promoting its expression via transcription [27]. Consistently, our RIP assay demonstrated that NCL interacted with c-Myc mRNA. Thus, we investigated whether SNHG26 transcriptionally modulates c-Myc expression via NCL. We subsequently found that lncRNA SNHG26 could modulate c-Myc expression at the protein level, whereas no difference was detected at the mRNA level. Moreover, no significant difference in the half-life between MKN-28 vector and MKN-28 SNHG26/overexpressing cells was observed, indicating that SNHG26 might regulate c-Myc expression translationally. In accordance with this hypothesis, the luciferase assay and polysome profiling demonstrated that SNHG26 promoted c-Myc translation in an IRES-dependent manner, indicating that SNHG26 could recruit NCL to Myc RNA and promote its translation. It has been reported in the literature that in viruses, NCL can bind to the IRES site of target gene RNA and recruit translation initiation factors to promote IRES-dependent translation of target genes [39]. Our IP experiments revealed that in human GC cells, SNHG26 promoted the binding of NCL to the translation initiation factor eIF4A and ribosomal protein, suggesting that SNHG26 recruits NCL to Myc RNA and promotes its binding to the translation initiation factor, in turn recruiting ribosomes to Myc RNA to initiate the translation process.
c-Myc, one of the most important oncoproteins, is well known to play critical roles in various biological processes, including energy metabolism [32]. Most tumor cells rely on glycolysis for energy production to maintain rapid proliferation even under aerobic conditions, and conventional mitochondrial oxidative phosphorylation is inhibited, leading to increased glucose consumption and lactate accumulation, a phenomenon called the Warburg effect [31, 42, 43]. In this study, we found that SNHG26 promoted glycolysis and ATP energy metabolism through the c-Myc/HK2 pathway in GC. Dysregulated energy metabolism is not only related to abnormal tumor proliferation but is also involved in tumor metastasis. For example, TIMMDC1 regulates abnormal glycolysis through the AKT/GSK3β/β-catenin signaling pathway, thus promoting GC metastasis [44]. However, the specific regulatory network of energy metabolism involved in tumor metastasis requires further exploration and clarification. In our study, the underlying mechanism is likely the SNHG26-mediated increase in c-Myc mRNA translation, which promotes c-Myc expression and thus mediates energy metabolism, promoting GC metastasis. Moreover, the increased glycolysis metabolism would supply energy to facilitate mRNA translation, forming a positive feedback loop. Recent studies have shown that glucose starvation could block the translation of proteins to save energy for cancer survival [43]. Consistently, our further studies revealed that metabolic and translation inhibitors can block the effect of this positive feedback regulatory loop, and the combination of both could increase the antitumor effect, suggesting potential future therapeutic targets.
In summary, we identified a novel lncRNA, SNHG26, which is upregulated in GC. SNHG26 interacts with NCL and promotes GC proliferation and metastasis by increasing c-Myc translation. Our study revealed a novel mechanism for lncRNAs in the association between translation and metastasis, implicating the SNHG26/NCL/c-Myc axis as a potential therapeutic target in GC.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Pineros M, et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144:1941–53.
McLean MH, El-Omar EM. Genetics of gastric cancer. Nat Rev Gastroenterol Hepatol. 2014;11:664–74.
Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F. Gastric cancer. Lancet. 2020;396:635–48.
Zhu XD, Huang MZ, Wang YS, Feng WJ, Chen ZY, He YF, et al. XELOX doublet regimen versus EOX triplet regimen as first-line treatment for advanced gastric cancer: an open-labeled, multicenter, randomized, prospective phase III trial (EXELOX). Cancer Commun. 2022;42:314–26.
Yuan L, Xu ZY, Ruan SM, Mo S, Qin JJ, Cheng XD. Long non-coding RNAs towards precision medicine in gastric cancer: early diagnosis, treatment, and drug resistance. Mol cancer. 2020;19:96.
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77:3965–81.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47:199–208.
Chen L, Dzakah EE, Shan G. Targetable long non-coding RNAs in cancer treatments. Cancer Lett. 2018;418:119–24.
Ma H, Hao Y, Dong X, Gong Q, Chen J, Zhang J, et al. Molecular mechanisms and function prediction of long noncoding RNA. Sci Worl J. 2012;2012:541786.
Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21:1253–61.
Zhen-Hua W, Yi-Wei G, Li-Qin Z, Jie-Yun Z, Zhe G, Wei-Jian G. Silencing of LncRNA C1RL-AS1 suppresses the malignant phenotype in gastric cancer cells via the AKT/beta-catenin/c-Myc pathway. Front Oncol. 2020;10:1508.
Huang G, **ang Z, Wu H, He Q, Dou R, Lin Z, et al. The lncRNA BDNF-AS/WDR5/FBXW7 axis mediates ferroptosis in gastric cancer peritoneal metastasis by regulating VDAC3 ubiquitination. Int J Biol Sci. 2022;18:1415–33.
Sun D, Gou H, Wang D, Li C, Li Y, Su H, et al. LncRNA TNFRSF10A-AS1 promotes gastric cancer by directly binding to oncogenic MPZL1 and is associated with patient outcome. Int J Biol Sci. 2022;18:3156–66.
Reticker-Flynn NE, Zhang W, Belk JA, Basto PA, Escalante NK, Pilarowski GOW, et al. Lymph node colonization induces tumor-immune tolerance to promote distant metastasis. Cell. 2022;185:1924–42.e23.
Dragomir MP, Kopetz S, Ajani JA, Calin GA. Non-coding RNAs in GI cancers: from cancer hallmarks to clinical utility. Gut. 2020;69:748–63.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Anders S, Pyl PT, Huber W. HTSeq–a python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Wu ZH, Liu CC, Zhou YQ, Hu LN, Guo WJ. OnclncRNA-626 promotes malignancy of gastric cancer via inactivated the p53 pathway through interacting with SRSF1. Am J cancer Res. 2019;9:2249–63.
Schmidt EK, Clavarino G, Ceppi M, Pierre P. SUnSET, a nonradioactive method to monitor protein synthesis. Nat methods. 2009;6:275–7.
Jia W, Yao Z, Zhao J, Guan Q, Gao L. New perspectives of physiological and pathological functions of nucleolin (NCL). Life Sci. 2017;186:1–10.
Wang X, Yu H, Sun W, Kong J, Zhang L, Tang J, et al. The long non-coding RNA CYTOR drives colorectal cancer progression by interacting with NCL and Sam68. Mol cancer. 2018;17:110.
Okur MN, Lee JH, Osmani W, Kimura R, Demarest TG, Croteau DL, et al. Cockayne syndrome group A and B proteins function in rRNA transcription through nucleolin regulation. Nucleic acids Res. 2020;48:2473–85.
Gonzalez V, Guo K, Hurley L, Sun D. Identification and characterization of nucleolin as a c-myc G-quadruplex-binding protein. J Biol Chem. 2009;284:23622–35.
Wu R, Li L, Bai Y, Yu B, **e C, Wu H, et al. The long noncoding RNA LUCAT1 promotes colorectal cancer cell proliferation by antagonizing nucleolin to regulate MYC expression. Cell death Dis. 2020;11:908.
Choi SH, Martinez TF, Kim S, Donaldson C, Shokhirev MN, Saghatelian A, et al. CDK12 phosphorylates 4E-BP1 to enable mTORC1-dependent translation and mitotic genome stability. Genes &. development 2019;33:418–35.
Lee HC, Wang H, Baladandayuthapani V, Lin H, He J, Jones RJ, et al. RNA polymerase I inhibition with CX-5461 as a novel therapeutic strategy to target MYC in multiple myeloma. Br J Haematol. 2017;177:80–94.
Cantor JR, Sabatini DM. Cancer cell metabolism: one hallmark, many faces. Cancer Discov. 2012;2:881–98.
Hung CL, Wang LY, Yu YL, Chen HW, Srivastava S, Petrovics G, et al. A long noncoding RNA connects c-Myc to tumor metabolism. Proc Natl Acad Sci USA. 2014;111:18697–702.
Sun Y, Wei G, Luo H, Wu W, Skogerbo G, Luo J, et al. The long noncoding RNA SNHG1 promotes tumor growth through regulating transcription of both local and distal genes. Oncogene. 2017;36:6774–83.
Lu Q, Shan S, Li Y, Zhu D, ** W, Ren T. Long noncoding RNA SNHG1 promotes non-small cell lung cancer progression by up-regulating MTDH via sponging miR-145-5p. FASEB J : Off Publ Fed Am Soc Exp Biol. 2018;32:3957–67.
Damas ND, Marcatti M, Come C, Christensen LL, Nielsen MM, Baumgartner R, et al. SNHG5 promotes colorectal cancer cell survival by counteracting STAU1-mediated mRNA destabilization. Nat Commun. 2016;7:13875.
Chen Z, Chen X, Chen P, Yu S, Nie F, Lu B, et al. Long non-coding RNA SNHG20 promotes non-small cell lung cancer cell proliferation and migration by epigenetically silencing of P21 expression. Cell death Dis. 2017;8:e3092.
Wang Y, Fu J, Yang L, Liang Z. Long noncoding RNA SNHG20 promotes colorectal cancer cell proliferation, migration and invasion via miR495/STAT3 axis. Mol Med Rep. 2021;23:31.
Jiang Q, Wang Z, Qi Q, Li J, **n Y, Qiu J. lncRNA SNHG26 promoted the growth, metastasis, and cisplatin resistance of tongue squamous cell carcinoma through PGK1/Akt/mTOR signal pathway. Mol Ther oncolytics. 2022;24:355–70.
Abdelmohsen K, Tominaga K, Lee EK, Srikantan S, Kang MJ, Kim MM, et al. Enhanced translation by nucleolin via G-rich elements in coding and non-coding regions of target mRNAs. Nucleic acids Res. 2011;39:8513–30.
Xu C, Wang Y, Tu Q, Zhang Z, Chen M, Mwangi J, et al. Targeting surface nucleolin induces autophagy-dependent cell death in pancreatic cancer via AMPK activation. Oncogene. 2019;38:1832–44.
Pichiorri F, Palmieri D, De Luca L, Consiglio J, You J, Rocci A, et al. In vivo NCL targeting affects breast cancer aggressiveness through miRNA regulation. J Exp Med. 2013;210:951–68.
Liu H, Luo J, Luan S, He C, Li Z. Long non-coding RNAs involved in cancer metabolic reprogramming. Cell Mol Life Sci: CMLS. 2019;76:495–504.
Guo T, Bai YH, Cheng XJ, Han HB, Du H, Hu Y, et al. Insulin gene enhancer protein 1 mediates glycolysis and tumorigenesis of gastric cancer through regulating glucose transporter 4. Cancer Commun. 2021;41:258–72.
Liu Y, Huang Y, Zhang J, Pei C, Hu J, Lyu J, et al. TIMMDC1 knockdown inhibits growth and metastasis of gastric cancer cells through metabolic inhibition and AKT/GSK3beta/beta-catenin signaling pathway. Int J Biol Sci. 2018;14:1256–67.
Acknowledgements
We would like to take our special thanks to Pro. He **anghuo and Huang Shenglin from Fudan University Shanghai Cancer Center and Institutes of Biomedical Sciences, for their kindness help and constructive advice in our study. We are also like to thank Pro. Zhang Honghe from Zhejiang University for his gift of the NCL truncated plasmids and Pro. Lan Fei from Institutes of Biomedical Sciences, Fudan University for the gift of pRF and pRmF reporter plasmids.
Funding
This work was supported by grants from the National Natural Science Foundation of China (82002542, 82173296).
Author information
Authors and Affiliations
Contributions
WZ-H and WY-X contributed experiments and writing; SJ-J contributed formal analysis; ZL-Q contributed data curation; ZY-J contributed validation; LY-F contributed methodology; GW-J contributed writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The Medical Ethics Committee of Fudan University and the Affiliated Cancer Hospital of Fudan University approved the collection of clinical materials for research purposes. All samples were collected and analyzed after obtaining written informed consent from each patient. All animal experiments adhered to the Principles of Care and Use of Laboratory Animals of Fudan University and were approved by the Subcommittee on the Ethics of Experimental Animal Welfare within the Medical Ethics Committee of Fudan University (approval no. USCC-IACUC-S20210090, approval date: 23 February 2021).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by professor Nickolai Barlev
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, ZH., Wang, YX., Song, JJ. et al. LncRNA SNHG26 promotes gastric cancer progression and metastasis by inducing c-Myc protein translation and an energy metabolism positive feedback loop. Cell Death Dis 15, 236 (2024). https://doi.org/10.1038/s41419-024-06607-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06607-8
- Springer Nature Limited