Abstract
Cancer cells develop multiple strategies to evade T cell-mediated killing. On one hand, cancer cells may preferentially rely on certain amino acids for rapid growth and metastasis. On the other hand, sufficient nutrient availability and uptake are necessary for mounting an effective T cell anti-tumor response in the tumor microenvironment (TME). Here we demonstrate that tumor cells outcompete T cells for cystine uptake due to high Slc7a11 expression. This competition induces T-cell exhaustion and ferroptosis, characterized by diminished memory formation and cytokine secretion, increased PD-1 and TIM-3 expression, as well as intracellular oxidative stress and lipid-peroxide accumulation. Importantly, either Slc7a11 deletion in tumor cells or intratumoral cystine supplementation improves T cell anti-tumor immunity. Mechanistically, cystine deprivation in T cells disrupts glutathione synthesis, but promotes CD36 mediated lipid uptake due to dysregulated cystine/glutamate exchange. Moreover, enforced expression of glutamate-cysteine ligase catalytic subunit (Gclc) promotes glutathione synthesis and prevents CD36 upregulation, thus boosting T cell anti-tumor immunity. Our findings reveal cystine as an intracellular metabolic checkpoint that orchestrates T-cell survival and differentiation, and highlight Gclc as a potential therapeutic target for enhancing T cell anti-tumor function.

Similar content being viewed by others
Introduction
Adoptive T-cell transfer and immune checkpoint blockade have achieved durable clinical outcomes in a certain fraction of cancer patients [1, 2], but the majority of patients do not benefit from those treatments due to the highly immunosuppressive tumor microenvironment (TME), which poses a major challenge for effective immunotherapies against solid tumors [3,4,5]. In the TME, multiple factors contribute to immune suppression, including low oxygen and pH levels, limited nutrient availability (e.g., glucose, fatty acids, and amino acids), accumulation of immunosuppressive metabolites (e.g., ROS, lactate, and lipids), and increased secretion of immunosuppressive cytokines as well as chemokines [45], our findings suggested that cystine starvation severely impairs T cell anti-tumor immunity. Hence, anti-tumor therapies in terms of cystine depletion should be treated with caution. Notably, different tumor types exhibit varying response to cystine deprivation. For instance, tumor cells with high Nrf2 expression are more resistant to ferroptosis induced by cystine depletion [33]. In such cases, cystine supplementation may enhance immune response and improve therapeutic efficacy. Therefore, future studies are needed to precisely identify the appropriate tumor types for cystine depletion or supplementation therapies. Additionally, our study emphasizes the impact of cystine transport on intracellular lipid accumulation in T cells. CD36 is the key factor inducing T-cell ferroptosis via excessive uptake of lipid peroxides [40, 41]. Our data suggested that cystine/glutamine exchange dysfunction exacerbates CD36-mediated lipid accumulation, consequently inducing T-cell ferroptosis. These findings provide important insights into rational design of future immunotherapy strategies.
The strategies targeting cystine metabolism to enhance T-cell immunity have attracted increasing attention. Several studies have reported that T cells overexpressing either Slc7a11 or cystathionine-gamma-lyase, the key enzyme converting methionine to cysteine, exhibit enhanced tumor suppression [17, 46]. Our study presents a strategy for T cells to counteract cystine deprivation. Gclc overexpression not only counters oxidative stress induced by cystine deficiency, but also mitigates the damage caused by cystine/glutamine exchange dysfunction, thereby enhancing T cell anti-tumor effects. This was in line with previous studies that heightened Gclc expression improves redox balance and prolongs mammalian cell lifespans [47,48,49,50]. Conversely, Gclc ablation in T cells severely impairs inflammatory responses [21]. Furthermore, overexpression of Nrf2, a key regulator of Gclc, significantly enhances T cell anti-tumor functionality [51]. Collectively, these findings highlight Gclc as a potential therapeutic target for boosting T cell anti-tumor immunity.
Nevertheless, this study has certain limitations. While the decrease of cystine in the TME has been demonstrated, the specific concentration of cystine may vary across different tumor types [17, 23]. Besides, plasma amino acid levels undergo dynamic changes as the tumor progress [52], implying that amino acid abundance in the TME may be also influenced by tumor development. Thus, the in vitro setting used in this study to investigate the impacts of cystine deprivation on T cells may not fully capture the dynamic and intricate TME. The variations in cystine content across different tumor types or stages should also be taken into account in future studies. Furthermore, while analysis from specific human cancer datasets supports our findings, the impact of cystine deprivation on T cells in human cancers requires further investigations.
In conclusion, our results reveal the impaired T cell-immunity due to cystine competition by tumor cells. We elucidate a novel mechanism of ferroptosis in tumor-infiltrating T cells: cystine deprivation leads to glutamate accumulation, subsequently exacerbating CD36-mediated lipid peroxides production. These findings provide a theoretical foundation for immunotherapeutic strategies targeting amino acid metabolism or CD36 blockade. Furthermore, we prove that elevated Gclc expression protects T cells from cystine deprivation induced damage, presenting a potential strategy for engineering T cells for effective anti-tumor immunotherapy.
Materials and methods
Cell lines
HEK-293FT cells (#CRL-3249, ATCC, USA) and MC38 cells (#ENH204-FP, Kerafast, USA) were cultured in DMEM (#C11995500CP, Gibco, USA) containing 10% FBS (#SH30396, Cytia, USA), and penicillin-streptomycin (#SV30010, Hyclone, USA) at 37 °C with 5% CO2. B16F10-OVA cells were kindly provided by Professor Bo Huang (Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Bei**g, China). B16F10 cells (#CRL-6475, ATCC, USA) and B16F10-OVA cells were cultured in RPMI (#11875-093, Gibco, USA) containing 10% FBS and penicillin-streptomycin at 37 °C with 5% CO2. Cell lines were regularly checked for mycoplasma contamination.
Mice
The animal protocol was approved by the Institutional Animal Care and Use Committee (IACUC) of Suzhou Institute of Systems Medicine (ISM-IACUC-0018 and ISM-IACUC-0055). We confirm that all experiments were conducted in accordance with the relevant regulations of the committee. CD45.1+ OT-1 TCR transgenic mice (C57BL/6 N background) and CD45.2+ female C57BL/6 N mice (6–8 weeks old, WT) were purchased from Vital River Company (Bei**g, China) and housed under specific pathogen-free conditions at Suzhou Institute of Systems Medicine.
Quantification of cystine concentration
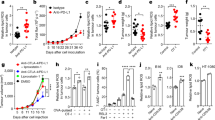
Tumor interstitial fluid was isolated from B16F10 tumors as previously described [23]. Serum was obtained by centrifuging blood at 3500 rpm for 5 min and collecting the supernatant. Cystine concentration was quantified by liquid chromatography-tandem mass spectrometry (LC-MS). LC-MS analysis was conducted at the APTBIO Biotechnology Inc (Shanghai, China).
CD8+ T-cell isolation and activation
OT-1 mouse spleens were processed into single-cell suspensions through 40 μm filters. The splenocytes were incubated in 2 mL of red blood cell lysis buffer (#420301, Biolegend, USA) for 10 min. Subsequently, splenocytes were washed by PBS and resuspended at 1 × 106/mL in T-cell medium (formula listed below) containing OVA peptide (1 μg/mL, #SP1050a, Abcepta Biotech, USA) and IL-2 (10 ng/mL, #200-02-500, Pepro Tech, USA) to activate CD8+ T cells. After 3 days, dead cells were removed by Ficoll-Paque separation (#GE17-1440-03, Sigma-Aldrich, USA), and alive CD8+ T cells were collected for subsequent experiments.
Formula of T-cell medium: The RPMI medium contained 10% FBS (#SH30396, Cytiva, USA), penicillin-streptomycin (1:100, #SH40003, Cytiva, USA), Hepes (10 mmol/L, #03-025-1B, Biological Industries, Israel), Sodium pyruvate (1 mmol/L, #11360-070, Gibco, USA), MEM non-essential amino acids (1:100, #11140-050, Gibco, USA), L-glutamine (2 mmol/L, #25030081, Gibco, USA), and β-mercaptoethanol (5 μmol/L, #B6891, Sigma, USA).
Cystine deprivation in CD8+ T cells
The RPMI medium lacking cystine, methionine, and glutamine was customized from Duoning Biological Company (Shanghai, China). Methionine (100 μmol/L, #M9625, Sigma, USA), cystine hydrochloride (#C6727, Sigma, USA), and other reagents according to the T-cell medium formula were added to the customized medium. Activated T cells were cultured in either normal (200 μmol/L) or cystine-deprived (20 μmol/L) medium with IL-2 and IL-7 (10 ng/mL) for 72 h.
Sensitivity of T cells and tumor cells to cystine deprivation
Activated T cells, B16F10 cells, and MC38 cells were cultured separately in T-cell medium with varying concentrations of cystine (0.001, 0.01, 0.1, 1, 10, 20, 40, 70, 100, 150, and 200 μmol/L). After 48 h of incubation, the CCK8 reagent (1:100, #K1018, APE-BIO, USA) was added to measure the cell viability.
T-cell cultivation in tumor supernatant
B16F10 and MC38 cells were cultured for 48 h in T-cell medium with cystine concentrations of 200, 50, and 30 μmol/L, and the medium (tumor supernatant) was collected. Subsequently, activated T cells were cultured for 24 h in either fresh medium or tumor supernatants with 200, 50, and 30 μmol/L cystine. The percentage of dead cells was assessed using flow cytometry.
Tumor and T cell co-culture
B16F10 and MC38 tumor cells were planted in 24-well plates (106 cells per well). After cell adhesion, the medium was replaced with either normal medium or 20 μmol/L cystine-deprived medium. Subsequently, 106 activated T cells were added to the upper chamber of transwell inserts, which were then placed into the 24-well plate for co-culture with tumor cells (Supplementary Fig. 1g). After 24 h, T cells from the upper chambers were collected, and cell viability was assessed using the CCK-8 assay.
Cell death pathway inhibition
Activated T cells were individually cultured with 10 μmol/L cystine and the following regents: ferrostatin-1 (10 μmol/L, #HY-100579, MCE, USA), N-Acetyl Cysteine (10 mmol/L, HY-B0215, MCE, USA), Z-VAD-FMK (10 μmol/L, #HY-16658B, MCE, USA), necrostatin-1 (10 μmol/L, #HY-15760, MCE, USA), MCC950 (10 μmol/L, #HY-12815, MCE, USA). Additionally, T cells were cultured in normal medium as control. After 48 h, T-cell viability was detected by CCK-8 and lipid peroxidation was detected by flow cytometry assay.
CD36 blockade of T cells
Activated T cells were cultured in normal medium, cystine-deprived medium (20 μmol/L), and cystine-deprived medium supplemented with CD36 antibody (10 μg/mL, #163002, Biolegend, USA) for 48 h. Subsequently, T cells were collected for flow cytometry analysis.
Glutamine supplementation and RSL3 treatment
For glutamine supplementation treatment, activated T cells were cultured in normal medium, normal medium with 20 mmol/L glutamine (#25030081, Gibco, USA), cystine-deprived medium, and cystine-deprived medium with 20 mmol/L glutamine for 48 h. For RSL3 treatment, activated T cells were cultured in normal medium, normal medium with 20 mmol/L glutamine, normal medium containing RSL3 (10 nmol/L, #HY-100218A, MCE, USA), and normal medium containing RSL3 and 20 mmol/L glutamine for 48 h.
Intratumoral cystine supplementation assay
C57BL/6 N mice were subcutaneously inoculated with 5 × 105 B16F10 cells, because B16F10 tumor was loose and soft, allowing effective intratumoral injection. On day 10 after tumor inoculation, 0.1 mg cystine hydrochloride in 100 μL PBS or PBS alone was intratumoral administered to each mouse every other day and the tumor size was measured. Tumor volume was calculated using the formula V = (L × W2) / 2, V is tumor volume, L is tumor length, and W is tumor width. Tumor size did not exceed 2 cm in any dimension in any of the experiments.
WT and Slc7a11-KD tumor inoculation
To generate pLKO.1-Slc7a11shRNA plasmid, forward oligonucleotide (5′-CCGGGCCCTGTCCTATGCAGAATTACTCGAGTAATTCTGCATAGGACAGGGCTTTTTG-3′) was annealed with reverse oligonucleotide (5′-AATTCAAAAAGCCCTGTCCTATGCAGAATTACTCGAGTAATTCTGCATAGGACAGGGC-3′), and then subcloned into an AgeI (#R0552S, NEB, USA)/EcoRI (#R0101S, NEB, USA) digested pLKO.1-TRC cloning vector (#10879, Addgene, USA). To produce pLKO.1 Lentiviral particles, 293FT cells were co-transfected in a 4:3:1 ratio with pLKO.1-shRNA (or pLKO.1-Slc7a11shRNA), psPAX2, and pMD2.G. The supernatant containing the viral particles was collected 48 and 72 h post-transfection. B16F10 cells were infected with the virus supernatant for 48 h, followed by an additional 7-day selection with 500 μg/mL G418 (#10131035, Invitrogen, USA). Subsequently, C57BL/6 N mice were randomly inoculated with either 5 × 105 WT or Slc7a11-KD B16F10 cells, and tumor volumes were measured since the 10th day post-inoculation. For CD8 depletion assay, 200 μg CD8 antibody was intraperitoneally injected into each mouse three days before tumor inoculation and on days 1, 7, and 12 post-inoculation (Supplementary Fig. 3j).
Gclc-overexpressing T cell generation and adoptive T-cell therapy
The Gclc sequence was amplificated (Forward primer: 5′-GGGTGGACCATCCTCTAGCCCTCGAGATGGGGCTGCTGTCCCAAG-3′; Reverse primer: 5′-GCTTCCGGCTAGCCCTGCGCAAGCTTCCGGCTGAAGGGTCGCTTTTACCTC-3′) and inserted into XhoI (#R0146L, NEB, USA) and HindIII (#R3104S, NEB, USA) digested pMSGV-Thy1.1 plasmid. pCL-Eco (#HG-VNC0832, Clontech, USA) and pMSGV-Thy1.1 were 1:1 co-transfected into 293FT cells. The virus-containting supernatant was collected at 48 and 72 h post-transfection. Naïve CD8+ T cells were sorted from mouse spleens using the CD8+ naïve T-cell enrichment kit (#480044, Biolegend, USA), and were planted into αCD3 (1 μg/mL, #16-0031-86, Invitrogen, USA) and αCD28(1 μg/mL, #16-0281-85, Invitrogen, USA) precoated plate. On the 3rd day, activated T cells were planted into RetroNectin (0.1 mg/mL, #T100A, TaKaRa, Japan) precoated 24-well plates, culturing with a 1:1 mixture of virus supernatant and T-cell medium containing polybrene (1 μg/mL, #SC-134220, Santa Cruz, USA) and IL-2 (10 ng/mL). After 24 h, the infected T cells were cultured in T-cell medium. After 48 h, the frequency of Thy1.1+ T cells was detected as infection efficiency.
For adoptive T-cell therapy, 5 × 105 B16-OVA tumor cells were subcutaneously injected into CD45.2+ C57BL/6 mice. Mice were 5 Gy irradiated when the tumor diameter was approximately 4–5 mm. Subsequently, vector or Gclc-OE T cells were intravenously transferred into recipients (1 × 106 cells per mouse). Tumor size was measured every 3 days.
Flow cytometry analysis
Spleens, lymph nodes, and tumor tissues were processed into single-cell suspensions using 40μm filters. Lymphocytes from tumor tissues were enriched using Percoll density gradient media (#17089109, Cytiva, Sweden). All samples were stained with Fixable Viability Dye eFluor™ 506 (#65-0866-18, Invitrogen, USA) on ice for 20 min. For cell surface staining, antibodies (listed in Supplementary Table 1) were diluted to 1:200 in FACS buffer (PBS with 2% FBS) and incubated on ice for 25 min. For intracellular cytokine staining, cells were incubated with Brefeldin A (1:1000, #00-4506-51, Invitrogen, USA), Monensin (1:1000, #00-4505-51, Invitrogen, USA), PMA (10 ng/mL, #P8139, Sigma, USA) and ionomycin (500 ng/mL, #FMS-FZ208, Fcmacs, Nan**g, China) at 37 °C for 3.5 h. The cells were then fixed on ice with Fixation Buffer (#420801, Biolegend, USA) for 20 min, and stained with cytokine antibodies (listed in Supplementary Table 1) in permeabilization buffer (#421002, Biolegend, USA). For intracellular transcription factor staining, cells were fixed with Foxp3/Transcription Factor Staining Buffer (#00-5223-56, #00-5123-43, Invitrogen, USA), and then stained with transcription factor antibodies (listed in Supplementary Table 1) in Foxp3/Transcription Factor Permeabilization Buffer (#00-8333-56, Invitrogen, USA). All samples were resuspended in FACS buffer, loaded in an LSR Fortessa flow cytometer (Becton-Dickinson, San Jose, CA) and analyzed using FlowJo software.
oxLDL uptake and lipid peroxidation assay
To measure oxLDL uptake, cells were incubated at 37 °C in PBS containing oxLDL-DyLight-488 (1:1000, #601181, Cayman, USA) for 20 min. To detect lipid peroxidation, cells were incubated at 37 °C in PBS containing BODIPY FLC11 (1.5 μmol/L, #D3861, Invitrogen, USA) for 20 min. After incubation, the cells were washed with FACS buffer for cell surface staining.
ROS and glutathione measurement
For ROS measurement, T cells were incubated at 37 °C for 20 min in PBS containing DCFHDA (1:1000, #S0033S, Beyotime, China) for cellular ROS detection or with Mito-SOX (1:200, #M36008, Invitrogen, USA) for mitochondrial ROS detection. After incubation, T cells were resuspended in FACS buffer for cell surface staining. For glutathione measurement, 106 alive T cells were collected for each group, and the total glutathione levels were determined according to the protocol of the GSH and GSSG assay kit (#S0053, Beyotime, China).
RNA sequencing
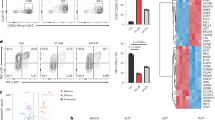
Total RNA from CD8+ T cells were extracted using the RNeasy Mini Kit (#74104, Qiagen, Germany). Three biological replicates were performed for each studied condition. RNA sequencing was conducted and the data were processed by the bioinformatic core at the Suzhou Institute of Systems Medicine. Source data were published on the Figshare website, Doi: 10.6084 / m9 Figshare. 24925728. Gene sets related to lipid peroxidation/ferroptosis activation, T-cell memory/effector differentiation, and T-cell dysfunction/exhaustion were selected from existing publications [53,54,55]. Heatmap analysis were performed by Hiplot Pro (https://hiplot.com.cn/), a comprehensive web service for biomedical data analysis and visualization.
Quantitative lipidomics assay
T cells were cultured in NM or CD for 3 days, and the alive cells were collected for lipidomics assay. LC-MS analysis was conducted at the Metware Biotechnology Inc (Wuhan, China). For the detection of free fatty acids, the samples were homogenized and mixed with an internal standard mixture in ice-cold methanol. The extracted free fatty acids were then converted to acyl chloride intermediates and detected using previously described methods [40].
Real-time quantitative PCR and western blot assay
CD8+ T cells from spleens or tumor tissues were sorted using PE-anti-CD45.1 antibody (#110708, Biolegend, USA) and anti-PE magnetic beads (#480080, Biolegend, USA), or T cells from in vitro culture were collected for RT-qPCR. Total RNA was isolated using the RNeasy Mini Kit, and subjected to reverse transcription with PrimeScript RT Master Mix Kit (#RR036B, TaKaRa, Japan). Reactions were conducted on the LightCycler 480 system (Roche, Switzerland) using Perfectstart Green qP CR SuperMix (#AQ601-04, TransGen Bitotech, China). Specific primers are listed in Supplementary Table 2.
For western blot assay, cells were lysed using RIPA buffer (#P0013B, Beyotime, China). Protein was quantified by BCA protein quantification kit (#P0010S, Beyotime, China) and denatured at 95 °C for 10 min. Protein was separated via 10% SDS-polyacrylamide gel and then transferred to PVDF membranes. The membranes were blocked in 5% non-fat milk for 1 h and then incubated with specified primary antibodies (listed in Supplementary Table 1) at 4 °C overnight. After incubation with HRP-secondary antibody, target protein was visualized by the chemiluminescent detection kit (#180–501, Tanon, China) and a ChemiDoc western imaging system (Bio-rad, USA).
Statistical analysis
All data were analyzed from at least three experiments and presented as mean ± s.e.m. The sample size was not predetermined using a statistical method, it was based on prior experimental observations. Mice were randomly allocated to experimental groups and no blinding method was used for animal experiments. Differences were assessed by two-tailed student’s unpaired t-test among two groups. One-way ANOVA with Turkey’s multiple comparison test was used to examine difference between more than two groups. The variance is similar between the groups undergoing statistical comparisons. p value < 0.05 was considered significant (ns, no significant, *p < 0.05; **p < 0.01; ***p < 0.001). All statistics were performed using GraphPad Prism 9 (GraphPad, San Diego, USA).
Data availability
Data reported in this paper will be shared by the lead contact upon request. The analysis of the correlation between Slc7a11 and immune infiltration in human tumors was performed by the TIMER2.0 online database (http://timer.cistrome.org/). The scRNA sequencing data from melanoma patients were found at the GEO accession number GSE120575.
References
Rosenberg SA, Restifo NP. Adoptive cell transfer as personalized immunotherapy for human cancer. Science. 2015;348:62–8.
Finck AV, Blanchard T, Roselle CP, Golinelli G, June CH. Engineered cellular immunotherapies in cancer and beyond. Nat Med. 2022;28:678–89.
Met Ö, Jensen KM, Chamberlain CA, Donia M, Svane IM. Principles of adoptive T cell therapy in cancer. Semin Immunopathol. 2019;41:49–58.
DePeaux K, Delgoffe GM. Metabolic barriers to cancer immunotherapy. Nat Rev Immunol. 2021;21:785–97.
Kishton RJ, Sukumar M, Restifo NP. Metabolic regulation of T cell longevity and function in tumor immunotherapy. Cell Metab. 2017;26:94–109.
**a L, Oyang L, Lin J, Tan S, Han Y, Wu N, et al. The cancer metabolic reprogramming and immune response. Mol Cancer. 2021;20:28.
Pitt JM, Marabelle A, Eggermont A, Soria J-C, Kroemer G, Zitvogel L. Targeting the tumor microenvironment: removing obstruction to anticancer immune responses and immunotherapy. Ann Oncol. 2016;27:1482–92.
** M-Z, ** W-L. The updated landscape of tumor microenvironment and drug repurposing. Signal Transduct Target Ther. 2020;5:166.
Chen Z, Ji Z, Ngiow SF, Manne S, Cai Z, Huang AC, et al. TCF-1-centered transcriptional network drives an effector versus exhausted CD8 T cell-fate decision. Immunity. 2019;51:840–55.e5.
McLane LM, Abdel-Hakeem MS, Wherry EJ. CD8 T cell exhaustion during chronic viral infection and cancer. Annu Rev Immunol. 2019;37:457–95.
Sckisel GD, Mirsoian A, Minnar CM, Crittenden M, Curti B, Chen JQ, et al. Differential phenotypes of memory CD4 and CD8 T cells in the spleen and peripheral tissues following immunostimulatory therapy. J Immunother Cancer. 2017;5:33.
Han C, Ge M, Ho P-C, Zhang L. Fueling T-cell antitumor immunity: amino acid metabolism revisited. Cancer Immunol Res. 2021;9:1373–82.
Wang W, Zou W. Amino acids and their transporters in T cell immunity and cancer therapy. Mol Cell. 2020;80:384–95.
Yang L, Chu Z, Liu M, Zou Q, Li J, Liu Q, et al. Amino acid metabolism in immune cells: essential regulators of the effector functions, and promising opportunities to enhance cancer immunotherapy. J Hematol Oncol. 2023;16:59.
Srivastava MK, Sinha P, Clements VK, Rodriguez P, Ostrand-Rosenberg S. Myeloid-derived suppressor cells inhibit T-cell activation by depleting cystine and cysteine. Cancer Res. 2010;70:68–77.
Yan Z, Garg SK, Banerjee R. Regulatory T cells interfere with glutathione metabolism in dendritic cells and T cells. J Biol Chem. 2010;285:41525–32.
Lancien M, Gueno L, Salle S, Merieau E, Beriou G, Nguyen TH, et al. Cystathionine-gamma-lyase overexpression in T cells enhances antitumor effect independently of cysteine autonomy. Cancer Sci. 2021;112:1723–34.
Angelini G, Gardella S, Ardy M, Ciriolo MR, Filomeni G, Di Trapani G, et al. Antigen-presenting dendritic cells provide the reducing extracellular microenvironment required for T lymphocyte activation. Proc Natl Acad Sci USA. 2002;99:1491–6.
Levring TB, Hansen AK, Nielsen BL, Kongsbak M, von Essen MR, Woetmann A, et al. Activated human CD4+ T cells express transporters for both cysteine and cystine. Sci Rep. 2012;2:266.
Castellano F, Molinier-Frenkel V. Control of T-cell activation and signaling by amino-acid catabolizing enzymes. Front Cell Dev Biol. 2020;8:613416.
Mak TW, Grusdat M, Duncan GS, Dostert C, Nonnenmacher Y, Cox M, et al. Glutathione primes T cell metabolism for inflammation. Immunity. 2017;46:675–89.
Levring TB, Kongsbak M, Rode AKO, Woetmann A, Ødum N, Bonefeld CM, et al. Human CD4+ T cells require exogenous cystine for glutathione and DNA synthesis. Oncotarget. 2015;6:21853–64.
Sullivan MR, Danai LV, Lewis CA, Chan SH, Gui DY, Kunchok T, et al. Quantification of microenvironmental metabolites in murine cancers reveals determinants of tumor nutrient availability. Elife. 2019;8:e44235.
Robert SM, Buckingham SC, Campbell SL, Robel S, Holt KT, Ogunrinu-Babarinde T, et al. SLC7A11 expression is associated with seizures and predicts poor survival in patients with malignant glioma. Sci Transl Med. 2015;7:289ra86.
Shi Z, Naowarojna N, Pan Z, Zou Y. Multifaceted mechanisms mediating cystine starvation-induced ferroptosis. Nat Commun. 2021;12:4792.
Koppula P, Zhuang L, Gan B. Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell. 2021;12:599–620.
Doll S, Freitas FP, Shah R, Aldrovandi M, da Silva MC, Ingold I, et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature. 2019;575:693–8.
Bersuker K, Hendricks JM, Li Z, Magtanong L, Ford B, Tang PH, et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature. 2019;575:688–92.
Zhang Y, Swanda RV, Nie L, Liu X, Wang C, Lee H, et al. mTORC1 couples cyst(e)ine availability with GPX4 protein synthesis and ferroptosis regulation. Nat Commun. 2021;12:1589.
Gao M, Yi J, Zhu J, Minikes AM, Monian P, Thompson CB, et al. Role of mitochondria in ferroptosis. Mol Cell. 2019;73:354–63.e3.
Gao M, Monian P, Quadri N, Ramasamy R, Jiang X. Glutaminolysis and transferrin regulate ferroptosis. Mol Cell. 2015;59:298–308.
Fan G, Liu M, Liu J, Huang Y. The initiator of neuroexcitotoxicity and ferroptosis in ischemic stroke: glutamate accumulation. Front Mol Neurosci. 2023;16:1113081.
Kang YP, Mockabee-Macias A, Jiang C, Falzone A, Prieto-Farigua N, Stone E, et al. Non-canonical glutamate-cysteine ligase activity protects against ferroptosis. Cell Metab. 2021;33:174–89.e7.
Zhang L, Romero P. Metabolic control of CD8+ T cell fate decisions and antitumor immunity. Trends Mol Med. 2018;24:30–48.
Miotto G, Rossetto M, Di Paolo ML, Orian L, Venerando R, Roveri A, et al. Insight into the mechanism of ferroptosis inhibition by ferrostatin-1. Redox Biol. 2020;28:101328.
Cowburn AS, White JF, Deighton J, Walmsley SR, Chilvers ER. z-VAD-fmk augmentation of TNF alpha-stimulated neutrophil apoptosis is compound specific and does not involve the generation of reactive oxygen species. Blood. 2005;105:2970–2.
Cao L, Mu W. Necrostatin-1 and necroptosis inhibition: pathophysiology and therapeutic implications. Pharm Res. 2021;163:105297.
Corcoran SE, Halai R, Cooper MA. Pharmacological inhibition of the nod-like receptor family pyrin domain containing 3 inflammasome with MCC950. Pharm Rev. 2021;73:968–1000.
Stockwell BR, Jiang X, Gu W. Emerging mechanisms and disease relevance of ferroptosis. Trends Cell Biol. 2020;30:478–90.
Ma X, **ao L, Liu L, Ye L, Su P, Bi E, et al. CD36-mediated ferroptosis dampens intratumoral CD8+ T cell effector function and impairs their antitumor ability. Cell Metab. 2021;33:1001–12.e5.
Xu S, Chaudhary O, Rodríguez-Morales P, Sun X, Chen D, Zappasodi R, et al. Uptake of oxidized lipids by the scavenger receptor CD36 promotes lipid peroxidation and dysfunction in CD8+ T cells in tumors. Immunity. 2021;54:1561–77.e7.
Li S, Lu Z, Sun R, Guo S, Gao F, Cao B, et al. The role of SLC7A11 in cancer: friend or foe? Cancers. 2022;14:3059.
Hope HC, Salmond RJ. The role of non-essential amino acids in T cell function and anti-tumour immunity. Arch Immunol Ther Exp. 2021;69:29.
Zhang J, Pavlova NN, Thompson CB. Cancer cell metabolism: the essential role of the nonessential amino acid, glutamine. EMBO J. 2017;36:1302–15.
Badgley MA, Kremer DM, Maurer HC, DelGiorno KE, Lee H-J, Purohit V, et al. Cysteine depletion induces pancreatic tumor ferroptosis in mice. Science. 2020;368:85–9.
Panetti S, McJannett N, Fultang L, Booth S, Gneo L, Scarpa U, et al. Engineering amino acid uptake or catabolism promotes CAR T-cell adaption to the tumor environment. Blood Adv. 2023;7:1754–61.
Moskalev A, Shaposhnikov M, Proshkina E, Belyi A, Fedintsev A, Zhikrivetskaya S, et al. The influence of pro-longevity gene Gclc overexpression on the age-dependent changes in Drosophila transcriptome and biological functions. BMC Genom. 2016;17:1046.
Zhang Z, Kuang Y, Ma K, Li Y, Liu X, Shi Y, et al. Gclc overexpression inhibits apoptosis of bone marrow mesenchymal stem cells through the PI3K/AKT/Foxo1 pathway to alleviate inflammation in acute lung injury. Int Immunopharmacol. 2022;110:109017.
Tran POT, Parker SM, LeRoy E, Franklin CC, Kavanagh TJ, Zhang T, et al. Adenoviral overexpression of the glutamylcysteine ligase catalytic subunit protects pancreatic islets against oxidative stress*. J Biol Chem. 2004;279:53988–93.
Cortes-Wanstreet MM, Giedzinski E, Limoli CL, Luderer U. Overexpression of glutamate–cysteine ligase protects human COV434 granulosa tumour cells against oxidative and γ-radiation-induced cell death. Mutagenesis. 2009;24:211–24.
Gnanaprakasam JNR, Kushwaha B, Liu L, Chen X, Kang S, Wang T, et al. Asparagine restriction enhances CD8+ T cell metabolic fitness and antitumoral functionality through an NRF2-dependent stress response. Nat Metab. 2023;5:1423–39.
Sosnowska A, Chlebowska-Tuz J, Matryba P, Pilch Z, Greig A, Wolny A, et al. Inhibition of arginase modulates T-cell response in the tumor microenvironment of lung carcinoma. Oncoimmunology. 2021;10:1956143.
Galletti G, De Simone G, Mazza EMC, Puccio S, Mezzanotte C, Bi TM, et al. Two subsets of stem-like CD8+ memory T cell progenitors with distinct fate commitments in humans. Nat Immunol. 2020;21:1552–62.
**ao L, Ma X, Ye L, Su P, **ong W, Bi E, et al. IL-9/STAT3/fatty acid oxidation-mediated lipid peroxidation contributes to Tc9 cell longevity and enhanced antitumor activity. J Clin Invest. 2022;132:e153247.
Pilipow K, Scamardella E, Puccio S, Gautam S, De Paoli F, Mazza EM, et al. Antioxidant metabolism regulates CD8+ T memory stem cell formation and antitumor immunity. JCI Insight. 2018;3:e122299.
Sade-Feldman M, Yizhak K, Bjorgaard SL, Ray JP, de Boer CG, et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell. 2018;175:998–1013.e20.
Acknowledgements
Lianjun Zhang was in part supported by the National Key R&D Program of China (2022YFA0807300), the National Natural Science Foundation of China (82350114 and 82271775), the Natural Science Foundation Outstanding Youth Fund of Jiangsu Province (BK20220049), the Suzhou Municipal Key Laboratory (SZS2023005) and the CAMS Innovation Fund for Medical Sciences (CIFMS 2021-I2M-1-061 and 2022-I2M-2-004). Liyuan Zhang was in part supported by the National Natural Science Foundation of China (82171828) and Jiangsu Provincial Key Research and Development Program (BE2021652). We also thank the NCTIB Fund for R&D Platform for Cell and Gene Therapy.
Author information
Authors and Affiliations
Contributions
LZ and CH conceived the experiments, LZ designed the experiments; CH and MG performed most of the experiments. CZ and SL performed bioinformatics analysis of the RNA-seq data. PX, TX, YM, KM, WL and XL helped for the experiments and data collection. BZ and LZ helped for data interpretation and valuable discussions. LZ and CH wrote the manuscript. All authors provided intellectual input to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
The animals used in our study were treated in accordance with protocols approved by the Institutional Animal Care and Use Committee (IACUC) of Suzhou Institute of Systems Medicine.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Hans-Uwe Simon
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, C., Ge, M., **ng, P. et al. Cystine deprivation triggers CD36-mediated ferroptosis and dysfunction of tumor infiltrating CD8+ T cells. Cell Death Dis 15, 145 (2024). https://doi.org/10.1038/s41419-024-06503-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06503-1
- Springer Nature Limited
This article is cited by
-
Establishment of disulfidptosis-related LncRNA signature as biomarkers in colon adenocarcinoma
Cancer Cell International (2024)