Abstract
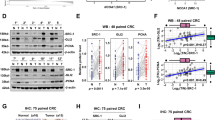
Disabled tumor suppressor genes and hyperactive oncogenes greatly contribute to cell fates during cancer development because of their genetic alterations such as somatic mutations. However, little is known about how tumor suppressor genes react to diverse oncogenes during tumor progression. Our previous study showed that RUNX3 inhibits invasiveness by preventing vascular endothelial growth factor secretion and suppressed endothelial cell growth and tube formation in colorectal cancer (CRC). Hedgehog signaling is crucial for the physiological maintenance and self-renewal of stem cells, and its deregulation is responsible for their tumor development. The mechanisms that inhibit this pathway during proliferation remain poorly understood. Here, we found that the tumor suppressor RUNX3 modulates tumorigenesis in response to cancer cells induced by inhibiting oncogene GLI1 ubiquitination. Moreover, we demonstrated that RUNX3 and GLI1 expression were inversely correlated in CRC cells and tissues. We observed a direct interaction between RUNX3 and GLI1, promoting ubiquitination of GLI1 at the intracellular level. Increased ubiquitination of GLI1 was induced by the E3 ligase β-TrCP. This novel RUNX3-dependent regulatory loop may limit the extent and duration of Hedgehog signaling during extension of the tumor initiation capacity. On the basis of our results, identification of agents that induce RUNX3 may be useful for develo** new and effective therapies for CRC.
Similar content being viewed by others
Introduction
The development of tumor cells often results from multiple genetic alterations that cause a cellular shift. Numerous specific genetic alterations have been identified that activate proto-oncogenes or inactivate tumor suppressor genes. Indeed, the sine qua non of a tumor gene is that it is affected by a mutational event with a vital prevalence. In addition, a ‘third’ pathway to tumorigenesis has been identified whereby the expression of key genes is regulated through promoter hypermethylation and silencing. Particularly, tumor suppressor genes may be subject to this mechanism of inactivation, in addition to mutational events. Oncogenes and tumor suppressor genes have classically been assigned distinct, independent roles in tumor progression. However, the relationships between these tumor-promoting processes remain poorly understood.
The human RUNX gene is homologous to the Drosophila genes Runt and Lozenge [1, 2] and encodes a subunit of the Runt-domain transcription factor PEBP2/CBF [3]. The Runt-related transcription factor (RUNX) family includes RUNX1, RUNX2, and RUNX3, which play a role in cell proliferation and differentiation in humans [4, 5]. RUNX3 has been shown to act as a tumor suppressor in gastric cancer [6], and previous studies have indicated that it is downregulated in various tumors [7]. Moreover, inactivation of RUNX3 is caused by promoter hypermethylation, loss of heterozygosity, or mislocalization [8, 9]. Various studies have demonstrated that RUNX3 can function as a tumor suppressor by regulating metastasis in cancer [44, 45]. The stability of GLI1 via SHh signaling is associated with a malignant phenotype in various cancers. In addition, it has been reported that increased GLI1 enhanced tumor induction in transgenic mice [46]. Unlike SHh and IHh signaling regulated GLI1 through interactions with RUNX2 and RUNX3 in a chondrogenesis mouse model [47].
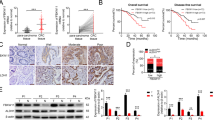
Our study investigated the possibility of using RUNX3 and GLI1 as biomarkers in the patients with CRC. We demonstrated a difference of survival by expression of RUNX3 in early stages such as stages I and II. Typically, patients in these stages are not administered adjuvant chemotherapy after corrective resection of the primary tumor and show a high recurrence rate. Clinical biomarkers that detected poor prognosis patients are urgently needed.
In summary, we showed that Hh-GLI signaling, a major regulator of tumorigenesis, is suppressed by RUNX3. This phenomenon involves the ubiquitin-regulated processing of GLI1, which is mediated by functional cooperation between RUNX3 and the E3 ubiquitin ligase β-TrCP (Fig. 6g). RUNX3 also regulates the transactivation of Notch1 and, consequently, the cell proliferation fate by Notch signaling [48]. Functional crosstalk between the Notch and Hh pathways is known to occur during development and tumorigenesis, but the underlying mechanisms are unclear. Ultimately, considering the roles of Hh signaling in tumorigenesis, appropriate molecular targets should be considered for evaluation in clinical anticancer drug trials focusing on Hh signaling suppression. Our founding improves the understanding of the mechanism by which connecting to RUNX3 as a tumor suppressor and GLI1 as an oncogene occurs as it relates to the metastatic and drug resistance of CRC.
References
Kania MA, Bonner AS, Duffy JB, Gergen JP. The Drosophila segmentation gene runt encodes a novel nuclear regulatory protein that is also expressed in the develo** nervous system. Genes Dev. 1990;4:1701–13.
Daga A, Karlovich CA, Dumstrei K, Banerjee U. Patterning of cells in the Drosophila eye by Lozenge, which shares homologous domains with AML1. Genes Dev. 1996;10:1194–205.
Ito Y. Molecular basis of tissue-specific gene expression mediated by the runt domain transcription factor PEBP2/CBF. Genes Cells. 1999;4:685–96.
Tracey WD, Speck NA. Potential roles for RUNX1 and its orthologs in determining hematopoietic cell fate. Semin Cell Dev Biol. 2000;11:337–42.
Yang S, Wei D, Wang D, Phimphilai M, Krebsbach PH, Franceschi RT. In vitro and in vivo synergistic interactions between the Runx2/Cbfa1 transcription factor and bone morphogenetic protein-2 in stimulating osteoblast differentiation. J Bone Min Res. 2003;18:705–15.
Bae SC, Choi JK. Tumor suppressor activity of RUNX3. Oncogene. 2004;23:4336–40.
Li QL, Ito K, Sakakura C, Fukamachi H, Inoue K, Chi XZ, et al. Causal relationship between the loss of RUNX3 expression and gastric cancer. Cell. 2002;109:113–24.
Ito K, Liu Q, Salto-Tellez M, Yano T, Tada K, Ida H, et al. RUNX3, a novel tumor suppressor, is frequently inactivated in gastric cancer by protein mislocalization. Cancer Res. 2005;65:7743–50.
Yanada M, Yaoi T, Shimada J, Sakakura C, Nishimura M, Ito K, et al. Frequent hemizygous deletion at 1p36 and hypermethylation downregulate RUNX3 expression in human lung cancer cell lines. Oncol Rep. 2005;14:817–22.
Chen F, Wang M, Bai J, Liu Q, ** Y, Li W, et al. Role of RUNX3 in suppressing metastasis and angiogenesis of human prostate cancer. PLoS One. 2014;9:e86917.
He L, Zhao X, Wang H, Zhang P, Guo C, Huang C, et al. RUNX3 mediates suppression of tumor growth and metastasis of human CCRCC by regulating cyclin related proteins and TIMP-1. PLoS One. 2012;7:e32961.
Lobo NA, Shimono Y, Qian D, Clarke MF. The biology of cancer stem cells. Annu Rev Cell Dev Biol. 2007;23:675–99.
O’Brien CA, Kreso A, Jamieson CH. Cancer stem cells and self-renewal. Clin Cancer Res. 2010;16:3113–20.
Liu S, Dontu G, Mantle ID, Patel S, Ahn NS, Jackson KW, et al. Hedgehog signaling and Bmi-1 regulate self-renewal of normal and malignant human mammary stem cells. Cancer Res. 2006;66:6063–71.
Pannuti A, Foreman K, Rizzo P, Osipo C, Golde T, Osborne B, et al. Targeting Notch to target cancer stem cells. Clin Cancer Res. 2010;16:3141–52.
Valkenburg KC, Graveel CR, Zylstra-Diegel CR, Zhong Z, Williams BO. Wnt/beta-catenin signaling in normal and cancer stem cells. Cancers (Basel). 2011;3:2050–79.
Ingham PW, McMahon AP. Hedgehog signaling in animal development: paradigms and principles. Genes Dev. 2001;15:3059–87.
Cochrane CR, Szczepny A, Watkins DN, Cain JE. Hedgehog signaling in the maintenance of cancer stem cells. Cancers (Basel). 2015;7:1554–85.
Ruiz i Altaba A. Hedgehog signaling and the Gli code in stem cells, cancer, and metastases. Sci Signal. 2011;4:pt9.
Yoo YA, Kang MH, Lee HJ, Kim BH, Park JK, Kim HK, et al. Sonic hedgehog pathway promotes metastasis and lymphangiogenesis via activation of Akt, EMT, and MMP-9 pathway in gastric cancer. Cancer Res. 2011;71:7061–70.
Ruiz i Altaba A. Gli proteins and Hedgehog signaling: development and cancer. Trends Genet. 1999;15:418–25.
Aza-Blanc P, Lin HY, Ruiz i Altaba A, Kornberg TB. Expression of the vertebrate Gli proteins in Drosophila reveals a distribution of activator and repressor activities. Development. 2000;127:4293–301.
Ruiz i Altaba A, Sanchez P, Dahmane N. Gli and hedgehog in cancer: tumours, embryos and stem cells. Nat Rev Cancer. 2002;2:361–72.
Di Marcotullio L, Ferretti E, Greco A, De Smaele E, Po A, Sico MA, et al. Numb is a suppressor of Hedgehog signalling and targets Gli1 for Itch-dependent ubiquitination. Nat Cell Biol. 2006;8:1415–23.
Pandolfi S, Stecca B. Cooperative integration between HEDGEHOG-GLI signalling and other oncogenic pathways: implications for cancer therapy. Expert Rev Mol Med. 2015;17:e5.
Ito Y, Bae SC, Chuang LS. The RUNX family: developmental regulators in cancer. Nat Rev Cancer. 2015;15:81–95.
Pratap J, Wixted JJ, Gaur T, Zaidi SK, Dobson J, Gokul KD, et al. Runx2 transcriptional activation of Indian Hedgehog and a downstream bone metastatic pathway in breast cancer cells. Cancer Res. 2008;68:7795–802.
Kim BR, Kang MH, Kim JL, Na YJ, Park SH, Lee SI, et al. RUNX3 inhibits the metastasis and angiogenesis of colorectal cancer. Oncol Rep 2016;36:2601–8.
Fidler IJ. Critical factors in the biology of human cancer metastasis: twenty-eighth G.H.A. Clowes memorial award lecture. Cancer Res. 1990;50:6130–8.
Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331:1559–64.
Visvader JE, Lindeman GJ. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer. 2008;8:755–68.
Valenti G, Quinn HM, Heynen G, Lan L, Holland JD, Vogel R, et al. Cancer stem cells regulate cancer-associated fibroblasts via activation of hedgehog signaling in mammary gland tumors. Cancer Res. 2017;77:2134–47.
Merchant AA, Matsui W. Targeting Hedgehog—a cancer stem cell pathway. Clin Cancer Res. 2010;16:3130–40.
Zhao C, Chen A, Jamieson CH, Fereshteh M, Abrahamsson A, Blum J, et al. Hedgehog signalling is essential for maintenance of cancer stem cells in myeloid leukaemia. Nature. 2009;458:776–9.
Papadopoulos V, Tsapakidis K, Riobo Del Galdo NA, Papandreou CN, Del Galdo F, Anthoney A, et al. The prognostic significance of the hedgehog signaling pathway in colorectal cancer. Clin Colorectal Cancer. 2016;15:116–27.
Varnat F, Duquet A, Malerba M, Zbinden M, Mas C, Gervaz P, et al. Human colon cancer epithelial cells harbour active HEDGEHOG-GLI signalling that is essential for tumour growth, recurrence, metastasis and stem cell survival and expansion. EMBO Mol Med. 2009;1:338–51.
Mazumdar T, DeVecchio J, Shi T, Jones J, Agyeman A, Houghton JA. Hedgehog signaling drives cellular survival in human colon carcinoma cells. Cancer Res. 2011;71:1092–102.
Yoshimoto AN, Bernardazzi C, Carneiro AJ, Elia CC, Martinusso CA, Ventura GM, et al. Hedgehog pathway signaling regulates human colon carcinoma HT-29 epithelial cell line apoptosis and cytokine secretion. PLoS One. 2012;7:e45332.
Chatel G, Ganeff C, Boussif N, Delacroix L, Briquet A, Nolens G, et al. Hedgehog signaling pathway is inactive in colorectal cancer cell lines. Int J Cancer. 2007;121:2622–7.
Gerling M, Buller NV, Kirn LM, Joost S, Frings O, Englert B, et al. Stromal Hedgehog signalling is downregulated in colon cancer and its restoration restrains tumour growth. Nat Commun. 2016;7:12321.
van den Brink GR, Bleuming SA, Hardwick JC, Schepman BL, Offerhaus GJ, Keller JJ, et al. Indian Hedgehog is an antagonist of Wnt signaling in colonic epithelial cell differentiation. Nat Genet. 2004;36:277–82.
Lee JH, Pyon JK, Kim DW, Lee SH, Nam HS, Kang SG, et al. Expression of RUNX3 in skin cancers. Clin Exp Dermatol. 2011;36:769–74.
Salto-Tellez M, Peh BK, Ito K, Tan SH, Chong PY, Han HC, et al. RUNX3 protein is overexpressed in human basal cell carcinomas. Oncogene. 2006;25:7646–9.
Marigo V, Tabin CJ. Regulation of patched by sonic hedgehog in the develo** neural tube. Proc Natl Acad Sci USA. 1996;93:9346–51.
Goodrich LV, Johnson RL, Milenkovic L, McMahon JA, Scott MP. Conservation of the hedgehog/patched signaling pathway from flies to mice: induction of a mouse patched gene by Hedgehog. Genes Dev. 1996;10:301–12.
Nilsson M, Unden AB, Krause D, Malmqwist U, Raza K, Zaphiropoulos PG, et al. Induction of basal cell carcinomas and trichoepitheliomas in mice overexpressing GLI-1. Proc Natl Acad Sci USA. 2000;97:3438–43.
Kim EJ, Cho SW, Shin JO, Lee MJ, Kim KS, Jung HS. Ihh and Runx2/Runx3 signaling interact to coordinate early chondrogenesis: a mouse model. PLoS One. 2013;8:e55296.
Gao J, Chen Y, Wu KC, Liu J, Zhao YQ, Pan YL, et al. RUNX3 directly interacts with intracellular domain of Notch1 and suppresses Notch signaling in hepatocellular carcinoma cells. Exp Cell Res. 2010;316:149–57.
Acknowledgements
Vectors of pCS4-3myc-RUNX3 (full-length) and RUNX3 deletion mutant vectors fused with the Myc tag were kindly provided by Dr. SC Bae at Chungbuk National University. This research is based on data used in Bo Ram Kim’s doctoral dissertation (Korea University).
Funding
This work was supported by a National Research Foundation (NRF) of Korea grant funded by the Korean government (MSIP) [NRF-2017R1A6A3A11030765] and a Korea University Grant.
Author contributions
BRK conceived and designed the study, provided financial support, collected and assembled the data, analyzed and interpreted the data, and wrote the paper. YJN, YAJ, SHP, and MJJ conceived and designed the study, and analyzed and interpreted the data. SHK conceived and designed the study. JLK and SYJ collected and assembled the data, and analyzed and interpreted the data. SCO and DHL conceived and designed the study, provided financial support, collected and assembled the data, analyzed and interpreted the data, wrote the paper, and provided final approval of paper. All authors discussed the results and commented on the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Del Sal
Rights and permissions
About this article
Cite this article
Kim, B.R., Na, Y.J., Kim, J.L. et al. RUNX3 suppresses metastasis and stemness by inhibiting Hedgehog signaling in colorectal cancer. Cell Death Differ 27, 676–694 (2020). https://doi.org/10.1038/s41418-019-0379-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0379-5
- Springer Nature Limited
This article is cited by
-
RUNX3 pathway signature predicts clinical benefits of immune checkpoint inhibition plus tyrosine kinase inhibition in advanced renal cell carcinoma
BMC Urology (2024)
-
RUNX transcription factors: biological functions and implications in cancer
Clinical and Experimental Medicine (2024)
-
Regulation and signaling pathways in cancer stem cells: implications for targeted therapy for cancer
Molecular Cancer (2023)
-
The involvement of E3 ubiquitin ligases in the development and progression of colorectal cancer
Cell Death Discovery (2023)
-
RUNX3 inactivates oncogenic MYC through disruption of MYC/MAX complex and subsequent recruitment of GSK3β-FBXW7 cascade
Communications Biology (2023)