Abstract
Background
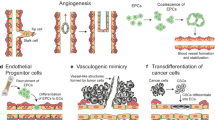
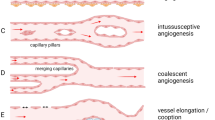
The eukaryotic elongation factor, EEF1A2, has been identified as an oncogene in various solid tumors. Here, we have identified a novel function of EEF1A2 in angiogenesis.
Methods
Chick chorioallantoic membrane, tubulogenesis, aortic ring, Matrigel plug, and skin wound healing assays established EEF1A2’s role in angiogenesis.
Result
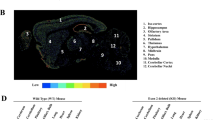
Higher EEF1A2 levels in breast cancer cells enhanced cell growth, movement, blood vessel function, and tubule formation in HUVECs, as confirmed by ex-ovo and in-vivo tests. The overexpression of EEF1A2 could be counteracted by Plitidepsin. Under normoxic conditions, EEF1A2 triggered HIF1A expression via ERK-Myc and mTOR signaling in TNBC and ER/PR positive cells. Hypoxia induced the expression of EEF1A2, leading to a positive feedback loop between EEF1A2 and HIF1A. Luciferase assay and EMSA confirmed HIF1A binding on the EEF1A2 promoter, which induced its transcription. RT-PCR and polysome profiling validated that EEF1A2 affected VEGF transcription and translation positively. This led to increased VEGF release from breast cancer cells, activating ERK and PI3K-AKT signaling in endothelial cells. Breast cancer tissues with elevated EEF1A2 showed higher microvessel density.
Conclusion
EEF1A2 exhibits angiogenic potential in both normoxic and hypoxic conditions, underscoring its dual role in promoting EMT and angiogenesis, rendering it a promising target for cancer therapy.








Similar content being viewed by others
Data availability
All data and materials related to this research are available in the manuscript or supplementary information.
References
Wang R, Zhu Y, Liu X, Liao X, He J, Niu L. The clinicopathological features and survival outcomes of patients with different metastatic sites in stage IV breast cancer. BMC Cancer. 2019;19:1–12.
Foote RL, Weidner N, Harris J, Hammond E, Lewis JE, Vuong T, et al. Evaluation of tumor angiogenesis measured with microvessel density (MVD) as a prognostic indicator in nasopharyngeal carcinoma: results of RTOG 9505. Int J Radiat Oncol Biol Phys. 2005;61:745–53.
Bais C, Mueller B, Brady MF, Mannel RS, Burger RA, Wei W, et al. Tumor microvessel density as a potential predictive marker for bevacizumab benefit: GOG-0218 biomarker analyses. J Natl Cancer Inst. 2017;109:djx066.
Mucci LA, Powolny A, Giovannucci E, Liao Z, Kenfield SA, Shen R, et al. Prospective study of prostate tumor angiogenesis and cancer-specific mortality in the health professionals follow-up study. J Clin Oncol. 2009;27:5627.
Krock BL, Skuli N, Simon MC. Hypoxia-induced angiogenesis: good and evil. Genes Cancer. 2011;2:1117–33.
Shibuya M. Vascular endothelial growth factor (VEGF) and its receptor (VEGFR) signaling in angiogenesis: a crucial target for anti-and pro-angiogenic therapies. Genes Cancer. 2011;2:1097–105.
Vasudev NS, Reynolds AR. Anti-angiogenic therapy for cancer: current progress, unresolved questions and future directions. Angiogenesis. 2014;17:471–94.
Abbas W, Kumar A, Herbein G. The eEF1A proteins: at the crossroads of oncogenesis, apoptosis, and viral infections. Front Oncol. 2015;5:75.
Anand N, Murthy S, Amann G, Wernick M, Porter LA, Cukier IH, et al. Protein elongation factor EEF1A2 is a putative oncogene in ovarian cancer. Nat Genet. 2002;31:301–5.
Cao H, Zhu Q, Huang J, Li B, Zhang S, Yao W, et al. Regulation and functional role of eEF1A2 in pancreatic carcinoma. Biochem Biophys Res Commun. 2009;380:11–6.
Pellegrino R, Calvisi DF, Neumann O, Kolluru V, Wesely J, Chen X, et al. EEF1A2 inactivates p53 by way of PI3K/AKT/mTOR‐dependent stabilization of MDM4 in hepatocellular carcinoma. Hepatology. 2014;59:1886–99.
Kulkarni G, Turbin DA, Amiri A, Jeganathan S, Andrade-Navarro MA, Wu TD, et al. Expression of protein elongation factor eEF1A2 predicts favorable outcome in breast cancer. Breast Cancer Res Treat. 2007;102:31–41.
Hassan MK, Kumar D, Patel SA, Dixit M. EEF1A2 triggers stronger ERK mediated metastatic program in ER negative breast cancer cells than in ER positive cells. Life Sci. 2020;262:118553.
Knowles HJ, Cleton-Jansen A, Korsching E, Athanasou NA. Hypoxia‐inducible factor regulates osteoclast‐mediated bone resorption: role of angiopoietin‐like 4. FASEB J. 2010;24:4648–59.
Kondo K, Klco J, Nakamura E, Lechpammer M, Kaelin WG Jr. Inhibition of HIF is necessary for tumor suppression by the von Hippel-Lindau protein. Cancer Cell. 2002;1:237–46.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Kumar D, Patel SA, Khan R, Chawla S, Mohapatra N, Dixit M. IQ Motif-containing GTPase-activating protein 2 inhibits breast cancer angiogenesis by suppressing VEGFR2–AKT signaling. Mol Cancer Res. 2022;20:77–91.
Naik M, Brahma P, Dixit M. A cost-effective and efficient chick ex-ovo CAM assay protocol to assess angiogenesis. Methods Protoc. 2018;1:19.
Rio DC, Ares M, Hannon GJ, Nilsen TW. Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb Protoc. 2010;2010:pdb-prot5439.
Kang H. Sample size determination and power analysis using the G* Power software. J Educ Eval Health Prof. 2021;18:17.
De Francesco EM, Lappano R, Santolla MF, Marsico S, Caruso A, Maggiolini M. HIF-1α/GPER signaling mediates the expression of VEGF induced by hypoxia in breast cancer associated fibroblasts (CAFs). Breast Cancer Res. 2013;15:1–18.
Wang Y, Yang C, Gu Q, Sims M, Gu W, Pfeffer LM, et al. KLF4 promotes angiogenesis by activating VEGF signaling in human retinal microvascular endothelial cells. PLoS One. 2015;10:e0130341.
Masoud GN, Li W. HIF-1α pathway: role, regulation and intervention for cancer therapy. Acta Pharm Sin B. 2015;5:378–89.
Wang X, Bove AM, Simone G, Ma B. Molecular bases of VEGFR-2-mediated physiological function and pathological role. Front Cell Dev Biol. 2020;8:599281.
Doe MR, Ascano JM, Kaur M, Cole MD. Myc posttranscriptionally induces HIF1 protein and target gene expression in normal and cancer cells. Cancer Res. 2012;72:949–57.
Lee T, Yao G, Nevins J, You L. Sensing and integration of Erk and PI3K signals by Myc. PLoS Comput Biol. 2008;4:e1000013.
Dodd KM, Yang J, Shen MH, Sampson JR, Tee AR. mTORC1 drives HIF-1α and VEGF-A signalling via multiple mechanisms involving 4E-BP1, S6K1 and STAT3. Oncogene. 2015;34:2239–50.
Xu C, Hu D, Zhu Q. eEF1A2 promotes cell migration, invasion and metastasis in pancreatic cancer by upregulating MMP-9 expression through Akt activation. Clin Exp Metastasis. 2013;30:933–44.
Worst TS, Waldbillig F, Abdelhadi A, Weis C-A, Gottschalt M, Steidler A, et al. The EEF1A2 gene expression as risk predictor in localized prostate cancer. BMC Urol. 2017;17:1–9.
Jia L, Ge X, Du C, Chen L, Zhou Y, **ong W, et al. EEF1A2 interacts with HSP90AB1 to promote lung adenocarcinoma metastasis via enhancing TGF-β/SMAD signalling. Br J Cancer. 2021;124:1301–11.
Losada A, Muñoz-Alonso MJ, Martínez-Díez M, Gago F, Domínguez JM, Martínez-Leal JF, et al. Binding of eEF1A2 to the RNA-dependent protein kinase PKR modulates its activity and promotes tumour cell survival. Br J Cancer. 2018;119:1410–20.
Jun JC, Rathore A, Younas H, Gilkes D, Polotsky VY. Hypoxia-inducible factors and cancer. Curr Sleep Med Rep. 2017;3:1–10.
Lee Y-H, Bae HC, Noh KH, Song K-H, Ye S, Mao C-P, et al. Gain of HIF-1α under normoxia in cancer mediates immune adaptation through the AKT/ERK and VEGFA AxesHIF-1α mediates cancer immune adaptation. Clin Cancer Res. 2015;21:1438–46.
Yang J, Tang J, Li J, Cen Y, Chen J, Dai G. Effect of activation of the Akt/mTOR signaling pathway by EEF1A2 on the biological behavior of osteosarcoma. Ann Transl Med. 2021;9:158.
Li Z, Qi C-F, Shin D-M, Zingone A, Newbery HJ, Kovalchuk AL, et al. Eef1a2 promotes cell growth, inhibits apoptosis and activates JAK/STAT and AKT signaling in mouse plasmacytomas. PLoS One. 2010;5:e10755.
Tsai W-B, Aiba I, Long Y, Lin H-K, Feun L, Savaraj N, et al. Activation of Ras/PI3K/ERK pathway induces c-Myc stabilization to upregulate argininosuccinate synthetase, leading to arginine deiminase resistance in melanoma cells. Cancer Res. 2012;72:2622–33.
Nilsson M, Heymach JV. Vascular endothelial growth factor (VEGF) pathway. J Thorac Oncol. 2006;1:768–70.
Baudino TA, McKay C, Pendeville-Samain H, Nilsson JA, Maclean KH, White EL, et al. c-Myc is essential for vasculogenesis and angiogenesis during development and tumor progression. Genes Dev. 2002;16:2530–43.
Li Z, Van Calcar S, Qu C, Cavenee WK, Zhang MQ, Ren B. A global transcriptional regulatory role for c-Myc in Burkitt’s lymphoma cells. Proc Natl Acad Sci. 2003;100:8164–9.
Shen Z, Li Y, Fang Y, Lin M, Feng X, Li Z, et al. SNX16 activates c‐Myc signaling by inhibiting ubiquitin‐mediated proteasomal degradation of eEF1A2 in colorectal cancer development. Mol Oncol. 2020;14:387–406.
Forsythe JA, Jiang B-H, Iyer NV, Agani F, Leung SW, Koos RD, et al. Activation of vascular endothelial growth factor gene transcription by hypoxia-inducible factor 1. Mol Cell Biol. 1996;16:4604–13.
Mezquita P, Parghi SS, Brandvold KA, Ruddell A. Myc regulates VEGF production in B cells by stimulating initiation of VEGF mRNA translation. Oncogene. 2005;24:889–901.
Land SC, Tee AR. Hypoxia-inducible factor 1α is regulated by the mammalian target of rapamycin (mTOR) via an mTOR signaling motif. J Biol Chem. 2007;282:20534–43.
Sengupta T, Abraham G, Xu Y, Clurman BE, Minella AC. Hypoxia-inducible factor 1 is activated by dysregulated cyclin E during mammary epithelial morphogenesis. Mol Cell Biol. 2011;31:3885–95.
Yu Y, Zhao D, Li K, Cai Y, Xu P, Li R, et al. E2F1 mediated DDX11 transcriptional activation promotes hepatocellular carcinoma progression through PI3K/AKT/mTOR pathway. Cell Death Dis. 2020;11:273.
Mavria G, Vercoulen Y, Yeo M, Paterson H, Karasarides M, Marais R, et al. ERK-MAPK signaling opposes Rho-kinase to promote endothelial cell survival and sprouting during angiogenesis. Cancer Cell. 2006;9:33–44.
Vera M, Pani B, Griffiths LA, Muchardt C, Abbott CM, Singer RH, et al. The translation elongation factor eEF1A1 couples transcription to translation during heat shock response. Elife. 2014;3:e03164.
Binet F, Sapieha P. ER stress and angiogenesis. Cell Metab. 2015;22:560–75.
**ao W, Wang J, Ou C, Zhang Y, Ma L, Weng W, et al. Mutual interaction between YAP and c-Myc is critical for carcinogenesis in liver cancer. Biochem Biophys Res Commun. 2013;439:167–72.
Henze A-T, Acker T. Feedback regulators of hypoxia-inducible factors and their role in cancer biology. Cell cycle. 2010;9:2821–35.
Hassan MK, Kumar D, Naik M, Dixit M. The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS One. 2018;13:e0191377.
Losada A, Muñoz-Alonso MJ, García C, Sánchez-Murcia PA, Martínez-Leal JF, Domínguez JM, et al. Translation elongation factor eEF1A2 is a novel anticancer target for the marine natural product plitidepsin. Sci Rep. 2016;6:1–15.
Acknowledgements
We thank Dr. Saurabh Chawla, NISER, Bhubaneswar, India for hel** with animal experiments. We thank Dr. Anup Kumar Ram and Dr. Pankaj V Alone for guidance in polysome analysis. We thank Ananya Palo, Soham Choudhury, Deeptima Jaiswar, and Talina Mohapatra for measuring absorbance of polysome fractions. We thank Ruchika Rai for hel** in power analysis. We thank Shubhanjali and Anamika Singh for the bioinformatic data analysis. This work was supported by intramural funding from the National Institute of Science Education and Research (NISER), Department of Atomic Energy (DAE), Government of India (GOI). SAP, MKH, and MN received fellowships from NISER, DAE, GOI.
Funding
We thank National Institute of Science Education and Research (Department of Atomic Energy), the Government of India (GOI) for providing infrastructure, and intramural support.
Author information
Authors and Affiliations
Contributions
SAP performed majority of the in vivo experiments; wound healing assay, matrigel plug assay, tumor development in mice, IHC of 92 samples and mice tissue, represented in Figs. 2c–f, 7a–h, 8a–f, Supplementary Fig. 9A–E and in vitro experiments; Western blot, EMSA, Luciferase, polysome assay, qRT-PCR, tubulogenesis, proliferation assay, ELISA, represented in Figs. 1a, c, d, 3a–n, 4a–i, 5a–j, 6a–n, and Supplementary Figs. 1C–G, 2A, B, 3A–D, 4B, 6A–L, 7A–F, 8A–L, data curation, formal analysis, and writing original draft and editing; MKH performed the in vitro experiments of Figs. 1b, c, 3a, 8e, Supplementary Fig. 1A, 1B, 1H, 1I, 5A-D, and performed IHC of 96 samples (Fig. 8e), wrote the original draft partly; MN performed the CAM assay (Fig. 2a) and Matrigel Plug assay (Supplementary Fig. 4A, B, D); NM analysed and scored patient IHC samples, PB performed and analysed the experiment given in Fig. 2b; PSK supervised the experiments carried out for generating Fig. 2b; MD conceptualized the whole project, designed experiments, planned and guided the research, formal analysis, supervision, funding acquisition, resources, review and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The animal experiment was approved by Institutional Animal Ethics Committee, NISER, India (protocol-NISER/SBS/AH184), (protocol-NISER/SBS/AH/224), and (protocol-NISER/SBS/AH/170) and conducted at NISER animal house facility. Human samples-based study was approved by the Institutional Ethics Committee, NISER, Bhubaneswar, India (protocol-NISER/IEC/2016–02).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patel, S.A., Hassan, M.K., Naik, M. et al. EEF1A2 promotes HIF1A mediated breast cancer angiogenesis in normoxia and participates in a positive feedback loop with HIF1A in hypoxia. Br J Cancer 130, 184–200 (2024). https://doi.org/10.1038/s41416-023-02509-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02509-2
- Springer Nature Limited