Abstract
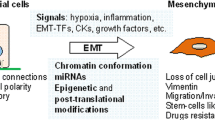
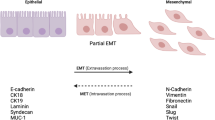
Epithelial–mesenchymal transition (EMT) is an important biological process that has been implicated in cancer metastasis. Epithelial cell adhesion molecule (EpCAM) is expressed at the basolateral membrane of most normal epithelial cells but is overexpressed in many epithelial cancers. In our studies on the role of EpCAM in cancer biology, we observed that EpCAM expression is decreased in mesenchymal-like primary cancer specimens in vivo and following induction of EMT in cancer cell lines in vitro. Extracellular signal-related kinase (ERK) is a key regulator of EMT. We observed that EpCAM expression is decreased with activation of the ERK pathway in primary cancer specimens in vivo and in cancer cell lines in vitro. In experimental models, growth factor stimulation and/or oncogene-induced ERK2 activation suppressed EpCAM expression, whereas genetic or pharmacological inhibition of the ERK pathway restored EpCAM expression. In detailed studies of the EpCAM promoter region, we observed that ERK2 suppresses EpCAM transcription directly by binding to a consensus ERK2-binding site in the EpCAM promoter and indirectly through activation of EMT-associated transcription factors SNAI1, SNAI2, TWIST1 and ZEB1, which bind to E-box sites in the EpCAM promoter. Surprisingly, EpCAM appears to modulate ERK activity. Using multiple cell lines, we demonstrated that specific ablation of EpCAM resulted in increased ERK pathway activity and SNAI2 expression, migration and invasion, whereas forced expression of EpCAM resulted in decreased ERK pathway activity and SNAI2 expression, migration and invasion. These observations provide important insights into the regulation of EpCAM expression during EMT, demonstrate an unexpected role for EpCAM in the regulation of ERK and define a novel double-negative feedback loop between EpCAM and ERK that contributes to the regulation of EMT. These studies have important translational implications as both EpCAM and ERK are currently being targeted in human clinical trials.






Similar content being viewed by others
References
Klymkowsky MW, Savagner P . Epithelial-mesenchymal transition: a cancer researcher's conceptual friend and foe. Am J Pathol 2009; 174: 1588–1593.
Polyak K, Weinberg RA . Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer 2009; 9: 265–273.
Thiery JP, Acloque H, Huang RY, Nieto MA . Epithelial-mesenchymal transitions in development and disease. Cell 2009; 139: 871–890.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008; 133: 704–715.
Shin J, Yang J, Lee J, Baek KH . Depletion of ERK2 but not ERK1 abrogates oncogenic Ras-induced senescence. Cell Signal 2013; 25: 2540–2547.
Shin S, Dimitri CA, Yoon SO, Dowdle W, Blenis J . ERK2 but not ERK1 induces epithelial-to-mesenchymal transformation via DEF motif-dependent signaling events. Mol Cell 2010; 38: 114–127.
Brabletz S, Brabletz T . The ZEB/miR-200 feedback loop—a motor of cellular plasticity in development and cancer? EMBO Rep 2010; 11: 670–677.
Feldman S, Krishnamurthy S, Gillanders W, Gittleman M, Beitsch PD, Young PR et al. A novel automated assay for the rapid identification of metastatic breast carcinoma in sentinel lymph nodes. Cancer 2011; 117: 2599–2607.
Armstrong A, Eck SL . EpCAM: a new therapeutic target for an old cancer antigen. Cancer Biol Ther 2003; 2: 320–326.
Munz M, Murr A, Kvesic M, Rau D, Mangold S, Pflanz S et al. Side-by-side analysis of five clinically tested anti-EpCAM monoclonal antibodies. Cancer Cell Int 2010; 10: 44.
Cristofanilli M, Budd GT, Ellis MJ, Stopeck A, Matera J, Miller MC et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N Engl J Med 2004; 351: 781–791.
Osta WA, Chen Y, Mikhitarian K, Mitas M, Salem M, Hannun YA et al. EpCAM is overexpressed in breast cancer and is a potential target for breast cancer gene therapy. Cancer Res 2004; 64: 5818–5824.
Sankpal NV, Willman MW, Fleming TP, Mayfield JD, Gillanders WE . Transcriptional repression of epithelial cell adhesion molecule contributes to p53 control of breast cancer invasion. Cancer Res 2009; 69: 753–757.
Sankpal NV, Mayfield JD, Willman MW, Fleming TP, Gillanders WE . Activator protein 1 (AP-1) contributes to EpCAM-dependent breast cancer invasion. Breast Cancer Res 2011; 13: R124.
Sankpal NV, Fleming TP, Gillanders WE . EpCAM modulates NF-kappaB signaling and interleukin-8 expression in breast cancer. Mol Cancer Res 2013; 11: 418–426.
Maetzel D, Denzel S, Mack B, Canis M, Went P, Benk M et al. Nuclear signalling by tumour-associated antigen EpCAM. Nat Cell Biol 2009; 11: 162–171.
Cheng JC, Chang HM, Leung PC . Egr-1 mediates epidermal growth factor-induced downregulation of E-cadherin expression via Slug in human ovarian cancer cells. Oncogene 2013; 32: 1041–1049.
Zhang K, Corsa CA, Ponik SM, Prior JL, Piwnica-Worms D, Eliceiri KW et al. The collagen receptor discoidin domain receptor 2 stabilizes SNAIL1 to facilitate breast cancer metastasis. Nat Cell Biol 2013; 15: 677–687.
Caramel J, Papadogeorgakis E, Hill L, Browne GJ, Richard G, Wierinckx A et al. A switch in the expression of embryonic EMT-inducers drives the development of malignant melanoma. Cancer Cell 2013; 24: 466–480.
Byers LA, Diao L, Wang J, Saintigny P, Girard L, Peyton M et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin Cancer Res 2013; 19: 279–290.
Gao J, Yan Q, Wang J, Liu S, Yang X . Epithelial-to-mesenchymal transition induced by TGF-beta1 is mediated by AP1-dependent EpCAM expression in MCF-7 cells. J Cell Physiol 2015; 230: 775–782.
Santisteban M, Reiman JM, Asiedu MK, Behrens MD, Nassar A, Kalli KR et al. Immune-induced epithelial to mesenchymal transition in vivo generates breast cancer stem cells. Cancer Res 2009; 69: 2887–2895.
Ye X, Tam WL, Shibue T, Kaygusuz Y, Reinhardt F, Ng Eaton E et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 2015; 525: 256–260.
Lyons JG, Patel V, Roue NC, Fok SY, Soon LL, Halliday GM et al. Snail up-regulates proinflammatory mediators and inhibits differentiation in oral keratinocytes. Cancer Res 2008; 68: 4525–4530.
Gemmill RM, Roche J, Potiron VA, Nasarre P, Mitas M, Coldren CD et al. ZEB1-responsive genes in non-small cell lung cancer. Cancer Lett 2011; 300: 66–78.
Vannier C, Mock K, Brabletz T, Driever W . Zeb1 regulates E-cadherin and Epcam (epithelial cell adhesion molecule) expression to control cell behavior in early zebrafish development. J Biol Chem 2013; 288: 18643–18659.
Massoner P, Thomm T, Mack B, Untergasser G, Martowicz A, Bobowski K et al. EpCAM is overexpressed in local and metastatic prostate cancer, suppressed by chemotherapy and modulated by MET-associated miRNA-200c/205. Br J Cancer 2014; 111: 955–964.
Hook KE, Garza SJ, Lira ME, Ching KA, Lee NV, Cao J et al. An integrated genomic approach to identify predictive biomarkers of response to the aurora kinase inhibitor PF-03814735. Mol Cancer Ther 2012; 11: 710–719.
Raponi M, Zhang Y, Yu J, Chen G, Lee G, Taylor JM et al. Gene expression signatures for predicting prognosis of squamous cell and adenocarcinomas of the lung. Cancer Res 2006; 66: 7466–7472.
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res 2010; 12: R68.
Asiedu MK, Ingle JN, Behrens MD, Radisky DC, Knutson KL . TGFbeta/TNF(alpha)-mediated epithelial-mesenchymal transition generates breast cancer stem cells with a claudin-low phenotype. Cancer Res 2011; 71: 4707–4719.
Dry JR, Pavey S, Pratilas CA, Harbron C, Runswick S, Hodgson D et al. Transcriptional pathway signatures predict MEK addiction and response to selumetinib (AZD6244). Cancer Res 2010; 70: 2264–2273.
Hu S, **e Z, Onishi A, Yu X, Jiang L, Lin J et al. Profiling the human protein-DNA interactome reveals ERK2 as a transcriptional repressor of interferon signaling. Cell 2009; 139: 610–622.
Grotegut S, von Schweinitz D, Christofori G, Lehembre F . Hepatocyte growth factor induces cell scattering through MAPK/Egr-1-mediated upregulation of Snail. EMBO J 2006; 25: 3534–3545.
Hajra KM, Chen DY, Fearon ER . The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res 2002; 62: 1613–1618.
Taube JH, Herschkowitz JI, Komurov K, Zhou AY, Gupta S, Yang J et al. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc Natl Acad Sci USA 2010; 107: 15449–15454.
Zhang P, Sun Y, Ma L . ZEB1: at the crossroads of epithelial-mesenchymal transition, metastasis and therapy resistance. Cell Cycle 2015; 14: 481–487.
Kusewitt DF, Choi C, Newkirk KM, Leroy P, Li Y, Chavez MG et al. Slug/Snai2 is a downstream mediator of epidermal growth factor receptor-stimulated reepithelialization. J Invest Dermatol 2009; 129: 491–495.
Maghzal N, Kayali HA, Rohani N, Kajava AV, Fagotto F . EpCAM controls actomyosin contractility and cell adhesion by direct inhibition of PKC. Dev Cell 2013; 27: 263–277.
Tarcic G, Avraham R, Pines G, Amit I, Shay T, Lu Y et al. EGR1 and the ERK-ERF axis drive mammary cell migration in response to EGF. FASEB J 2012; 26: 1582–1592.
Fischer KR, Durrans A, Lee S, Sheng J, Li F, Wong ST et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 2015; 527: 472–476.
Singh A, Settleman J . EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene 2010; 29: 4741–4751.
Zheng X, Carstens JL, Kim J, Scheible M, Kaye J, Sugimoto H et al. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature 2015; 527: 525–530.
Lamouille S, Xu J, Derynck R . Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 2014; 15: 178–196.
Carlson SM, Chouinard CR, Labadorf A, Lam CJ, Schmelzle K, Fraenkel E et al. Large-scale discovery of ERK2 substrates identifies ERK-mediated transcriptional regulation by ETV3. Sci Signal 2011; 4: rs11.
Courcelles M, Fremin C, Voisin L, Lemieux S, Meloche S, Thibault P . Phosphoproteome dynamics reveal novel ERK1/2 MAP kinase substrates with broad spectrum of functions. Mol Syst Biol 2013; 9: 669.
Roskoski R Jr . ERK1/2 MAP kinases: structure, function, and regulation. Pharmacol Res 2012; 66: 105–143.
Munz M, Baeuerle PA, Gires O . The emerging role of EpCAM in cancer and stem cell signaling. Cancer Res 2009; 69: 5627–5629.
Eberlein C, Rooney C, Ross SJ, Farren M, Weir HM, Barry ST . E-Cadherin and EpCAM expression by NSCLC tumour cells associate with normal fibroblast activation through a pathway initiated by integrin alphavbeta6 and maintained through TGFbeta signalling. Oncogene 2014; 34: 704–716.
Soysal SD, Muenst S, Barbie T, Fleming T, Gao F, Spizzo G et al. EpCAM expression varies significantly and is differentially associated with prognosis in the luminal B HER2(+), basal-like, and HER2 intrinsic subtypes of breast cancer. Br J Cancer 2013; 108: 1480–1487.
Songun I, Litvinov SV, van de Velde CJ, Pals ST, Hermans J, van Krieken JH . Loss of Ep-CAM (CO17-1A) expression predicts survival in patients with gastric cancer. Br J Cancer 2005; 92: 1767–1772.
Spizzo G, Went P, Dirnhofer S, Obrist P, Simon R, Spichtin H et al. High Ep-CAM expression is associated with poor prognosis in node-positive breast cancer. Breast Cancer Res Treat 2004; 86: 207–213.
Yang X, Lin X, Zhong X, Kaur S, Li N, Liang S et al. Double-negative feedback loop between reprogramming factor LIN28 and microRNA let-7 regulates aldehyde dehydrogenase 1-positive cancer stem cells. Cancer Res 2010; 70: 9463–9472.
Shi L, Jackstadt R, Siemens H, Li H, Kirchner T, Hermeking H . p53-induced miR-15a/16-1 and AP4 form a double-negative feedback loop to regulate epithelial-mesenchymal transition and metastasis in colorectal cancer. Cancer Res 2014; 74: 532–542.
Siemens H, Jackstadt R, Hunten S, Kaller M, Menssen A, Gotz U et al. miR-34 and SNAIL form a double-negative feedback loop to regulate epithelial-mesenchymal transitions. Cell Cycle 2011; 10: 4256–4271.
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 2008; 68: 7846–7854.
Liu S, Cong Y, Wang D, Sun Y, Deng L, Liu Y et al. Breast cancer stem cells transition between epithelial and mesenchymal states reflective of their normal counterparts. Stem Cell Rep 2014; 2: 78–91.
Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA . Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res 2008; 68: 3645–3654.
Li S, Shen D, Shao J, Crowder R, Liu W, Prat A et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep 2013; 4: 1116–1130.
Acknowledgements
We thank Dr Kerry Kornfeld, Dr Jason Weber and Dr Gregory Longmore for reviewing the manuscript and providing valuable advice. This work was supported by a grant from Susan G Komen for the Cure (BCTR0707917) and Siteman Cancer Center fund (P30CA91842).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Sankpal, N., Fleming, T., Sharma, P. et al. A double-negative feedback loop between EpCAM and ERK contributes to the regulation of epithelial–mesenchymal transition in cancer. Oncogene 36, 3706–3717 (2017). https://doi.org/10.1038/onc.2016.504
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.504
- Springer Nature Limited