Abstract
The global pandemic of COVID-19 is a serious public health concern. Over 625 million confirmed cases and more than 6 million deaths have been recorded worldwide. Although several vaccines and antiviral medications have been developed, their efficacy is limited by the emerging new SARS-CoV-2 strains. Peptide-based therapeutics is a fast-growing class of new drugs and have unique advantages over large proteins and small molecules. Antiviral peptides (AVPs) are short polycationic antivirals with broad-spectrum effects, which have been shown to exert both prophylactic and therapeutic actions against reported coronaviruses. The potential therapeutic targets of AVPs are located either on the virus (e.g., E-protein and S-protein) to prohibit viral binding or host cells, particularly, those present on the cell surface (e.g., ACE2 and TMPRSS2). Despite AVPs having promising antiviral effects, their efficacy is limited by low bioavailability. Thus, nanoformulation is a prerequisite for prolonged bioavailability and efficient delivery. This review aimed to present an insight into the therapeutic AVP targets on both virus and host cells by discussing their antiviral activities and associated molecular mechanisms. Besides, it described the technique for discovering and develo** possible AVPs based on their targets, as well as the significance of using nanotechnology for their efficient delivery against SARS-CoV-2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the last 2 decades, there have been numerous epidemics, including severe acute respiratory syndrome (SARS) coronavirus, H1N1 influenza virus, Middle East respiratory syndrome (MERS) coronavirus, Ebola virus, and Zika virus [1,2,3,4,5]. Among these epidemics, the current global coronavirus disease 2019 (COVID-19) is the most contagious. The pathogen behind the COVID-19 epidemic is severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [6]. An excess of 625 million individuals have been affected by the current pandemic due to SARS-CoV-2, with more than 6 million (approximately 1.04%) deaths recorded as of October 2022 [7]. Since its identification, the world scientific community has continually searched for treatments against SARS-CoV-2. Traditional approaches for drug discovery and development are time-consuming and can take an average of 15 years due to regulatory standards to analyze drug safety and efficacy. Currently, the US Food and Drug Administration (US-FDA) has approved Veklury, also known as remdesivir, an anti-viral drug for certain adult and pediatric cases. Additionally, the US-FDA has also approved monoclonal antibodies under the category of Emergency Use Authorization (EUA) for prophylactic and therapeutic use against COVID-19 [8].
Several vaccines against COVID-19 have recently been administered to human subjects under emergency use globally. According to the World Health Organization (WHO), approximately 60% of the world population has been fully vaccinated as of October 2022 [7]. Currently, three types of vaccines are employed to prevent SARS-CoV-2 infections, including messenger ribonucleic acid (mRNA), adenovirus vector-based, and inactivated virus-based vaccines [9]. Current US-FDA and WHO approved vaccines are produced by Pfizer-BioNTech (USA), Moderna (USA), Johnson & Johnson (USA), AstraZeneca and Oxford University (USA and UK), Sinopharm (China), Sinovac (China), CanSino Biologics (China), and Gamaleya (Russia), Novavax COVID-19 Vaccine, Adjuvanted (USA), Serum Institute of India (India), Bharat Biotech International (India) [10,11,12,13]. Despite the rapid production and dissemination of vaccines, multiple new variants, such as alpha, beta, gamma, delta, and omicron, have emerged from various regions in the world, constituting the requirement for booster shots for enhanced prophylactic efficacy [14]. Other therapeutic approaches, such as the use of antiviral drugs, have been investigated for the treatment of COVID-19 patients. Remdesivir, hydroxychloroquine, and lopinavir-ritonavir are the three major antiviral drugs that have been utilized to treat COVID-19 [12]. However, hydroxychloroquine and lopinavir–ritonavir are not used as often because of their uncertain effectiveness [14]. Recently, the EUA of molnupiravir (Merck) as a new oral antiviral has been considered by the FDA [15]. On the other hand, the FDA has approved the EUA of paxlovid (Pfizer) as an oral antiviral treatment for patients who are at a high risk of develo** severe COVID-19 [15]. Paxlovid is an inhibitor of SARS-Cov-2–3 CL protease, while molnupiravir acts as a nucleoside analogue that leads to lethal mutagenesis of the viral genome [15].
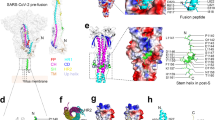
Apart from vaccines and antiviral drugs, antiviral peptides (AVPs) have demonstrated preventive benefits against coronaviruses [16]. Antiviral polypeptides have been reported to have a significantly broad range of mechanism of action. For example, mucroporin-M1 induced destruction of the viral envelope [17], whereas targeting the heptad repeat 2 (HR2)-M2, EK1, EK1C4, and TMPRSS2 within the viral Spike (S) protein to inhibit protein-mediated fusion, the S2 subunit HR1 domain, and S-protein priming processes, respectively [18, SARS-CoV-2 is the causative agent of the ongoing global COVID-19 pandemic that infected 480 million individuals by April 2022 and caused over 6 million deaths [7]. The outbreak was first reported in China, where the source of infection of several patients with pneumonia of unknown origin was all linked to a Wuhan seafood market [24]. It was first identified as the novel coronavirus (2019-nCoV), and the name of the virus and disease were later changed to SARS-CoV-2 and COVID-19, respectively [25]. The virus falls under the genus betacoronavirus and family Coronaviridae, and it has a positive sense, single-stranded RNA genome of approximately 29.9 kb. The nucleotide sequence of SARS-CoV-2 shows 79 and 50% sequence homology with SARS-CoV and MERS, respectively [26]. Specifically, the SARS-CoV-2 and SARS-CoV S1 subunit of the S protein share approximately 50 conserved amino acids that otherwise showed mutational differences among other coronaviruses. Additionally, despite being categorized into the same phylogenetic clades as bat-derived coronaviruses, the RBD of SARS-CoV-2 shared higher genetic similarity with SARS-CoV, both of which bind to ACE2 receptor on the host cell membrane for viral entry. These findings suggested that there might be a similarity between the immune responses generated by these three coronaviruses [26]. The SARS-CoV-2 encodes 16 non-structural proteins (NSP-1–16), four structural proteins, such as the S, membrane (M), envelope (E), and nucleocapsid (N) proteins, and accessory proteins (Fig. 1). The 5′ untranslated region (UTR) of the SARS-CoV-2 genome consists of two open-reading frames (ORFs) (i.e., ORF1a and ORF1b), which make up two-thirds of the viral genome, giving rise to the large polypeptides 1a (pp1a), and 1ab (pp1ab). These polyproteins are then cleaved into a total of 16 NSPs, where NSPs 1–11 are encoded by pp1a and pp1ab gives rise to NSPs 12–16 [27]. The functional involvement of 16 NSPs includes viral replication and transcription [28, 29]. The 3′ end of the viral genome consists of conserved regions, such as the stem-loop-II-like motif (s2), hyper-variable region (HVR), mutually exclusive bulged stem-loop (BSL), which are involved in the synthesis of viral RNA [30]. The genetic structure and function of SARS-CoV-2. SARS-CoV-2 is made up of structural proteins, such as the envelope (E), spike (S), and membrane (M) proteins, as well as a nucleocapsid (N) containing the viral RNA genome. The structural and accessory proteins are essential for SARS-CoV-2 pathogenicity The structural S protein is approximately 180–200 kDa in size and is made up of an extracellular N-terminal domain, receptor-binding domain (RBD) in the S1 subunit, an intracellularly located C-terminal domain, transmembrane (TM) domain, and heptad repeat 1 (HR1) and HR2 located in the S2 subunit [51,52,53]. The dominant clinical manifestations of SARS-CoV-2 include fever, cough, and shortness of breath. Subsequent clinical manifestations such as chills, muscle pain, headache, sore throat, diarrhea, skin rash, or discoloration of fingers or toes, red or irritated eyes, and loss of taste or smell were included by the US Center for Disease Control and Prevention (CDC) [53, 54]. Some patients show early symptoms like rhinorrhea, nausea, chest tightness, loss of speech or mobility, or confusion and vomiting before the onset of fever, suggesting that fever is not only considered as an early symptom but also a critical condition. The incubation period of SARS-CoV-2 is between 1 and 14 days, and in most cases, it is about three to seven days. Most of the time, the infected patients showed symptoms of infection after the incubation period, while shortness of breath and pneumonia developed within an average time of eight days [54, 55]. On the contrary, some patients only show mild fever, mild fatigue, or even asymptomatic. This group of patients generally recover after one week, while severe cases have been documented that may develop progressive respiratory failure, as a result of alveolar destruction which can cause death [56]. AVPs are peptides that can inhibit viral infection or replication. The mechanism of action for each AVP type is different based on its target. These sites of action or inhibition can be found on the virus or host cell and are involved in essential steps of the viral replication cycle. Increasing interest towards the use of AVPs against many viral infections, including SARS-CoV-2, has been generated during the pandemic. This phenomenon is primarily due to their high specificity and potent antiviral activity even at low doses. Furthermore, many AVPs can be derived from natural or biological sources and have fewer associated adverse effects and cytotoxicity [57]. Intriguingly, the FDA has approved protease inhibitors, including both peptides (e.g., Enfuvirtide) and peptidomimetics mimicking the structural and functional properties of peptides (e.g., Boceprevir, and Telaprevir) for use against infectious diseases due to their ease of synthesis, greater efficacy, safety, and tolerability [58, 59]. Protein–protein interactions are essential for many cellular processes, including viral infections, and have been identified as central targets for drug discovery and development [60]. For instance, therapeutic peptides are target-specific peptides that are composed of less than 100 amino acids and are highly specific [61]. Furthermore, they are also non-toxic, because they can easily be hydrolyzed by normal host peptidases, thus preventing accumulation in organs that can lead to toxicity and other adverse effects. Some essential properties for AVPs include hydrophobicity, which is considered essential for targeting enveloped viruses specifically, as well as high target specificity, which is required for all types of AVPs to be effective [62]. Due to the complete dependence of viruses on the host cell machinery to complete most of the stages in their replication cycle, AVPs can be developed by targeting either the virus or host cells. Stages in viral replication that can be targeted include viral attachment, entry, uncoating, synthesis, and assembly [63]. The mechanism of action used by AVPs is generally summarized as being virucidal, as they are specifically designed or chosen based on their ability to interfere with essential steps in the viral life cycle to prevent viruses from establishing successful infection within the host cells. AVPs can be designed to interact with the viral membrane or structural proteins to successfully prevent viral attachment or entry. On the other hand, develo** host-directed antiviral peptides is a promising alternative strategy with a reduced likelihood of resistance [64]. Several strategies are currently used in the development of AVPs, including computationally derived peptides, peptides derived from plants, animals, and microorganisms, or peptides isolated from biological sources [59, 63]. Computational methods of AVP identification and design include molecular docking and simulations, as well as peptidomimetics [63, 65]. The process of molecular docking, where the interactions between molecules are analyzed by computational modeling focusing on their structural orientations and conformations, has accelerated drug development. This method provides the means to identify potential target sites and peptides with the highest binding affinity via an in silico approach. While peptidomimetics is another approach in which the biological target is mimicked, and its interaction with other proteins is examined. Both in silico and computational methods allow early identification of potential targets for AVPs. However, AVPs identified based on these methods are required to be validated using in vitro and in vivo approaches to gain a deeper understanding of their biological activities and antiviral efficacies. The second approach that has been used in AVP development is identifying natural compounds showing potential antiviral activity [63]. The natural compounds are usually antimicrobial peptides (AMPs) that may have potent antiviral activity and can be isolated from natural sources, such as plants, animals, fungi, and bacteria. Depending on the mechanism of action, the properties of these natural AMPs and AVPs allow them to target viral proteins that are prerequisites for viral replication. One example of a naturally occurring peptide family with a broad-spectrum antiviral activity is cyclotides. These plant-derived AVPs have cationic and amphipathic properties, and they have been reported to prevent dengue, human immunodeficiency virus (HIV), and influenza infections by causing rupture of the viral membrane [66,67,68]. Comparatively, AMPs are often preferred over synthetic antiviral drugs due to their higher resistance to mutations and show a broad spectrum of antimicrobial activity [69]. Besides, the biological approach using in vitro display methodologies can also be used to identify and generate potential AVPs. They are usually genetically encoded to generate peptides with higher binding affinities to their targets. The most common techniques used in the biological approach include peptides displayed by phage, mRNA, ribosome, and yeast peptide libraries [59]. The phage display method fuses exogenous peptide sequences into the genome of a phage for surface expression, whereas mRNA display method extracts transcription and translation machinery from prokaryotic or eukaryotic cells to perform in vitro translation of covalently bonded mRNA polypeptide complexes linked through puromycin. The ribosome display method uses in vitro translation of non-covalent-ribosome-mRNA-polypeptide complexes to couple genotypes and phenotypes which can select high-affinity peptides. Lastly, the yeast display method relies on the integration of the protein or peptide of interest on the cell surface glycoproteins via N- or C-terminal fusion. Yeast proteins that are used include both Ag-alpha-1 and aga-2, due to their role in yeast cell mating. The biological approach is considered the most appropriate method of screening different peptides for their potential antiviral activity, as the high affinity for their targets is determined using in vitro techniques. As mentioned earlier, AVPs can be designed to target proteins that play an essential role in the viral replication cycle. The potential therapeutic targets of AVPs against SARS-CoV-2 can be categorized based on their location in either the viral cells or host cells, particularly, on the cell surface. The AVP viral targets are those that are involved in the prevention of viral infections by inhibiting viral entry into the host cells, including viral E-protein and S-protein sites (e.g., RBD and HR1/HR2 domains), as well as the viral membrane. Viral glycoproteins have been identified as preferred target sites for antiviral drugs in the past due to their importance in viral attachment and entry [70]. In agreement with this, two major sites that have been identified as potential targets in the AVP development against SARS-CoV-2 are the S1 and S2 subunits of S protein, which can prevent the virus from binding to host cells [71]. These targets are more favorable due to their extracellular location as they induce a lower risk of adverse toxicity to host cells. AVP host cell targets, on the other hand, are those that are involved in facilitating and allowing the viral S protein binding and entry (e.g., ACE2 and TMPRSS2). The following sections describe the therapeutic targets of AVPs on both viral and host cells, as well as their mechanistic activities, both of which have been summarized in Table 1. SARS-CoV-2 S protein. S protein is a major site of interest in the identification and development of binding and/or attachment-inhibiting AVPs that have broad-spectrum activity against SARS-CoV-2 and other coronaviruses. This is because S proteins of viruses in the Coronaviridae family have similar nucleotide sequences. Particularly, the RBD of SARS-CoV and SARS-CoV-2 share 73–76% similarity in their nucleotide sequences [32]. Jaiswal and Kumar [72] used in silico methods to design an AVP targeting the S protein of SARS-CoV-2. The AVP of interest, ΔABP-D25Y, was designed based on the ACE2 α-helical region [72]. Using molecular docking simulations, they found that the synthetically designed AVP might have the ability to inhibit SARS-CoV-2 infection by competitively blocking RBD interaction with ACE2 receptors on host cells. Additionally, Chowdhury et al. [73] also used molecular docking to screen 51 AVPs with known antiviral activity against SARS-CoV for their ability to bind to SARS-CoV-2. They found that S2P25, S2P26, and 13 other peptides that have higher binding affinity for the α-helical region of the RBD showed the most promising SARS-CoV-2-specific antiviral activity [73]. Contradictory to the appeal of using the same AVP against SARS-CoV and SARS-CoV-2 due to the similarity in their S protein, both SARS-CoV specific murine monoclonal antibodies and polyclonal antibodies were found to lack the ability to interact with SARS-CoV-2 RBD [73]. The S2 subunit of SARS-CoV and SARS-CoV-2 have approximately 90% nucleotide sequence identity, which is higher than the RBD in the S1 subunit, hence, it is a more desirable target site for AVPs with broad-spectrum anti-coronavirus activity [74]. Due to the critical role of HR1 and HR2 of the S2 subunit in membrane fusion, they have been identified as potential targets in AVP development [32]. In fact, AVPs that target these fusion sites have the potential to confer broad-spectrum antiviral activity against coronaviruses. One such fusion inhibitor is EK1 which has pan-coronavirus antiviral activity and is effective against SARS-CoV, SARS-CoV-2, and MERS-CoV infection [32, 82]. SARS-CoV-2 Mpro is a homodimer that is consisted of two protomers and three domains, with domains I and II being composed of six antiparallel β-barrels, while domain III is made of five α helices. The proteolytic activity of Mpro is conferred by N-terminus and Cys-His catalytic diad located in the cleft between domains I and II. It is conserved across all coronaviruses, making it an ideal target for antiviral drugs with broad-spectrum activity against all human coronaviruses [83, 84]. Zhang et al. used in silico methods to predict the antiviral activity of SARS-CoV-2 Mpro inhibiting α-ketoamide, which is also known as 13a [83]. The design of the compound was based on modifying 11r, which was previously identified as having broad-spectrum activity against coronaviruses and adenoviruses by specifically targeting main proteases of both alpha and beta coronaviruses and 3C protease of enteroviruses. To design 13a, 11r was modified by hiding the P3-P2 amide bond in a pyridine ring, followed by replacing hydrophobic cinnamoyl moiety with a relatively less hydrophobic Boc group to prevent cellular proteases from cleaving the amide bond and increasing solubility in plasma for a decreased plasma protein binding, respectively. Using peptidomimetics, it was shown that 13a potentially had SARS-CoV-2 specific antiviral activity by binding with the SARS-CoV-2 Mpro to prevent viral maturation by inhibiting RNA replication. One other approach that has been utilized to confer a broad-spectrum antiviral activity is the use of peptides to interfere with the viral uncoating stage, which in turn inhibited the release of genetic material into the host cells. For instance, P9 peptide, derived from mouse beta-defensin-4, could bind to SARS-CoV, MERS-CoV S-proteins, and multiple influenza viruses (e.g., H1N1, H3N2, H5N1, H7N7, and H7N9) [85]. However, rather than blocking the binding sites needed for cell–cell interaction, P9 remained attached to the viral surface as it entered the host cell. Once inside the endosome, the polycationic properties of P9 resulted in a basic microenvironment rather than the acidic microenvironment, which is vital for these viruses to release their genetic material. The basic pH prevented viral membrane fusion with the host endosomal membrane, thus preventing viral RNA release. ACE2 receptor. Rather than targeting structures on the viral surface, the ACE2 receptor of host cells also can be targeted, as this would also prevent viral entry. An AVP targeting the host cell receptor required for viral binding and entry has already been approved for treating HIV-1 [86]. Maraviroc, an HIV-1 entry and fusion inhibiting AVP, was able to prevent HIV-1 entry into host cells by selectively binding to human chemokine receptor CCR5, which directly interferes with the crucial step of HIV-1 gp 120 binding to CCR5. Similarly, a hexapeptide Tyr-Lys-Tyr-Arg-Tyr-Leu designed from a naturally occurring hexapeptide found in the RBD of SARS-CoV was shown to reduce viral infection in vitro using epithelial cell lines [87]. Based on the results, this AVP bound to host cell receptors and interfered with necessary interactions for viral attachment. However, the activity of this hexapeptide against SARS-CoV-2 has not yet been examined. Next, HD5 peptide is a peptide secreted by Paneth cells in the crypts of Lieberkühn that has a high affinity toward ACE2 receptors [74]. The molecular dynamic simulation was used to show that HD5 is competitively bound to ligand-binding domains on the ACE2 receptor. Additionally, it was found that HD5 and ACE2 binding occurred through the formation of multiple hydrogen bonds to confer protection against SARS-CoV-2 that would otherwise bind via the free ACE2 receptor. Host cell proteases. Due to the crucial role of proteolytic cleavage in host cell entry, TMPRSS2 also has been targeted and shown to effectively inhibit SARS-CoV-2 entry into host cells [28]. An abundant naturally occurring serine protease inhibitor, α1-antitrypsin, was shown to prevent SARS-CoV-2 entry into host cells by inhibiting the protease activity of TMPRSS2, leaving the S2 domain uncleaved and therefore unable to undergo membrane fusion [88]. Other than SARS-CoV-2, TMPRSS2 is also an important host cell protease in the replication of influenza A [89]. The role of TMPRSS2 in several viral infections would allow AVPs that target the host cell protease to have broad-spectrum activity against respiratory viruses. Bestle et al. demonstrated the role of peptide mimetic inhibitors of TMPRSS2 (MI-432 and MI-1900), that could successfully inhibit SARS-CoV-2 from causing infection in human airway cells [90]. Interestingly, the antiviral activity was improved when both TMPRSS2 inhibitors were combined with the furin inhibitor MI-1851, indicating potential synergistic effects of peptides that target different cleavage sites. Another host protease that can be targeted is CatB/L, which is a lysosomal cysteine protease that plays a role in SARS-CoV-2 host cell entry via the endosomal pathway [28, 91]. CatB/L is required for S1 subunit cleavage prior to fusion of host cell endosomal and viral membrane. Therefore, the release of viral genetic material can be inhibited by blocking CatB/L [92]. Other than designing peptides that competitively bind to the CatB/L protease, peptides that inhibit endosomal acidification can also be used to inhibit the activity of CatB/L and other host cell proteases that is pH-dependent [93, 94]. Zhao et al. (2020) reported the potent broad-spectrum activity of P9R against enveloped viruses. The mechanism of action used by this AVP is the inhibition of endosomal acidification by reducing protons within the endosome [95]. The results showed that P9R, which was modified to have a more positive charge, had higher SARS-CoV-2 specific antiviral activity than the previously tested P9, which was also designed to bind to viral glycoproteins and prevent endosomal acidification [85, 95]. In addition to inhibiting either TMPRSS2 or CatB/L using antiviral peptides, Padmanabhan et al. (2020) showed better antiviral activity and complete inhibition of viral entry when both proteases were targeted simultaneously [96]. Targeting both TMPRSS2 and CatB/L have been suggested to be a more effective method in preventing viral infection, as it inhibits both routes of entry that can be used by SARS-CoV-2 to enter host cells. This is particularly more advantageous than targeting a single route of entry, as the virus particles can enter host cells via another route. It has been reported that the use of AVPs as therapeutics is limited due to their instability, short half-life, and easy degradability, all of which lead to poor bioavailability. On top of that, they are also associated with having low potency and poor ability to cross membrane barriers [79]. Particularly, naturally derived AVPs are unstable and have demonstrated weak binding affinity to their targets, sensitive to conformational changes in response to environmental stimuli, limiting the ability to cross membranes due to large size, and poorly excreted [97]. Thus, these physicochemical properties may limit their clinical application. Besides, AVPs have limited systemic delivery that may prevent them from reaching the target site with the correct dose due to the presence of numerous proteases, as well as rapid clearance from blood circulation by opsonization and agglutination [96]. These limitations can be resolved using nanoformulations (Fig. 3). For instance, nanoparticles (NPs) are known for having physical properties that can be manipulated, making them optimized drug carriers. Advantages of using NPs include protecting peptides from proteolytic and enzymatic degradation, prolonged bioavailability via sustained peptide release, and preventing peptides from premature clearance, all of which extend their bioavailability. Furthermore, NP encapsulation can be used to enhance the delivery of hydrophobic or insoluble AVPs. Systemic side effects induced by peptide-based therapeutics also can be reduced using NP encapsulation, as it can ensure targeted drugs to release in a controlled and consistent manner. Examples of nanocarriers that can be used include polymeric NPs, such as poly(lactic-co-glycolic acid) (PLGA), poly(lactic acid) (PLA), chitosan, liposomes, and micelles [98]. These NPs can be selected or modified to ensure they are non-toxic, non-immunogenic, biodegradable, and suitable for peptide and protein encapsulation. On top of that, their size, surface area, stimuli-responsiveness, and surface charge also can be modified to enhance their delivery to specific target sites and further improve the efficacy of antiviral peptides [99]. Among different types of NP-encapsulated AVPs, polymeric peptides have demonstrated antiviral activity against influenza, herpes simplex virus, human papillomavirus, respiratory syncytial virus, dengue, and lentivirus [100, 101]. SARS-CoV-2 virus transmission mechanism. SARS-CoV-2 is airborne, and once inhaled, the virus's spike proteins may have a strong affinity for the mucins in the mucus that lines the airways. Nanoparticles can be used to load drugs and AVPs against SARS-CoV-2 due to their capacity to shield the cargo from proteolytic and enzymatic degradation, as well as prolonging and increasing its bioavailability Using NPs as drug carriers also may potentially allow for non-conventional/needle-free administration, such as intranasal administration [99, 102]. The intranasal route is favorable for drug administration, as it eliminates the need for trained healthcare professionals to deliver the vaccine and reduces the cost of immunization programs, making it easily accessible to third-world countries and regions with low socioeconomic status [103]. Furthermore, ease of drug administration has been associated with increased patient compliance [104]. Additionally, the main route of entry used by SARS-CoV-2 includes the airways, from which they travel to the lungs prior to entering the systemic circulation. In this regard, AVPs against SARS-CoV-2 can be administered intranasally, as the nasal mucosa provides a large surface area for nanoparticle absorption for both local and systemic delivery of peptides [105]. Intranasal delivery of AVPs was shown to be effective against respiratory infections caused by MERS in animal models. The use of HR2P-M2, a MERS fusion inhibitor was shown to exhibit high efficacy as both a prophylactic and therapeutic [106]. It was also demonstrated that intranasal application of EK1C4 to mice could protect them from pre- and post-human coronavirus OC43 infections, indicating that it has both prophylactic and therapeutic effects against currently circulating SARS-CoV-2 and other emerging SARS-related coronaviruses [63]. Comparatively, there is a relatively higher risk for viruses to develop resistance as compared to AVPs designed to target the host cell proteins [62] due to the fact that a high rate of mutations may occur in the viral genome [57]. It has been reported that therapeutics application of AVPs is limited due to their instability, short half-life, easy degradability, and poor bioavailability. To overcome these limitations, nanoformulation approaches have emerged as a promising biological strategy to improve their distribution and stability against SARS-CoV-2, particularly, using intranasal and pulmonary delivery, and loading into PLGA nanoparticles. NP-encapsulated AVPs have shown encouraging results for both preclinical and clinical applications [110]. Eleven known accessory proteins have been identified, but their biological and regulatory roles in SARS-CoV-2 pathogenesis are still largely unknown. Thus, it is necessary to develop AVPs targeting them and evaluate their associated anti-SARS CoV-2 activities and molecular mechanisms. Accumulating studies demonstrate that non-coding RNAs, particularly, long non-coding RNAs, are essential regulators of SARS-CoV-2 infection by affecting viral gene expression, replication, and pathology in the host cells, as well as to evade the immune response of host cells [108, 112,113,114]. Besides, further exploration of potential natural or biological sources such as cyclotides (plant-derived AVPs) which could exert broad-spectrum or unique promising antiviral effects with less associated adverse effects and toxicity should be conducted. The current studies investigating the potential application of NPs to deliver AVPs for treating COVD-19 are limited, thus more investigations for their preclinical efficacy and pharmacokinetic profiles should be performed to enable clinical translation. Interestingly, the application of NPs to deliver AVPs allows them to be structurally and functionally versatile, which could serve as the molecular template for the development of advanced therapeutic applications in the face of the current pandemic threat. Additionally, the potential application of aptamers, a type of bio-inspired receptor comprised of single-stranded DNA or RNA, should also be considered as an approach to deliver therapeutic AVPs or formulated as a SARS-CoV-2 targeting agent. For instance, AVP-based aptamers can bind to identical amino acids of RBD, thus providing a promising biological tool for COVID-19 prevention, and treatment [115].SARS-CoV-2
Origin, genome, and structure
Viral target identification and validation of antiviral peptides
SARS-CoV-2-specific targets for AVPs
Targeting the SARS-CoV-2 S-protein
Antiviral peptides targeting host cell targets
Efficient delivery of potential SARS-COV-2 using nanoformulations
Data availability statement
Data sharing is not applicable to this article as no datasets were generated or analysed during the current study.
Abbreviations
- AMPs:
-
Antimicrobial peptides
- ARDS:
-
Acute respiratory distress syndrome
- AVPs:
-
Antiviral peptides
- BSL:
-
Bulged stem loop
- CDC:
-
Centre for disease control
- COVID-19:
-
Coronavirus disease 2019
- EUA:
-
Emergency Use Authorization
- FP:
-
Fusion peptide
- HIV:
-
Human immunodeficiency virus
- HVR:
-
Hyper-variable region
- MERS:
-
Middle Eastern respiratory syndrome
- MPs:
-
Microparticles
- NPs:
-
Nanoparticles
- NSPs:
-
Non-structural proteins
- PLA:
-
Poly(lactic acid)
- PLGA:
-
Poly(lactic-co-glycolic acid)
- RBD:
-
Receptor binding domain
- SARS:
-
Severe acute respiratory syndrome
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- TM:
-
Transmembrane
- US-FDA:
-
United States Food and Drug Agency
- UTR:
-
Untranslated region
References
Hui DC, Lee N, Chan PKS. A clinical approach to the threat of emerging influenza viruses in the Asia-Pacific region. Respirology. 2017;22(7):1300–12.
Marston BJ, Dokubo EK, van Steelandt A, Martel L, Williams D, Hersey S, et al. Ebola response impact on Public Health Programs West Africa 2014–2017. Emerg Infect Dis. 2017;23(13):S25–32.
Suijkerbuijk AWM, Swaan CM, Mangen M-JJ, Polder JJ, Timen A, Ruijs WLM. Ebola in the Netherlands 2014–2015: costs of preparedness and response. Eur J Health Econ. 2018;19(7):935–43.
de Souza WV, Vazquez E, Bezerra LCA, Mendes ADCG, Lyra TM, de Araujo TVB, et al. Microcephaly epidemic related to the Zika virus and living conditions in Recife Northeast Brazil. BMC Public Health. 2018;18(1):1–7.
Lowe R, Barcellos C, Brasil P, Cruz OG, Honório NA, Kuper H, Carvalho MS. The Zika virus epidemic in Brazil: From discovery to future implications. Int J Environ Res Public Health. 2018;15(1):96.
Lai C-C, Shih T-P, Ko W-C, Tang H-J, Hsueh P-R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents. 2020;55(3): 105924.
Coronavirus WHO Dashboard. WHO coronavirus (COVID-19) dashboard with vaccination data 2021. https://covid19.who.int.
ACS know your treatment options for COVID-19. U.S Food and Drug Administration. https://www.fda.gov/consumers/consumer-updates/know-your-treatment-options-covid-19.
Krammer F. SARS-CoV-2 vaccines in development. Nature. 2020;586(7830):516–27.
Forchette L, Sebastian W, Liu T. A comprehensive review of COVID-19 virology, vaccines, variants, and therapeutics. Curr Med Sci. 2021;41(6):1037–105.
Narayanan DKL, Kayarohanam S, Fuloria S, Fuloria NK, Janakiraman AK, Djearamane S, et al. Covid-19 vaccine candidates under clinical evaluation—a review. Int J Pharm Res. 2021;13(1):4588–98.
Subramaniyan V, Chakravarthi S, Seng WY, Kayarohanam S, Fuloria NK, Fuloria S. Impact of COVID-19 in public health: prevalence and preventive approaches. Pak J Pharm Sci. 2020;33(4):1739–45.
COVID-19 vaccines with WHO emergency use listing. World Health Organization. https://extranet.who.int/pqweb/vaccines/vaccinescovid-19-vaccine-eul-issued.
Costanzo M, De Giglio ARM, Roviello NG. SARS-CoV-2: recent reports on antiviral therapies based on lopinavir/ritonavir darunavir/umifenovir hydroxychloroquine remdesivir favipiravir and other drugs for the treatment of the new coronavirus. Curr Med Chem. 2020;27(27):4536–41.
Coronavirus (COVID-19) Update: FDA authorizes first oral antiviral for treatment of COVID-19. https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-authorizes-first-oral-antiviral-treatment-covid-19.
Huang Y, Yang C, Xu X-F, Xu W, Liu S-W. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin. 2020;41(9):1141–9.
Li Q, Zhao Z, Zhou D, Chen Y, Hong W, Cao L, et al. Virucidal activity of a scorpion venom peptide variant mucroporin-M1 against measles SARS-CoV and influenza H5N1 viruses. Peptides. 2011;32(7):1518–25.
Wohlford-Lenane CL, Meyerholz DK, Perlman S, Zhou H, Tran D, Selsted ME, McCray PB Jr. Rhesus theta-defensin prevents death in a mouse model of severe acute respiratory syndrome coronavirus pulmonary disease. J Virol. 2009;83(21):11385–90.
**a S, Yan L, Xu W, Agrawal AS, Algaissi A, Tseng CT, Wang Q, Du L, Tan W, Wilson IA, Jiang S. A pan-coronavirus fusion inhibitor targeting the HR1 domain of human coronavirus spike. Sci Adv. 2019;5(4):eaav4580.
**a S, Liu M, Wang C, Xu W, Lan Q, Feng S, Qi F, Bao L, Du L, Liu S, Qin C. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 2020;30(4):343–55.
Wang C, Wang S, Li D, Wei DQ, Zhao J, Wang J. Human intestinal defensin 5 inhibits SARS-CoV-2 invasion by cloaking ACE2. Gastroenterology. 2020;159(3):1145–7.
Zhao H, Zhou J, Zhang K, Chu H, Liu D, Poon VK, Chan CC, Leung HC, Fai N, Lin YP, Zhang AJ. A novel peptide with potent and broad-spectrum antiviral activities against multiple respiratory viruses. Sci Rep. 2016;6(1):1–3.
Maiti BK. Potential role of peptide-based antiviral therapy against SARS-CoV-2 infection. ACS Pharmacol Transl Sci. 2020;3(4):783–5.
Xu X, Ong YK, Wang DY. Role of adjunctive treatment strategies in COVID-19 and a review of international and national clinical guidelines. Mil Med Res. 2020;7(1):22.
Thomas S. The structure of the membrane protein of SARS-CoV-2 resembles the sugar transporter SemiSWEET. Pathog Immun. 2020;5(1):342–63.
Sarkar M, Saha S. Structural insight into the role of novel SARS-CoV-2 E protein: a potential target for vaccine development and other therapeutic strategies. PLoS ONE. 2020;15(8): e0237300.
Cubuk J, Alston JJ, Incicco JJ, Singh S, Stuchell-Brereton MD, Ward MD, et al. The SARS-CoV-2 nucleocapsid protein is dynamic disordered and phase separates with RNA. Nat Commun. 2021;12(1):1936.
Hoffmann M, Kleine-Weber H, Pöhlmann SA. Multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol Cell. 2020;78(4):779-784e5.
White JM, Delos SE, Brecher M, Schornberg K. Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme. Crit Rev Biochem Mol Biol. 2008;43(3):189–219.
Bosch BJ, Martina BEE, van der Zee R, Lepault J, Haijema BJ, Versluis C, et al. severe acute respiratory syndrome coronavirus (SARS-CoV) infection inhibition using spike protein heptad repeat-derived peptides. Proc Natl Acad Sci USA. 2004;101(22):8455–60.
Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: their roles in pathogenesis. J Microbiol Immunol Infect. 2021;54(2):159–63.
Redondo N, Zaldívar-López S, Garrido JJ, Montoya M. SARS-CoV-2 accessory proteins in viral pathogenesis: knowns and unknowns. Front Immunol. 2021;12:708264–708264.
Silvas JA, Vasquez DM, Park JG, Chiem K, Allué-Guardia A, Garcia-Vilanova A, Platt RN, Miorin L, Kehrer T, Cupic A, Gonzalez-Reiche AS. Contribution of SARS-CoV-2 accessory proteins to viral pathogenicity in K18 human ACE2 transgenic mice. J Virol. 2021;95(17):e00402-e421.
Simmons G, Zmora P, Gierer S, Heurich A, Pöhlmann S. Proteolytic activation of the SARS-coronavirus spike protein: cutting enzymes at the cutting edge of antiviral research. Antiviral Res. 2013;100(3):605–14.
Abdelmoaty MM, Yeapuri P, Machhi J, Olson KE, Shah** F, Zhou Y, et al. Defining the immune responses for SARS-CoV-2–human macrophage interactions. Front Immunol. 2021;12: 741502.
Fajgenbaum DC, June CH. Cytokine storm. New Eng J Med. 2020;383(23):2255–73.
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan China. Lancet. 2020;395(10223):497–506.
Ruan Q, Yang K, Wang W, Jiang L, Song J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan China. Intensive Care Med. 2020;46(5):846–8.
Gao Y, Li T, Han M, Li X, Wu D, Xu Y, et al. Diagnostic utility of clinical laboratory data determinations for patients with the severe COVID-19. J Med Virol. 2020;92(7):791–6.
Laing AG, Lorenc A, Molino del Barrio I, Das A, Fish M, Monin L, Muñoz-Ruiz M, et al. A dynamic COVID-19 immune signature includes associations with poor prognosis. Nat Med. 2020;26(10):1623–35.
Bojkova D, Klann K, Koch B, Widera M, Krause D, Ciesek S, et al. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature. 2020;583(7816):469–72.
Varnaitė R, García M, Glans H, Maleki KT, Sandberg JT, Tynell J, et al. Expansion of SARS-CoV-2-specific antibody-secreting cells and generation of neutralizing antibodies in hospitalized COVID-19 patients. J Immunol. 2020;05(9):2437–46.
To KK-W, Tsang OT-Y, Leung W-S, Tam AR, Wu T-C, Lung DC, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20(5):565–74.
Sterlin D, Mathian A, Miyara M, Mohr A, Anna F, Claër L, et al. IgA dominates the early neutralizing antibody response to SARS-CoV-2. Sci Transl Med. 2021;13(577):eabd2223.
Sattler A, Angermair S, Stockmann H, Heim KM, Khadzhynov D, Treskatsch S, et al. SARS-CoV-2-specific T cell responses and correlations with COVID-19 patient predisposition. J Clin Investig. 2020;130(12):6477–89.
Grifoni A, Weiskopf D, Ramirez SI, Mateus J, Dan JM, Moderbacher CR, et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell. 2020;181(7):1489-1501e15.
Severa M, Diotti RA, Etna MP, Rizzo F, Fiore S, Ricci D, Iannetta M, et al. Differential plasmacytoid dendritic cell phenotype and type I interferon response in asymptomatic and severe COVID-19 infection. PLoS Pathog. 2021;17(9): e1009878.
Diao B, Wang C, Tan Y, Chen X, Liu Y, Ning L, et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front Immunol. 2020;11(827).
Cecchini R, Cecchini AL. SARS-CoV-2 infection pathogenesis is related to oxidative stress as a response to aggression. Med Hypotheses. 2020;143: 110102.
Short KR, Kroeze EJBV, Fouchier RAM, Kuiken T. Pathogenesis of influenza-induced acute respiratory distress syndrome. Lancet Infect Dis. 2014;14(1):57–69.
Hu B, Guo H, Zhou P, Shi Z-L. Characteristics of SARS-CoV-2 and COVID-19. Nat Rev Microbiol. 2021;19(3):141–54.
Adhikari SP, Meng S, Wu Y-J, Mao Y-P, Ye R-X, Wang Q-Z, et al. Epidemiology causes clinical manifestation and diagnosis prevention and control of coronavirus disease (COVID-19) during the early outbreak period: a sco** review. Infect Diseas Poverty. 2020;9(1):29.
Subramaniyan V, Fuloria S, Darnal H, Meenakshi D, Sekar M, Nordin R, et al. COVID-19-associated mucormycosis and treatments. Asian Pac J Trop Med. 2021;14(9):401–9.
Machhi J, Herskovitz J, Senan AM, Dutta D, Nath B, Oleynikov MD, et al. The natural history pathobiology and clinical manifestations of SARS-CoV-2 infections. J Neuroimmune Pharmacol. 2020;15(3):359–86.
Tu Y-F, Chien C-S, Yarmishyn AA, Lin Y-Y, Luo Y-H, Lin Y-T, et al. A Review of SARS-CoV-2 and the Ongoing Clinical Trials. Int J Mol Sci. 2020;21(7):2657.
Kordzadeh-Kermani E, Khalili H, Karimzadeh I. Pathogenesis clinical manifestations and complications of coronavirus disease 2019 (COVID-19). Future Microbiol. 2020;15(13):1287–305.
Heydari H, Golmohammadi R, Mirnejad R, Tebyanian H, Fasihi-Ramandi M, Moosazadeh MM. Antiviral peptides against Coronaviridae family: a review. Peptides. 2021;139: 170526.
Usmani SS, Bedi G, Samuel JS, Singh S, Kalra S, Kumar P, et al. THPdb: database of FDA-approved peptide and protein therapeutics. PLoS ONE. 2017;12(7): e0181748.
Agarwal G, Gabrani R. Antiviral peptides: identification and validation. Int J Pept Res Ther. 2021;27(1):149–68.
Lee H-G, Kim D-W, Park C-W. Dry powder inhaler for pulmonary drug delivery: human respiratory system approved products and therapeutic equivalence guideline. J Pharm Investig. 2018;48(6):603–61.
Donev R. Advances in protein chemistry and structural biology. New York: Academic Press; 2014.
Badani H, Garry RF, Wimley WC. Peptide entry inhibitors of enveloped viruses: the importance of interfacial hydrophobicity. Biochim Biophy Acta Biomembr. 2014;1838(9):2180–97.
Vilas Boas LCP, Campos ML, Berlanda RLA, de Carvalho NN, Franco OL. Antiviral peptides as promising therapeutic drugs. Cell Mol Life Sci. 2019;76(18):3525–42.
Kaufmann SHE, Dorhoi A, Hotchkiss RS, Bartenschlager R. Host-directed therapies for bacterial and viral infections. Nat Rev Drug Discov. 2018;17(1):35–56.
Mooney C, Haslam NJ, Pollastri G, Shields DC. Towards the improved discovery and design of functional peptides: common features of diverse classes permit generalized prediction of bioactivity. PLoS ONE. 2012;7(10): e45012.
Milan C, Timm CD. Virulence factors associated with biofilm formation by Salmonella enterica: mini-review. Sci Anim Health. 2015;3(1):94–102.
Gao Y, Cui T, Lam Y. Synthesis and disulfide bond connectivity–activity studies of a kalata B1-inspired cyclopeptide against dengue NS2B–NS3 protease. Bioorg Med Chem. 2010;18(3):1331–6.
Ireland DC, Wang CKL, Wilson JA, Gustafson KR, Craik DJ. Cyclotides as natural anti-HIV agents. Peptide Sci. 2008;90(1):51–60.
Magana M, Pushpanathan M, Santos AL, Leanse L, Fernandez M, Ioannidis A, et al. The value of antimicrobial peptides in the age of resistance. Lancet Infect Dis. 2020;20(9):e216–30.
Jesús T, Rogelio L, Abraham C, Uriel L, J-Daniel G, Alfonso M-T, et al. Prediction of antiviral peptides derived from viral fusion proteins potentially active against herpes simplex and influenza A viruses. Bioinformation. 2012;8(18):870–4.
Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm Sin B. 2020;10(5):766–88.
Jaiswal G, Kumar V. In-silico design of a potential inhibitor of SARS-CoV-2 S protein. PLoS ONE. 2020;15(10): e0240004.
Chowdhury P. In silico investigation of phytoconstituents from Indian medicinal herb ‘Tinospora cordifolia (giloy)’ against SARS-CoV-2 (COVID-19) by molecular dynamics approach. J Biomol Struct. 2021;39(17):6792–809.
Wang Q, Zhang Y, Wu L, Niu S, Song C, Zhang Z, et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell. 2020;181(4):894-904e9.
Liu S, **ao G, Chen Y, He Y, Niu J, Escalante CR, et al. Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet. 2004;363(9413):938–47.
Hollmann A, Matos PM, Augusto MT, Castanho MARB, Santos NC. Conjugation of cholesterol to HIV-1 fusion inhibitor C34 increases peptide–membrane interactions potentiating its action. PLoS ONE. 2013;8(4):e60302.
Teissier E, Penin F, Pécheur E-I. Targeting cell entry of enveloped viruses as an antiviral strategy. Molecules. 2011;16(1):221–50.
Channappanavar R, Lu L, **a S, Du L, Meyerholz DK, Perlman S, et al. Protective effect of intranasal regimens containing peptidic middle east respiratory syndrome coronavirus fusion inhibitor against MERS-CoV infection. J Infect Dis. 2015;212(12):1894–903.
Ling R, Dai Y, Huang B, Huang W, Yu J, Lu X, et al. In silico design of antiviral peptides targeting the spike protein of SARS-CoV-2. Peptides. 2020;130: 170328.
Feldman C, Anderson R. The role of co-infections and secondary infections in patients with COVID-19. Pneumonia. 2021;13(1):5.
Mukherjee S, Bhattacharyya D, Bhunia A. Host-membrane interacting interface of the SARS coronavirus envelope protein: immense functional potential of C-terminal domain. Biophys Chem. 2020;266: 106452.
Mengist HM, Dilnessa T, ** T. Structural basis of potential inhibitors targeting SARS-CoV-2 main protease. Front Chem. 2021;9:622898–622898.
Zhang L, Lin D, Sun X, Curth U, Drosten C, Sauerhering L, et al. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science. 2020;368(6489):409–12.
Ullrich S, Nitsche C. The SARS-CoV-2 main protease as drug target. Bioorg Med Chem Lett. 2020;30(17): 127377.
Zhao H, Zhou J, Zhang K, Chu H, Liu D, Poon VK-M, et al. A novel peptide with potent and broad-spectrum antiviral activities against multiple respiratory viruses. Sci Rep. 2016;6(1):22008.
Hardy WD, Gulick RM, Mayer H, Fätkenheuer G, Nelson M, Heera J, et al. Two-year safety and virologic efficacy of maraviroc in treatment-experienced patients with CCR5-tropic HIV-1 infection: 96-week combined analysis of MOTIVATE 1 and 2. J Acquir Immune Defic Syndr. 2010;55(5):558–64.
Struck A-W, Axmann M, Pfefferle S, Drosten C, Meyer B. A hexapeptide of the receptor-binding domain of SARS corona virus spike protein blocks viral entry into host cells via the human receptor ACE2. Antivir Res. 2012;94(3):288–96.
Wettstein L, Weil T, Conzelmann C, Müller JA, Groß R, Hirschenberger M, et al. Alpha-1 antitrypsin inhibits TMPRSS2 protease activity and SARS-CoV-2 infection. Nat Commun. 2021;12(1):1726.
Hatesuer B, Bertram S, Mehnert N, Bahgat MM, Nelson PS, Pöhlman S, et al. Tmprss2 is essential for influenza H1N1 virus pathogenesis in mice. PLoS Pathog. 2013;9(12): e1003774.
Bestle D, Heindl MR, Limburg H, Van Lam T, Pilgram O, Moulton H, et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci Alliance. 2020;3(9):e202000786.
Zhao J, Yang Y, Huang H, Li D, Gu D, Lu X, et al. Relationship between the ABO blood group and the coronavirus disease 2019 (COVID-19) susceptibility. Clin Infect Dis. 2020;73(2):328–31.
Muralidar S, Gopal G, Visaga AS. Targeting the viral-entry facilitators of SARS-CoV-2 as a therapeutic strategy in COVID-19. J Med Virol. 2021;93(9):5260–76.
Gomes CP, Fernandes DE, Casimiro F, da Mata GF, Passos MT, Varela P, et al. Cathepsin L in COVID-19: from pharmacological evidences to genetics. Front Cell Infect Microbiol. 2020;10:589505–589505.
Fonović M, Turk B. Cysteine cathepsins and extracellular matrix degradation. Biochim Biophys Acta Gen Subj. 2014;1840(8):2560–70.
Zhao B, Ni C, Gao R, Wang Y, Yang L, Wei J, et al. Recapitulation of SARS-CoV-2 infection and cholangiocyte damage with human liver ductal organoids. Protein Cell. 2020;11(10):771–5.
Padmanabhan P, Desikan R, Dixit NM. Targeting TMPRSS2 and cathepsin B/L together may be synergistic against SARS-CoV-2 infection. PLOS Comput Biol. 2020;16(12): e1008461.
Luong HX, Thanh TT, Tran TH. Antimicrobial peptides—advances in development of therapeutic applications. Life Sci. 2020;260: 118407.
Bruno BJ, Miller GD, Lim CS. Basics and recent advances in peptide and protein drug delivery. Ther Deliv. 2013;4(11):1443–67.
Correia-Pinto JF, Csaba N, Alonso MJ. Vaccine delivery carriers: insights and future perspectives. Int J Pharm. 2013;440(1):27–38.
Kumari A, Yadav SK, Yadav SC. Biodegradable polymeric nanoparticles based drug delivery systems. Colloids Surf B. 2010;75(1):1–18.
Lisuzzo L, Cavallaro G, Parisi F, Milioto S, Fakhrullin R, Lazzara G. Core/shell gel beads with embedded halloysite nanotubes for controlled drug release. Coatings. 2019;9(2):70.
Cagno V, Andreozzi P, D’Alicarnasso M, Jacob Silva P, Mueller M, Galloux M, et al. Broad-spectrum non-toxic antiviral nanoparticles with a virucidal inhibition mechanism Nat. Mater. 2018;17(2):195–203.
Vila A, Sánchez A, Tobío M, Calvo P, Alonso MJ. Design of biodegradable particles for protein delivery. J Control Release. 2002;78(1):15–24.
De Smet R, Allais L, Cuvelier CA. Recent advances in oral vaccine development. Hum Vaccin Immunother. 2014;10(5):1309–18.
Giudice EL, Campbell JD. Needle-free vaccine delivery. Adv Drug Del Rev. 2006;58(1):68–89.
Keller L-A, Merkel O, Popp A. Intranasal drug delivery: opportunities and toxicologic challenges during drug development. Drug Deliv Transl Res. 2021;12(4):735–57.
Ali M. Pulmonary drug delivery. In: Kulkarni VS, editor. Handbook of non-invasive drug delivery systems. Boston: William Andrew Publishing; 2010. p. 209–46.
Harush-Frenkel O, Bivas-Benita M, Nassar T, Springer C, Sherman Y, Avital A, et al. A safety and tolerability study of differently-charged nanoparticles for local pulmonary drug delivery. Toxicol Appl Pharmacol. 2010;246(1):83–90.
Al-Qadi S, Grenha A, Carrión-Recio D, Seijo B, Remuñán-López C. Microencapsulated chitosan nanoparticles for pulmonary protein delivery: In vivo evaluation of insulin-loaded formulations. J Control Release. 2012;157(3):383–90.
Emami F, Mostafavi Yazdi SJ, Na DH. Poly(lactic acid)/poly(lactic-co-glycolic acid) particulate carriers for pulmonary drug delivery. J Pharm Investig. 2019;49(4):427–42.
Pergolizzi JV Jr, Magnusson P, LeQuang JA, Breve F, Paladini A, Rekatsina M, et al. The current clinically relevant findings on COVID-19 pandemic. Anesth Pain Med. 2020;10(2):e103819–e103819.
Yang Q, Lin F, Wang Y, Zeng M, Luo M. Long noncoding RNAs as emerging regulators of COVID-19. Front Immunol. 2021;12(3076).
Hamid UZ, Sim MS, Guad RM, Subramaniyan V, Sekar M, Fuloria NK, Fuloria S, Choy KW, Fareez IM, Bonam SR, Wu YS. Molecular regulatory roles of long non-coding RNA HOTTIP: an overview in gastrointestinal cancers. Curr Mol Med. 2022; 22(6):478–490. https://doi.org/10.2174/1566524021666210806162848.
Ramli S, Sim MS, Guad RM, Gopinath SC, Subramaniyan V, Fuloria S, Fuloria NK, Choy KW, Rana S, Wu YS. Long Noncoding RNA UCA1 in Gastrointestinal Cancers: Molecular Regulatory Roles and Patterns Mechanisms and Interactions. J Oncol. 2021;1–15. https://doi.org/10.1155/2021/5519720.
Rhouati A, Teniou A, Badea M, Marty JL. Analysis of recent bio-/nanotechnologies for coronavirus diagnosis and therapy. Sensors. 2021;21(4):1485.
Funding
This review does not receive any funding. The figures and graphical abstract in this manuscript were created with BioRender.com (with the support of https://biorender.com under a paid subscription).
Author information
Authors and Affiliations
Contributions
CLP, RZE, and YSW designed the flow of the review and planned the manuscript. CLP and YSW supervised the entire work. RZE, YSW, KV, and MS collected the data, wrote the manuscript, and made the table and figures for the manuscript. YSW, RZE and KV revised the format of the manuscript. All co-authors revised the final draft and agreed to be accountable for all aspects of work to ensure integrity and accuracy. Furthermore, all authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Essa, R.Z., Wu, Ys., Batumalaie, K. et al. Antiviral peptides against SARS-CoV-2: therapeutic targets, mechanistic antiviral activity, and efficient delivery. Pharmacol. Rep 74, 1166–1181 (2022). https://doi.org/10.1007/s43440-022-00432-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43440-022-00432-6