Abstract
Grevillea robusta is an important plant in Proteaceae, decoding and understanding the chloroplast genome of G. robusta is of great theoretical significance and practical value to the genetic diversity and phylogenetic relationships of Proteaceae. In this work, the chloroplast genome of G. robusta was sequenced, characterized, and compared to the other Proteaceae species to provide chloroplast genetic resources and because the information on chloroplast genes is scarce in the Proteaceae, we also examined the affinities between G. robusta and species of various families within Proteales to determine G. robusta’s phylogenetic position. Based on the illumina sequencing data of G. robusta, the sequencing results were assembled and annotated utilizing the software tools GetOrganelle and CPGAVAS2. The chloroplast genome data for the genera Macadamia, Helicia, and Protea were obtained from the NCBI database. Subsequently, the chloroplast genomes of four genera within the Proteaceae family were subjected to analysis using various programs including MISA, REPuter, IRscope, and IQtree. The chloroplast genome of G. robusta was 158,642 bp in length and consists of 129 genes, including 84 protein-coding genes, 37 tRNA genes and 8 rRNA genes. Fifty-six simple repeat sequences were obtained from G. robusta, of which single-nucleotide repeats were the most (66.07%) and the six nucleotide repeats were the least (1).Simultaneously, the chloroplast genome of G. robusta exhibited the presence of 34 repeats, primarily consisting of palindrome repeats (16). The inverted repeat (IR) region of G. robusta did not undergo a significant contraction/expansion event, in contrast to the notable contraction observed in Protea kilimandscharica. Analysis of gene selection pressure indicated positive selection signals in the ycf1 genes. Furthermore, examination of RNA editing sites revealed the occurrence of 148 RNA editing sites within the protein-coding genes of the chloroplast genome of G. robusta, with the majority consisting of C/U editing, accounting for 54.73% of the total. Phylogenetic analysis confirmed that G. robusta belongs to Proteaceae, and grouped with Helicia and Macadamia, with a support value of 100%. The chloroplast genome of G. robusta was assembled successfully, which is closely related to the chloroplast genomes of Helicia and Macadamia, and belongs to the same clade as Proteaceae. The results of this study laid a foundation for understanding the systematic evolution of Proteaceae plants and provide rich data to support the development of molecular biological information, such as molecular markers.







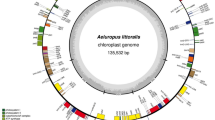
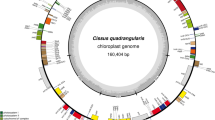
Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
Afzal M, Farman M, Rasib KZ, Qureshi NA (2019) Biocidal action of silver oak (Grevillea robusta) leaf extract on the termite Heterotermes indicola Wasmann (Blattodea: Rhinotermitidae). Int Biodeterior Biodegra 139:1–10. https://doi.org/10.1016/j.ibiod.2019.02.001
Ahmed AS, Nakamura N, Meselhy MR, Makhboul MA, El-Emary N, Hattori M (2000) Phenolic constituents from Grevillea robusta. Phytochemistry 53:149–154. https://doi.org/10.1016/S0031-9422(99)00484-7
Ahmed I, Biggs PJ, Matthews PJ, Collins LJ, Hendy MD, Lockhart PJ (2012) Mutational dynamics of aroid chloroplast genomes. Genome Biol Evol 4:1316–1323. https://doi.org/10.1093/gbe/evs110
Arseneau JR, Steeves R, Laflamme M (2017) Modified low-salt CTAB extraction of high-quality DNA from contaminant-rich tissues. Mol Ecol Resour 17:686–693. https://doi.org/10.1111/1755-0998.12616
Bhuvaneswari S, Sripriya N, Balamurugan A, Siril A, Prakash N (2014) Studies on the phytochemistry and bioefficacy of industrial crops—Coffea canephora and Gravillea robusta from Kolli hills. Int J Res Pharm Sci 5:147–151
Bock R, Khan MS (2004) Taming plastids for a green future. Trends Biotechnol 22:311–318. https://doi.org/10.1016/S0167-7799(04)00079-4
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Choi KS, Son O, Park S (2015) The chloroplast genome of Elaeagnus macrophylla and trnH duplication event in Elaeagnaceae. PLoS ONE 10:e0138727. https://doi.org/10.1371/journal.pone.0138727
Chuang TH, Wu PL (2007) Cytotoxic 5-alkylresorcinol metabolites from the leaves of Grevillea robusta. J Nat Prod 70:319–323. https://doi.org/10.1021/np0605687
Dabral A, Shamoon A, Meena RK, Kant R, Pandey S, Ginwal HS, Bhandari MS (2021) Genome skimming-based simple sequence repeat (SSR) marker discovery and characterization in Grevillea robusta. Physiol Mol Biol Plants 27:1623–1638. https://doi.org/10.1007/s12298-021-01035-w
Daniell H, Ruiz ON, Dhingra A (2005) Chloroplast genetic engineering to improve agronomic traits. Transgenic Plants Methods Protoc. https://doi.org/10.1385/1-59259-827-7:111
Emerman AB, Bowman SK, Barry A, Henig N, Patel KM, Gardner AF, Hendrickson CL (2017) NEBNext direct: a novel, rapid, hybridization-based approach for the capture and library conversion of genomic regions of interest. Curr Protoc Mol Biol 119:7–30. https://doi.org/10.1002/cpmb.39
George B, Bhatt BS, Awasthi M, George B, Singh AK (2015) Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr Genet 61:665–677. https://doi.org/10.1007/s00294-015-0495-9
Gong Z, Zhang WH, Zhang FQ, Li XY, Cheng YH, Xu WB (2013) Preliminary report on afforestation experiment of two Species of Guangdong Forestry Science and Technology 29:60–63. CNKI:SUN:GDLY.0.2013-04-013
Goulding SE, Olmstead RG, Morden CW, Wolfe KH (1996) Ebb and flow of the chloroplast inverted repeat. Mol Genet Genom 252:195–206. https://doi.org/10.1007/BF02173220
Jansen RK, Ruhlman TA (2012) Plastid genomes of seed plants. Genomics of chloroplasts and mitochondria. Springer, Dordrecht, pp 103–126. https://doi.org/10.1007/978-94-007-2920-9_5
Jia Y, Vázquez L, **aodan C, Huimin L, Hao Z, Zhanlin L, Guifang Z (2017) Development of chloroplast and nuclear DNA markers for Chinese Oaks (Quercus Subgenus Quercus) and assessment of their utility as DNA barcodes. Front Plant Sci 8:816. https://doi.org/10.3389/fpls.2017.00816
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:1–10. https://doi.org/10.1186/gb-2009-10-3-r25
Lawrie DS, Messer PW, Hershberg R, Petrov DA, Plotkin JB (2013) Strong purifying selection at synonymous sites in D. melanogaster. PLoS Genet 9:261–270. https://doi.org/10.1371/journal.pgen.1003527
Liu L, Wang Y, He P, Li P, Lee J, Soltis DE, Fu C (2018) Chloroplast genome analyses and genomic resource development for epilithic sister genera Oresitrophe and Mukdenia (Saxifragaceae), using genome skimming data. BMC Genom 19:235. https://doi.org/10.1186/s12864-018-4633-x
Lovin DD, Washington KO, deBruyn B, Hemme RR, Mori A, Epstein SR, Harker BW, Streit TG, Severson DW (2009) Genome-based polymorphic microsatellite development and validation in the mosquito Aedes aegypti and application to population genetics in Haiti. BMC Genom 10:590. https://doi.org/10.1186/1471-2164-10-590
Necşulea A, Lobry JR (2007) A new method for assessing the effect of replication on DNA base composition asymmetry. Mol Biol Evol 24:2169–2179. https://doi.org/10.1093/molbev/msm148
Petit JR, Duminil J, Fineschi S, Hampe A, Salvini D, Ven-dramin GG (2005) Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14:689–701. https://doi.org/10.1111/j.1365-294X.2004.02410.x
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277. https://doi.org/10.1016/s0168-9525(00)02024-2
Rozewicki J, Li S, Amada KM, Standley DM, Katoh K (2019) MAFFT-DASH:integrated protein sequence and structral alignment. Nucleic Acids Res 47:W5–W10. https://doi.org/10.1093/nar/gkz342
Shen J, Li X, Chen X, Huang X, ** S (2022) The complete chloroplast genome of carya cathayensis and phylogenetic analysis. Genes 13:369. https://doi.org/10.3390/genes13020369
Tae KH, Ki-Joong K, Marcel Q (2014) Chloroplast genome differences between Asian and American equisetum arvense (Equisetaceae) and the origin of the hypervariable trnY-trnE intergenic spacer. PLoS ONE 9:e103898. https://doi.org/10.1371/journal.pone.0103898
Terakami S, Matsumura Y, Kurita K, Kanamori H, Katayose Y, Yamamoto T, Katayama H (2012) Complete sequence of the chloroplast genome from pear (Pyrus pyrifolia): genome structure and comparative analysis. Tree Genet Genomes 8:841–854
Tillich M, Lehwark P, Pellizzer T, Ulbricht-Jones ES, Fischer A, Bock R, Greiner S (2017) GeSeq – versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45:W6–W11. https://doi.org/10.1093/nar/gkx391
Wen J, **e DF, Price M, Ren T, Deng YQ, Gui LJ, Guo XL, He XJ (2021) Backbone phylogeny and evolution of Apioideae (Apiaceae): new insights from phylogenomic analyses of plastome data. Mol Phylogenet Evol 161:107183. https://doi.org/10.1016/j.ympev.2021.107183
Weng QJ, Liu YC (2006) Biological characteristics and cultivation techniques of Grevillea robusta. Guangdong for Sci Technol 22:101–103. https://doi.org/10.3969/j.issn.1006-4427.2006.01.027
Xu JH, Liu Q, Hu W, Wang T, Xue Q, Messing J (2015) Dynamics of chloroplast genomes in green plants. Genomics 106:221–231. https://doi.org/10.1016/j.ygeno.2015.07.004
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556. https://doi.org/10.1093/bioinformatics/13.5.555
Acknowledgements
Thanks are due to Mr. Zuo Youwei from Southwest University for their help.
Funding
The authors did not receive support from any organization for the submitted work.
Author information
Authors and Affiliations
Contributions
All the authors participated in compiling and analyzing the data and preparing the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Liu, G., Yu, J. et al. Analysis on the complete chloroplast genome of Grevillea robusta. Braz. J. Bot 47, 133–143 (2024). https://doi.org/10.1007/s40415-023-00976-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-023-00976-8