Abstract
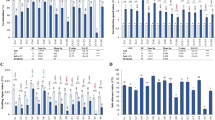
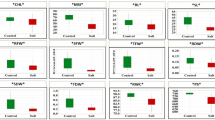
In this study, we analyzed the behavior of several neglected, ancestral, and domesticated wheat genotypes, including Ae. triuncialis, Ae. neglecta, Ae. caudata, Ae. umbellulata, Ae. tauschii, Ae. speltoides, T. boeoticum, T. urartu, T. durum, and T. aestivum under control and salinity stress to assess the mechanisms involved in salinity tolerance. Physiological and biochemical traits including root/shoot biomasses, root/shoot ion concentrations, activity of antioxidant enzymes APX, SOD, and GXP, and the relative expression of TaHKT1;5, TaSOS1, APX, GXP, and MnSOD genes were measured. Analysis of variance (ANOVA) revealed significant effects of the salinity treatments and genotypes for all evaluated traits. Salinity stress (350 mM NaCl) significantly decreased root/shoot biomasses, K+ concentration in root/shoot, and root/shoot K+/Na+ ratios. In contrast, salinity stress significantly increased Na+ concentration in root and shoot, activity of antioxidant enzymes (APX, SOD, and GPX) and relative expression of salt tolerance-related genes (TaHKT1;5, TaSOS1, APX, GPX, and MnSOD). Based on heat map and principal component analysis, the relationships among physiological traits and relative expression of salt-responsive genes were investigated. Remarkably, we observed a significant association between the relative expression of TaHKT1;5 with root K+ concentration and K+/Na+ ratio and with TaSOS1. Taken together, our study revealed that two neglected (Ae. triuncialis) and ancestral (Ae. tauschii) wheat genotypes responded better to salinity stress than other genotypes. Further molecular tasks are therefore essential to specify the pathways linked with salinity tolerance in these genotypes.





Similar content being viewed by others
Abbreviations
- SFW:
-
Shoot fresh weight
- RFW:
-
Root fresh weight
- SDW:
-
Shoot dry weight
- RDW:
-
Root dry weight
- LN:
-
Leaf Na+ concentration
- LK:
-
Leaf K+ concentration
- RN:
-
Root Na+ concentration
- RK:
-
Root K+ concentration
- LKN:
-
K+/Na+ ratio of leaf
- RKN:
-
K+/Na+ ratio of root
- APX activity:
-
Ascorbate peroxidase activity
- GPX activity:
-
Guaiacol peroxidase activity
- SOD activity:
-
Superoxide dismutase activity
- APX gene:
-
APX gene expression
- GPX gene:
-
GPX gene expression
- SOD gene:
-
MnSOD gene expression
- SOS1 gene:
-
TaSOS1 gene expression
- TaHKT1;5 gene:
-
TaHKT1;5 gene expression
References
Ahmadi J, Pour-Aboughadareh A, Fabriki Ourang S, Mehrabi AA, Siddique KHM (2018a) Screening wheat germplasm for seedling root architectural traits under contrasting water regimes: potential sources of variability for drought adaptation. Arch Agron Soil Sci 64:1351–1365
Ahmadi J, Pour-Aboughadareh A, Fabriki Ourang S, Mehrabi AA, Siddique KHM (2018b) Wild relatives of wheat: Aegilops–Triticum accessions disclose differential antioxidative and physiological responses to water stress. Acta Physiol Plant 40:90
Ahmadi J, Pour-Aboughadareh A, Fabriki-Ourang S, Mehrabi AA, Siddique KHM (2018c) Screening wild progenitors of wheat for salinity stress at early stages of plant growth: insight into potential sources of variability for salinity adaptation in wheat. Crop Pasture Sci 69:649–658
Ali AA, Alqurainy F (2006) Activities of antioxidants in plants under environmental stress. In: Motohashi N (ed) The lutein-prevention and treatment for diseases. Transworld Research Network, Trivandrum, pp 187–256
Arabbeigi M, Arzani A, Majidi MM, Kiani R, Tabatabaei BES, Habibi F (2014) Salinity tolerance of Aegilops cylindrica genotypes collected from hyper-saline shores of Uremia Salt Lake using physiological traits and SSR markers. Acta Physiol Plant 36:2243–2251
Arabbeigi M, Arzani A, Majidi MM, Seyed-Tabatabaei B, Saha P (2018) Expression pattern of salt tolerance-related genes in Aegilops cylindrica. Physiol Mol Biol Plants 24:61–73
Ariyarathna HACK, Oldach KH, Francki M (2016) A comparative gene analysis with rice identified orthologous group II HKT genes and their association with Na+ concentration in bread wheat. BMC Plant Biol 16:1–21
Assaha DVM, Ueda A, Saneoka H, Al-Yahyai R, Yaish MW (2017) The role of Na+ and K+ transporters in salt stress adaptation in glycophytes. Front Physiol 8:509
Baek KH, Skinner DZ (2003) Alteration of antioxidant enzyme gene expression during cold acclimation of near-isogenic wheat lines. Plant Sci 165:1221–1227
Blum A (2011) Drought resistance—is it really a complex trait? Funct Plant Biol 38:753–757
Bose J, Rodrigo-Moreno A, Shabala S (2014) ROS homeostasis in halophytes in the context of salinity stress tolerance. J Exp Bot 65:1241–1257
Bowler C, Van Montagu M, Inze D (1992) Superoxide dismutase and stress tolerance. Annu Rev Plant Physiol Plant Mol Biol 43:83–116
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, Tester M, Munns R (2007) HKT1;5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1. Plant Physiol 143:1918–1928
Chen GX, Asada K (1989) Ascorbate peroxidase in tea leaves: occurrence of two isozymes and the differences in their enzymatic and molecular properties. Plant Cell Physiol 30:987–998
Dhindsa RS, Plump-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased levels of membrane permeability and lipid peroxidation, and decreased levels of superoxide dismutase and catalase. J Exp Bot 32:93–101
Food Agriculture and Organization (FAO) (2015) Cereal supply and demand brief. http://www.fao.org/worldfoodsituation/csdb/en/. Accessed 20 Mar 2016
Gao J, Sun J, Cao P, Ren L, Liu L, Chen S, Chen F, Jiang J (2016) Variation in tissue Na+ concentration and the activity of SOS1 genes among two species and two related genera of Chrysanthemum. BMC Plant Biol 16:98–113
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Gong D, Guo Y, Schumaker KS, Zhu JK (2004) The SOS3 family of calcium sensors and SOS2 family of protein kinases in Arabidopsis. Plant Physiol 134:919–926
Goyal E, Amit SK, Singh RS (2016) Transcriptome profiling of the salt-stress response in Triticum aestivum cv. Kharchia Local. Sci Rep 6:27752
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. California Agricultural Experiment Station, Circular No. 374. The College of Agriculture, University of California, Berkeley, CA, USA
Horie T, Schroeder JI (2004) Sodium transporters in plants diverse genes and physiological functions. Plant Physiol 136:2457–2462
Hossain MS, Dietz KJ (2016) Tuning of redox regulatory mechanisms, reactive oxygen species and redox homeostasis under salinity stress. Front Plant Sci 7:548
Hossain MS, ElSayad AI, Moore M, Dietz KJ (2017) Redox and reactive oxygen species network in acclimation for salinity tolerance in sugar beet. J Exp Bot 68:1283–1298
Ji H, Pardo JM, Batelli G, Van Oseten MJ, Bressan RA, Li X (2013) The salt overly sensitive (SOS) pathway: established and emerging roles. Mol Plant 6:275–286
Johnson SM, Doherty SJ, Croy RRD (2003) Biphasic superoxide generation in potato tubers. A self-amplifying response to stress. Plant Physiol 13:1440–1449
Kang H-M, Saltveit ME (2002) Antioxidant enzymes and DPPH-radical scavenging activity in chilled and heat shocked rice (Oryza sativa L.) seedling radicles. J Agric Food Chem 50(3):513–518
Katschnig D, Bliek T, Rozema J, Schat H (2015) Constitutive highlevel SOS1 expression and absence of HKT1; 1 expression in the salt accumulating halophyte Salicornia dolichostachya. Plant Sci 234:144–154
Lee YP, Kim SH, Bang JW, Lee HS, Kwon SY (2007) Enhanced tolerance to oxidative stress in transgenic tobacco plants expressing three antioxidant enzymes in chloroplasts. Plant Cell Rep 26:591–598
Ludwig AA, Romeis T, Jones JD (2004) CDPK-mediated signaling pathways: specificity and cross talk. J Exp Bot 55:181–188
Maser P, Eckelmana BV, Vaidyanathan R, Horie T, Frbairn DJ, Kubo M, Yamagami M, Yamaguchi K, Nishimura M, Uozumi N, Robertson W, Mr Sussman, Schroeder JI (2002) Altered shoot/root Na+ distribution and bifurcating salt sensitivity in Arabidopsis by genetic disruption of the Na+ transporter AtHKT1. FEBS Lett 531:157–161
Mickelbart MV, Hasegawa PM, Bailey-Serres J (2005) Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16:237–251
Moller IS, Gilliham M, Jha D, Mayo GM, Roy SJ, Coates JC, Haseloff J, Tester M (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na+ transport in Arabidopsis. Plant Cell 21:2163–2178
Nakagami H, Pitzschke A, Hirt H (2005) Emerging MAP kinase pathways in plant stress signaling. Trends Plant Sci 7:339–346
Oh DH, Leidi E, Zhang Q, Hwang SM, Li Y, Quintero FH, Jiang X, Durzo MP, Lee SY, Zhao Y, Bahk JD, Bressan RA, Yun DJ, Pardo JM, Bohnert HJ (2009) Loss of halophytism by interference with SOS1 expression. Plant Physiol 151:210–222
Oyiga BC, Sharma RC, Shen J, Baum M (2016) Identification and characterization of salt tolerance of wheat germplasm using multivariable screening. J Agron Crop Sci 202:472–485
Pagnotta MA, Mondini L, Porceddu E (2009) Quantification and organization of WIS2-1A and BARE-1 retrotransposons in different genomes of Triticum and Aegilops species. Mol Genet Genomics 282:245–255
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucl Acids Res 29:e45
Pour-Aboughadareh A, Ahmadi J, Mehrabi AA, Moghaddam M, Etminan A (2017a) Physiological responses to drought stress in wild relatives of wheat: implications for wheat improvement. Acta Physiol Plant 39:106
Pour-Aboughadareh A, Mohmoudi AM, Ahmadi J, Mehrabi AA, Alavikia SS (2017b) Agro-morphological and molecular variability in Triticum boeoticum accessions from Zagros Mountains, Iran. Genet Resour Crop Evol 64:545–556
Ramezani A, Niazi A, Abolimoghadam AA, Zamani Bobgohari M, Deihimi T, Ebrahimi M, Akhtadanesh H, Ebrahimie E (2013) Quantitative expression analysis of TaSOS1 and TaSOS4 genes in cultivated and wild wheat plants under salt stress. Mol Biotechnol 53:189–197
Rampino P, Pataleo S, Gerardi C, Mita G, Perrotta C (2006) Drought stress response in wheat: physiological and molecular analysis of resistant and sensitive genotypes. Plant, Cell Environ 29:2143–2152
SAS Institute (2011) Base SAS 9.1 procedures guide. SAS Institute Inc, Cary
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279
Sharma P, Jha AB, Dubey RS, Pessarakli M (2012) Reactive oxygen species, oxidative damage and antioxidative defense mechanism in plants under stressful conditions. J Bot 217037:1–26
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Su Y, Luo W, Lin W, Ma L, Kabir MH (2015) Model of cation transportation mediated by high-affinity potassium transports (HKTs) in higher plants. Biol Proced Online 17:1–13
Suneja Y, Gupta AK, Bains NS (2017) Bread wheat progenitors: Aegilops tauschii (DD genome) and Triticum dicoccoides (AABB genome) reveal differential antioxidative response under water stress. Physiol Mol Biol Plants 23:99–114
Tester M, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot 91:503–527
Upadhyaya A, Sankhla D, Davis TD, Sankhla N, Smith BN (1985) Effect of paclobutrazol on the activities of some enzymes of activated oxygen metabolism and lipid peroxidation in senescing soybean leaves. J Plant Physiol 121:453–461
Verma KK, Singh M, Gupta RK, Verma CL (2014) Photosynthetic gas exchange, chlorophyll fluorescence, antioxidant enzymes, and growth responses of Jatropha curcas during soil flooding. Turk J Bot 38:130–140
Wang Y, Wisniewski M, Meilan R, Cui M, Webb R, Fuchigami L (2005) Overexpression of cytosolic ascorbate peroxidase in tomato confers tolerance to chilling and salt stress. J Am Soc Hortic Sci 130:167–173
Warnes GR, Bolker B, Bonebakker L, Gentlemean R, Huber W, Liaw A et al (2014) gplots: various R programming tools for plotting data. http://CRAN.R-project.org/package=gplots. Accessed 5 Feb 2019
Yan H, Li Q, Park SC, Wang X, Liu YJ, Zhang YG, Tang W, Kou M, Ma DF (2016) Overexpression of CuZnSOD and APX enhance salt stress tolerance in sweet potato. Plant Physiol Biochem 109:20–27
Yousfi S, Marquez AJ, Betti M (2016) Gene expression and physiological responses to salinity and water stress of contrasting durum wheat genotypes. J Integr Plant Biol 58:48–66
Zamani Bangohari M, Niazi A, Moghaddam AA, Deihimi T, Ebrahimie E (2013) Genome-wide analysis of key salinitytolerance transporter (HKT;5) in wheat and wild wheat relatives (A and D genomes). Vitro Cell Dev Biol Plant 49:97–106
Zhang Y, Li Y, Peng Y, Wang X, Peng D, Li Y, He X, Zhang X, Ma X, Huang L, Yan Y (2015) Clones of FeSOD, MDHAR, DHAR genes from white clover and gene expression analysis of ROS-scavenging enzymes during abiotic stress and hormone treatments. Molecules 20:20939–20954
Zhu JK (2000) Genetic analysis of plant salt tolerance using Arabidopsis. Plant Physiol 124:941–948
Zhu JK (2003) Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol 6:441–445
Zhu M, Shabala L, Cuin TA, Huang X, Zhou M, Munns R, Shabala S (2016) Nax loci affect SOS1-like Na+/H+ exchanger expression and activity in wheat. J Exp Bot 67(3):835–844
Acknowledgements
The authors acknowledge with thanks the technical and lab facilities support from the Genetic and Genomic laboratory of Department of Genetics and Plant Breeding, Imam Khomeini International University, Qazvin, Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ahmadi, J., Pour-Aboughadareh, A., Fabriki Ourang, S. et al. Unraveling salinity stress responses in ancestral and neglected wheat species at early growth stage: A baseline for utilization in future wheat improvement programs. Physiol Mol Biol Plants 26, 537–549 (2020). https://doi.org/10.1007/s12298-020-00768-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-020-00768-4