Abstract
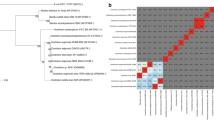
A new wild type solventogenic Clostridium bacterium, which could produce a large amount of solvent from a mixture of 70% glucose and 30% xylose, was isolated. Based on 16S rRNA gene analysis, this strain was identified as Clostridium beijerinckii. Batch fermentations with this strain (C. beijerinckii GSC1) resulted in the production of 19.65 g/L total solvents (Acetone-Butanol-Ethanol), which is 31% higher than that with the typical wild type strain Clostridium acetobutylicum ATCC 824. This new strain utilized glucose and xylose simultaneously without genetic modification. The selectivity of GSC1 for butanol was 81.8%, which is much higher than that of C. acetobutylicum ATCC 824 (68.7%). Simple genetic modification was performed to obtain a more improved performance. The acid production in batch fermentation by C. beijerinckii GSC1_R1 (gene-modified strain) was reduced to 2.6 g/L from 6.02 g/L. The solvent productivity of GSC1_R1 in continuous fermentation was 3.6 g/L/h. These results indicate that the newly isolated strain is very promising and applicable for the production of biobutanol from second-generation biomass owing to the superior performance of the wild type strain.
Similar content being viewed by others
References
Szulczyk, K. R. (2010) Which is a better transportation fuel — butanol or ethanol? Int. J. Energy Environ. 1: 501–512.
Keis, S., R. Shaheen, and D. T. Jones (2001) Emended descriptions of Clostridium acetobutylicum and Clostridium beijerinckii, and descriptions of Clostridium saccharoperbutylacetonicum sp. nov. and Clostridium saccharobutylicum sp. nov. Int. J. Syst. Evol. Microbiol. 51: 2095–2103.
Chen, C., C. Sun, and Y. R. Wu (2018) The draft genome sequence of a novel high-efficient butanol-producing bacterium Clostridium diolis strain WST. Curr. Microbiol. 75: 1011–1015.
Nasser Al-Shorgani, N. K., M. H. M. Isa, W. M. W. Yusoff, M. S. Kalil, and A. A. Hamid (2016) Isolation of a Clostridium acetobutylicum strain and characterization of its fermentation performance on agricultural wastes. Renew. Energy. 86: 459–465.
Dalal, J., M. Das, S. Joy, M. Yama, and J. Rawat (2019) Efficient isopropanol-butanol (IB) fermentation of rice straw hydrolysate by a newly isolated Clostridium beijerinckii strain C-01. Biomass Bioenergy. 127: 105292.
Li, H. G., F. K. Ofosu, K. T. Li, Q. Y. Gu, Q. Wang, and X. B. Yu (2014) Acetone, butanol, and ethanol production from gelatinized cassava flour by a new isolates with high butanol tolerance. Bioresour. Technol. 172: 276–282.
Zhang, J., W. Zhu, H. Xu, Y. Li, D. Hua, F. **, M. Gao, and X. Zhang (2016) Simultaneous glucose and xylose uptake by an acetone/butanol/ethanol producing laboratory Clostridium beijerinckii strain SE-2. Biotechnol. Lett. 38: 611–617.
Moon, H. G., Y. S. Jang, C. Cho, J. Lee, R. Binkley, and S. Y. Lee (2016) One hundred years of clostridial butanol fermentation. FEMS Microbiol. Lett. 363: fnw001.
Jang, Y. S., J. Lee, A. Malaviya, D. Y. Seung, J. H. Cho, and S. Y. Lee (2012) Butanol production from renewable biomass: Rediscovery of metabolic pathways and metabolic engineering. Biotechnol. J. 7: 186–198.
Sillers, R., A. Chow, B. Tracy, and E. T. Papoutsakis (2008) Metabolic engineering of the non-sporulating, non-solventogenic Clostridium acetobutylicum strain M5 to produce butanol without acetone demonstrate the robustness of the acid-formation pathways and the importance of the electron balance. Metab. Eng. 10: 321–332.
González-Pajuelo, M., I. Meynial-Salles, F. Mendes, J. C. Andrade, I. Vasconcelos, and P. Soucaille (2005) Metabolic engineering of Clostridium acetobutylicum for the industrial production of 1,3-propanediol from glycerol. Metab. Eng. 7: 329–336.
Kumar, M. and K. Gayen (2011) Developments in biobutanol production: New insights. Appl. Energy. 88: 1999–2012.
Gheshlaghi, R., J. M. Scharer, M. Moo-Young, and C. P. Chou (2009) Metabolic pathways of clostridia for producing butanol. Biotechnol. Adv. 27: 764–781.
Ezeji, T., N. Qureshi, and H. P. Blaschek (2007) Production of acetone-butanol-ethanol (ABE) in a continuous flow bioreactor using degermed corn and Clostridium beijerinckii. Process Biochem. 42: 34–39.
Nanda, S., D. Golemi-Kotra, J. C. McDermott, A. K. Dalai, I. Gökalp, and J. A. Kozinski (2017) Fermentative production of butanol: Perspectives on synthetic biology. N Biotechnol. 37: 210–221.
Xue, C., J. B. Zhao, L. J. Chen, F. W. Bai, S. T. Yan, and J. X. Sung (2014) Integrated butanol recovery for an advanced biofuel: current state and prospects. Appl. Microbiol. Biotechnol. 98: 3463–3474.
Jang, Y. S., A. Malaviya, C. Cho, J. Lee, and S. Y. Lee (2012) Butanol production from renewable biomass by clostridia. Bioresour. Technol. 123: 653–663.
Lee, S. Y., J. H. Park, S. H. Jang, L. K. Nielsen, J. Kim, and K. S. Jung (2008) Fermentative butanol production by clostridia. Biotechnol. Bioeng. 101: 209–228.
Heap, J. T., O. J. Pennington, S. T. Cartman, G. P. Carter, and N. P. Minton (2007) The ClosTron: A universal gene knock-out system for the genus Clostridium. J. Microbiol. Methods. 70: 452–464.
Wiesenborn, D. P., F. B. Rudolph, and E. T. Papoutsakis (1989) Coenzyme A transferase from Clostridium acetobutylicum ATCC 824 and its role in the uptake of acids. Appl. Environ. Microbiol. 55: 323–329.
Lee, S. H., S. Kim, J. Y. Kim, N. Y. Cheong, and K. H. Kim (2016) Enhanced butanol fermentation using metabolically engineered Clostridium acetobutylicum with ex situ recovery of butanol. Bioresour. Technol. 218: 909–917.
Lee, S. H., M. H. Eom, S. Kim, M. A. Kwon, J. D. R. Choi, J. Kim, Y. A. Shin, and K. H. Kim (2015) Ex situ product recovery and strain engineering of Clostridium acetobutylicum for enhanced production of butanol. Process Biochem. 50: 1683–1691.
Mermelstein, L. D., N. E. Welker, G. N. Bennett, and E. T. Papoutsakis (1992) Expression of cloned homologous fermentative genes in Clostridium acetobutylicum ATCC 824. Nat. Biotechnol. 10: 190–195.
Choi, J., Y. S. Jang, J. H. Cho, D. Seung, S. Y. Lee, E. T. Papoutsakis, G. N. Bennett, and H. Song (2013) Characterization and evaluation of corn steep liquid in acetone-butanol-ethanol production by Clostridium acetobutylicum. Biotechnol. Bioprocess Eng. 18: 266–271.
Nolling, J., G. Breton, M. V. Omelchenko, K. S. Makarova, Q. Zeng, R. Gibson, H. M. Lee, J. Dubois, D. Qiu, J. Hitti, Y. I. Wolf, R. L. Tatusov, F. Sabathe, L. Doucette-Stamm, P. Soucaille, M. J. Daly, G. N. Bennett, E. V. Koonin, and D. R. Smith (2001) Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J. Bacteriol. 183: 4823–4838.
Wang, Y., X. Li, Y. Mao, and H. P. Blaschek (2012) Genome-wide dynamic transcriptional profiling in Clostridium beijerinckii NCIMB 8052 using single-nucleotide resolution RNA-Seq. BMC Genomics. 13: 102.
Acknowledgements
This project (Project No. 2013001580001) is supported by the Ministry of Environment, Republic of Korea as “The Wastes to Energy Technology Development Program”.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
No ethical approval required.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Conflicts of Interest
The authors declare no conflict of interest.
Informed Consent
No informed consent required.
Rights and permissions
About this article
Cite this article
Shin, YA., Choi, S. & Han, M. Simultaneous Fermentation of Mixed Sugar by a Newly Isolated Clostridium beijerinckii GSC1. Biotechnol Bioproc E 26, 137–144 (2021). https://doi.org/10.1007/s12257-020-0183-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-020-0183-6