Abstract
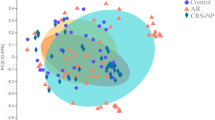
To evaluate the composition of the microbial community of the middle nasal in paranasal sinus fungus ball (FB), chronic sinusitis with nasal polyps (CRSwNP) and healthy controls, providing new insights into the pathogenesis of FB and CRSwNP. Through 16 s rRNA gene high-throughput sequencing to determine the microbial characterization from patients with FB (n = 29) and CRSwNP (n = 10), and healthy controls (n = 4). The FB group had significantly lower αdiversity and significantly different β diversity compared to the other groups. All three groups mainly consisted of four bacterial phyla (Firmicutes, Proteobacteria, Bacteroidetes, Actinobacteria). In the FB group, the highest relative abundance was found in Proteobacteria (47.04%). However, pairwise comparisons resulted in statistically significant differences only for Firmicutes (CRSwNP, p = 0.003, Control, p = 0.008). The CRSwNP group was statistically different from the control group in TM7(p = 0.010), Chloroflexi(p = 0.018) and Bacteroidete(p = 0.027). At the genus level, the FB group had the highest relative abundance of Haemophilus (11.53%), followed by Neisseria (7.39%), and Neisseria abundance (p < 0.001) was significantly different from the remaining two groups. Ruminococcacea abundance (p < 0.001) and Comamonadaceae abundance (p < 0.001) were increased in the CRSwNP group. The relative abundance of Lactobacillus (p < 0.001), Bacteroides S24_7 (p < 0.001), and Desulfovibrio (p < 0.001) was significantly decreased in the FB and CRSwNP groups compared to the control group. The imbalance of the microbial community is related to the pathogenesis of sinusitis.





Similar content being viewed by others
References
Ferreiro JA, Carlson BA, Cody DT 3rd (1997) Paranasal sinus fungus balls. Head Neck 19:481–486. https://doi.org/10.1002/(sici)1097-0347(199709)19:6%3c481::aid-hed4%3e3.0.co;2-v
Durrani Z, Naeem M, Khan MA (2018) Assessment of bacterial infection in patients operated for complications of chronic fungal rhinosinusitis. Pak J Med Health Scis 12:153–156
Lou H, Meng Y, Piao Y et al (2016) Cellular phenoty** of chronic rhinosinusitis with nasal polyps. Rhinology 54:150–159. https://doi.org/10.4193/Rhino15.271
Guarner F, Malagelada JR (2003) Gut flora in health and disease. Lancet 361:512–519. https://doi.org/10.1016/S0140-6736(03)12489-0
Maynard CL, Elson CO, Hatton RD, Weaver CT (2012) Reciprocal interactions of the intestinal microbiota and immune system. Nature 489:231–241. https://doi.org/10.1038/nature11551
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI (2007) The human microbiome project. Nature 449:804–810. https://doi.org/10.1038/nature06244
Kim DW, Kim YM, Min JY et al (2020) Clinicopathologic characteristics of paranasal sinus fungus ball: retrospective, multicenter study in Korea. Eur Arch Otorhinolaryngol 277:761–765. https://doi.org/10.1002/lary.27726
Agostinis C, Mangogna A, Bossi F, Ricci G, Kishore U, Bulla R (2019) Uterine Immunity and Microbiota: A Shifting Paradigm. Front Immunol 17:2387. https://doi.org/10.3389/fimmu.2019.02387
He XS, Shi WY (2009) Oral microbiology: past, present and future. Int J Oral Sci 1:47–58. https://doi.org/10.4248/ijos.09029
Gan W, Zhang H, Yang F et al (2021) The influence of nasal bacterial microbiome diversity on the pathogenesis and prognosis of chronic rhinosinusitis patients with polyps. Eur Arch Otorhinolaryngol 278:1075–1088. https://doi.org/10.1007/s00405-020-06370-4
Rom D, Bassiouni A, Eykman E et al (2019) The association between disease severity and microbiome in chronic Rhinosinusitis. Laryngoscope 129:1265–1273
Wagner Mackenzie B, Waite DW, Hoggard M et al (2017) Bacterial community collapse: a meta-analysis of the sinonasal microbiota in chronic rhinosinusitis. Environ Microbiol 19:381–392. https://doi.org/10.1111/1462-2920.13632
Stammberger H, Jakse R, Beaufort F (1984) Aspergillosis of the paranasal sinuses x-ray diagnosis, histopathology, and clinical aspects. Ann Otol Rhinol Laryngol 93:251–256. https://doi.org/10.1177/000348948409300313
Qaisar Sajjad SM, Suhail Z, Ahmed R (2020) Prevalence of fungal infection in nasal polyposis - A cross-sectional study, conducted at a tertiary care hospital in Karachi. J Pak Med Assoc 70:48–52. https://doi.org/10.5455/JPMA.296507
Abreu NA, Nagalingam NA, Song Y et al (2020) Sinus microbiome diversity depletion and Corynebacterium tuberculostearicum enrichment mediates rhinosinusitis. Sci Transl Med 4:151ra124. https://doi.org/10.1126/scitranslmed.3003783
Wagner Mackenzie B, West AG, Waite DW et al (2020) A novel description of the human sinus archaeome during health and chronic Rhinosinusitis. Front Cell Infect Microbiol 10:398. https://doi.org/10.3389/fcimb.2020.00398
Lai JC, Lee HS, Chen MK et al (2011) Patient satisfaction and treatment outcome of fungus ball rhinosinusitis treated by functional endoscopic sinus surgery. Eur Arch Otorhinolaryngol 268:227–230. https://doi.org/10.1007/s00405-010-1299-7
Gan W, Yang F, Meng J et al (2021) Comparing the nasal bacterial microbiome diversity of allergic rhinitis, chronic rhinosinusitis and control subjects. Eur Arch Otorhinolaryngol 278:711–718. https://doi.org/10.1007/s00405-020-06311-1
Wu Y, Chen Y, Li Q et al (2021) Tetrahydrocurcumin alleviates allergic airway inflammation in asthmatic mice by modulating the gut microbiota. Food Funct 12:6830–6840. https://doi.org/10.1039/d1fo00194a
Zhong Y, Li Y, **ao X (2014) The relationship of the expression of thymic stromal lymphopoietin in nasal polyps tissues and Th2 inflammatory response. Lin Chung Er Bi Yan Hou Tou **g Wai Ke Za Zhi 28:817–820
Salvetti E, O’Toole PW (2017) When regulation challenges innovation: the case of the genus Lactobacillus. Trends Food Sci Technol 66:S0924224416300759. https://doi.org/10.1016/j.tifs.2017.05.009
De Boeck I, van den Broek MFL, Allonsius CN et al (2020) Lactobacilli Have a Niche in the Human Nose. Cell Rep 31:107674. https://doi.org/10.1016/j.celrep.2020.107674
Lagkouvardos I, Lesker TR, Hitch TCA et al (2019) Sequence and cultivation study of Muribaculaceae reveals novel species, host preference, and functional potential of this yet undescribed family. Microbiome 7:28. https://doi.org/10.1186/s40168-019-0637-2
Baraniuk JN, Petrie KN, Le U et al (2005) Neuropathology in rhinosinusitis. Am J Respir Crit Care Med 171:5–11. https://doi.org/10.1164/rccm.200403-357OC
Pereira FC, Wasmund K, Cobankovic I et al (2020) Rational design of a microbial consortium of mucosal sugar utilizers reduces Clostridiodes difficile colonization. Nat Commun 11:5104. https://doi.org/10.1038/s41467-020-18928-1
Castillo Vizuete JA, Sastre J, Del Cuvillo BA et al (2019) Asthma, rhinitis, and nasal polyp multimorbidities. Arch Bronconeumol (Engl Ed) 55:146–155. https://doi.org/10.1016/j.arbres.2018.09.001
Weyand NJ (2017) Neisseria models of infection and persistence in the upper respiratory tract. Pathog Dis 75:ftx031. https://doi.org/10.1093/femspd/ftx031
Adriaens LM, Alessandri R, Spörri S et al (2009) Does pregnancy have an impact on the subgingival microbiota? J Periodontol 80:72–81. https://doi.org/10.1902/jop.2009.080012
Rodríguez-Rojas L, Suarez-López A, Cantón R et al (2020) Eikenella corrodens causing deep-seated infections Six-year experience in a University Hospital in Madrid. Enferm Infecc Microbiol Clin (Engl Ed) 38(2):76–78. https://doi.org/10.1016/j.eimc.2019.04.004
Peleg AY, Hogan DA, Mylonakis E (2010) Medically important bacterial-fungal interactions. Nat Rev Microbiol 8:340–349. https://doi.org/10.1038/nrmicro2313
Kroll JS, Wilks KE, Farrant JL et al (1998) Natural genetic exchange between Haemophilus and Neisseria: intergeneric transfer of chromosomal genes between major human pathogens. Proc Natl Acad Sci U S A 95:12381–12385. https://doi.org/10.1073/pnas.95.21.12381
Acknowledgements
The research was funded by the National Natural Science Foundation of China (grant award number 81070768) and Wu Jie** Clinical Research Special Assistance Fund (grant award number 320.6750); National Key Research & Development Program of China (2017YFC0112500), Bei**g Municipal Administration of Hospitals’ Mission Plan (SML20150201).
Ethical Approval
Ethical issues (Including plagiarism, informed consent, misconduct, data fabrication and/or falsification, double publication and/or submission, redundancy, etc.) have been completely observed by the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, P., Wei, H., Liu, X. et al. The Nasal Bacteria Microbiome Comparison Among Fungal Ball Sinusitis, Chronic Sinusitis with Polyps. Indian J Microbiol 63, 120–128 (2023). https://doi.org/10.1007/s12088-023-01062-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-023-01062-z