Abstract
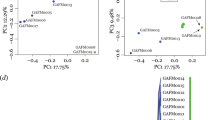
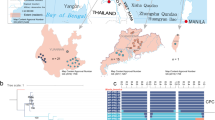
The Chinese pangolin (Manis pentadactyla, MP) has been extensively exploited and is now on the brink of extinction, but its population structure, evolutionary history, and adaptive potential are unclear. Here, we analyzed 94 genomes from three subspecies of the Chinese pangolin and identified three distinct genetic clusters (MPA, MPB, and MPC), with MPB further divided into MPB1 and MPB2 subpopulations. The divergence of these populations was driven by past climate change. For MPB2 and MPC, recent human activities have caused dramatic population decline and small population size as well as increased inbreeding, but not decrease in genomic variation and increase in genetic load probably due to strong gene flow; therefore, it is crucial to strengthen in situ habitat management for these two populations. By contrast, although human activities have a milder impact on MPA, it is at high risk of extinction due to long-term contraction and isolation, and genetic rescue is urgently needed. MPB1 exhibited a relatively healthy population status and can potentially serve as a source population. Overall, our findings provide novel insights into the conservation of the Chinese pangolin and biogeography of the mammals of eastern Asia.
Similar content being viewed by others
Data availability
The raw whole genome resequencing reads have been deposited in the CNGB Sequence Archive of China National GeneBank DataBase (CNGBdb) (accession number: CNP0001723).
References
Alexander, D.H., Novembre, J., and Lange, K. (2009). Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19, 1655–1664.
Challender, D.W.S., Harrop, S.R., and MacMillan, D.C. (2015). Understanding markets to conserve trade-threatened species in CITES. Biol Conservation 187, 249–259.
Challender, D.W., and Hywood, L. (2011). Asian pangolins: increasing affluence driving hunting pressure. Traffic Bull 23, 92–93.
Challender, D.W., Nash, H.C., and Waterman, C. (2019). Pangolins: Science, Society and Conservation. London: Academic Press.
Chen, S., Zhou, Y., Chen, Y., and Gu, J. (2018). fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890.
Cheng, F., Tian, J., He, J., He, H., Liu, G., Zhang, Z., and Zhou, L. (2023). The spatial and temporal distribution of China’s forest carbon. Front Ecol Evol 11, 1110594.
Cingolani, P., Platts, A., Wang, L.L., Coon, M., Nguyen, T., Wang, L., Land, S.J., Lu, X., and Ruden, D.M. (2012). A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92.
Danecek, P., Auton, A., Abecasis, G., Albers, C.A., Banks, E., DePristo, M.A., Handsaker, R.E., Lunter, G., Marth, G.T., Sherry, S.T., et al. (2011). The variant call format and VCFtools. Bioinformatics 27, 2156–2158.
Do, C., Waples, R.S., Peel, D., Macbeth, G.M., Tillett, B.J., and Ovenden, J.R. (2014). NEESTIMATOR v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resources 14, 209–214.
Do, R., Balick, D., Li, H., Adzhubei, I., Sunyaev, S., and Reich, D. (2015). No evidence that selection has been less effective at removing deleterious mutations in Europeans than in Africans. Nat Genet 47, 126–131.
Dobrynin, P., Liu, S., Tamazian, G., **ong, Z., Yurchenko, A.A., Krasheninnikova, K., Kliver, S., Schmidt-Küntzel, A., Koepfli, K.P., Johnson, W., et al. (2015). Genomic legacy of the African cheetah, Acinonyx jubatus. Genome Biol 16, 1–20.
Durand, E.Y., Patterson, N., Reich, D., and Slatkin, M. (2011). Testing for ancient admixture between closely related populations. Mol Biol Evol 28, 2239–2252.
Edge, P., and Bansal, V. (2019). Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing. Nat Commun 10, 4660.
Excoffier, L., and Foll, M. (2011). Fastsimcoal: a continuous-time coalescent simulator of genomic diversity under arbitrarily complex evolutionary scenarios. Bioinformatics 27, 1332–1334.
Fu, J., and Wen, L. (2023). Impacts of Quaternary glaciation, geological history and geography on animal species history in continental East Asia: a phylogeographic review. Mol Ecol 32, 4497–4514.
Harris, R.S. (2007). Improved pairwise alignment of genomic DNA. State College: The Pennsylvania State University.
Hahn, C., Bachmann, L., and Chevreux, B. (2013). Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads—a baiting and iterative map** approach. Nucleic Acids Res 41, e129.
Hijmans, R.J. (2015). Raster: Geographic data analysis and modeling. R Package Version 2.4-15.
Heinrich, S., Wittmann, T.A., Prowse, T.A.A., Ross, J.V., Delean, S., Shepherd, C.R., and Cassey, P. (2016). Where did all the pangolins go? International CITES trade in pangolin species. Glob Ecol Conserv 8, 241–253.
Hohenlohe, P.A., Funk, W.C., and Rajora, O.P. (2021). Population genomics for wildlife conservation and management. Mol Ecol 30, 62–82.
Hu, J.Y., Hao, Z.Q., Frantz, L., Wu, S.F., Chen, W., Jiang, Y.F., Wu, H., Kuang, W.M., Li, H., Zhang, Y.P., et al. (2020a). Genomic consequences of population decline in critically endangered pangolins and their demographic histories. Natl Sci Rev 7, 798–814.
Hu, Y., Thapa, A., Fan, H., Ma, T., Wu, Q., Ma, S., Zhang, D., Wang, B., Li, M., Yan, L., et al. (2020b). Genomic evidence for two phylogenetic species and long-term population bottlenecks in red pandas. Sci Adv 6, eaax5751.
Hu, Y., Wang, X., Xu, Y., Yang, H., Tong, Z., Tian, R., Xu, S., Yu, L., Guo, Y., Shi, P., et al. (2023). Molecular mechanisms of adaptive evolution in wild animals and plants. Sci China Life Sci 66, 453–495.
Kardos, M., Åkesson, M., Fountain, T., Flagstad, Ø, Liberg, O., Olason, P., Sand, H., Wabakken, P., Wikenros, C., and Ellegren, H. (2018). Genomic consequences of intensive inbreeding in an isolated wolf population. Nat Ecol Evol 2, 124–131.
Kirin, M., McQuillan, R., Franklin, C.S., Campbell, H., McKeigue, P.M., and Wilson, J. F. (2010). Genomic runs of homozygosity record population history and consanguinity. PLoS ONE 5, e13996.
Klein Goldewijk, K., Beusen, A., Doelman, J., and Stehfest, E. (2017). Anthropogenic land use estimates for the Holocene-HYDE 3.2. Earth Syst Sci Data 9, 927–953.
Knaus, B.J., and Grünwald, N.J. (2017). VCFR: a package to manipulate and visualize variant call format data in R. Mol Ecol Resources 17, 44–53.
Kondrashov, A.S., and Crow, J.F. (1993). A molecular approach to estimating the human deleterious mutation rate. Hum Mutat 2, 229–234.
Kuang, W., Ming, C., Li, H., Wu, H., Frantz, L., Roos, C., Zhang, Y., Zhang, C., Jia, T., and Yang, J.Y. (2019). The origin and population history of the endangered golden snub-nosed monkey (Rhinopithecus roxellana). Mol Biol Evol 36, 487–499.
Kumar, S., Stecher, G., Li, M., Knyaz, C., and Tamura, K. (2018). MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35, 1547.
Kyriazis, C.C., Wayne, R.K., and Lohmueller, K.E. (2021). Strongly deleterious mutations are a primary determinant of extinction risk due to inbreeding depression. Evol Lett 5, 33–47.
Li, H., and Durbin, R. (2009). Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760.
Li, H., and Durbin, R. (2011). Inference of human population history from individual whole-genome sequences. Nature 475, 493–496.
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., Marth, G., Abecasis, G., and Durbin, R. (2009). The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079.
Librado, P., and Rozas, J. (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452.
Liu, X., and Fu, Y.X. (2015). Exploring population size changes using SNP frequency spectra. Nat Genet 47, 555–559.
Lu, H.Y., Yi, S.W., Xu, Z.W., Zhou, Y.L., Zeng, L., Zhu, F.Y., Feng, H., Dong, L.N., Zhuo, H.X., Yu, K.F., et al. (2013). Chinese deserts and sand fields in Last Glacial Maximum and Holocene Optimum. Chin Sci Bull 58, 2775–2783.
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A., Garimella, K., Altshuler, D., Gabriel, S., Daly, M., et al. (2010). The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303.
Mills, L.S., and Allendorf, F.W. (1996). The one-migrant-per-generation rule in conservation and management. Conserv Biol 10, 1509–1518.
Muscarella, R., Galante, P.J., Soley-Guardia, M., Boria, R.A., Kass, J.M., Uriarte, M., and Anderson, R.P. (2014). ENM eval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5, 1198–1205.
Myers, N., Mittermeier, R.A., Mittermeier, C.G., da Fonseca, G.A.B., and Kent, J. (2000). Biodiversity hotspots for conservation priorities. Nature 403, 853–858.
Nigenda-Morales, S.F., Lin, M., Nuñez-Valencia, P.G., Kyriazis, C.C., Beichman, A.C., Robinson, J.A., Ragsdale, A.P., Urbán R., J., Archer, F.I., Viloria-Gómora, L., et al. (2023). The genomic footprint of whaling and isolation in fin whale populations. Nat Commun 14, 5465.
Patterson, N., Moorjani, P., Luo, Y., Mallick, S., Rohland, N., Zhan, Y., Genschoreck, T., Webster, T., and Reich, D. (2012). Ancient admixture in human history. Genetics 192, 1065–1093.
Phillips, S.J., Anderson, R.P., and Schapire, R.E. (2006). Maximum entropy modeling of species geographic distributions. Ecol Model 190, 231–259.
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M.A.R., Bender, D., Maller, J., Sklar, P., de Bakker, P.I.W., Daly, M.J., et al. (2007). PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575.
Ralls, K., Ballou, J.D., Dudash, M.R., Eldridge, M.D.B., Fenster, C.B., Lacy, R.C., Sunnucks, P., and Frankham, R. (2018). Call for a paradigm shift in the genetic management of fragmented populations. Conserv Lett 11, e12412.
Robinson, J., Kyriazis, C.C., Yuan, S.C., and Lohmueller, K.E. (2023). Deleterious variation in natural populations and implications for conservation genetics. Annu Rev Anim Biosci 11, 93–114.
Santiago, E., Novo, I., Pardiñas, A.F., Saura, M., Wang, J., and Caballero, A. (2020). Recent demographic history inferred by high-resolution analysis of linkage disequilibrium. Mol Biol Evol 37, 3642–3653.
Sharma, H.P., Rimal, B., Zhang, M., Sharma, S., Poudyal, L.P., Maharjan, S., Kunwar, R., Kaspal, P., Bhandari, N., Baral, L., et al. (2020). Potential distribution of the critically endangered Chinese Pangolin (Manis pentadactyla) in different land covers of Nepal: implications for conservation. Sustainability 12, 1282.
Sun, X., Liu, Y.C., Tiunov, M.P., Gimranov, D.O., Zhuang, Y., Han, Y., Driscoll, C.A., Pang, Y., Li, C., Pan, Y., et al. (2023). Ancient DNA reveals genetic admixture in China during tiger evolution. Nat Ecol Evol 7, 1914–1929.
Vaser, R., Adusumalli, S., Leng, S.N., Sikic, M., and Ng, P.C. (2016). SIFT missense predictions for genomes. Nat Protoc 11, 1–9.
von Seth, J., Dussex, N., Diez-del-Molino, D., van der Valk, T., Kutschera, V.E., Kierczak, M., Steiner, C.C., Liu, S., Gilbert, M.T.P., Sinding, M.H.S., et al. (2021). Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations. Nat Commun 12, 2393.
Wang, Q., Lan, T., Li, H., Sahu, S.K., Shi, M., Zhu, Y., Han, L., Yang, S., Li, Q., Zhang, L., et al. (2022). Whole-genome resequencing of Chinese pangolins reveals a population structure and provides insights into their conservation. Commun Biol 5, 821.
Wei, S., Li, Z., Momigliano, P., Fu, C., Wu, H., and Merilä, J. (2020). The roles of climate, geography and natural selection as drivers of genetic and phenotypic differentiation in a widespread amphibian Hyla annectans (Anura: Hylidae). Mol Ecol 29, 3667–3683.
Wei, F., Huang, G., Guan, D., Fan, H., Zhou, W., Wang, D., and Hu, Y. (2022a). Digital Noah’s Ark: last chance to save the endangered species. Sci China Life Sci 65, 2325–2327.
Wei, S., Sun, S., Dou, H., An, F., Gao, H., Guo, C., and Hua, Y. (2022b). Influence of Pleistocene climate fluctuations on the demographic history and distribution of the critically endangered Chinese pangolin (Manis pentadactyla). BMC Zool 7, 1.
Willi, Y., Kristensen, T.N., Sgrò, C.M., Weeks, A.R., Ørsted, M., and Hoffmann, A.A. (2022). Conservation genetics as a management tool: the five best-supported paradigms to assist the management of threatened species. Proc Natl Acad Sci USA 119, e2105076119.
Wu, S.B. (2004). Assessment of threatened status of Chinese Pangolin (Manis pentadactyla). Chin J Appl Environ Biol 10, 456–461.
**ng, Y., and Ree, R.H. (2017). Uplift-driven diversification in the Hengduan Mountains, a temperate biodiversity hotspot. Proc Natl Acad Sci USA 114, E3444–E3451.
Yang, J., Lee, S.H., Goddard, M.E., and Visscher, P.M. (2011). GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88, 76–82.
Yang, L., Chen, M., Challender, D.W.S., Waterman, C., Zhang, C., Huo, Z., Liu, H., and Luan, X. (2018). Historical data for conservation: reconstructing range changes of Chinese pangolin (Manispentadactyla) in eastern China (1970–2016). Proc R Soc B 285, 20181084.
Yang, L., Wei, F., Zhan, X., Fan, H., Zhao, P., Huang, G., Chang, J., Lei, Y., and Hu, Y. (2022). Evolutionary conservation genomics reveals recent speciation and local adaptation in threatened takins. Mol Biol Evol 39, msac111.
Yang, S., Lan, T., Zhang, Y., Wang, Q., Li, H., Dussex, N., Sahu, S.K., Shi, M., Hu, M., Zhu, Y., et al. (2023). Genomic investigation of the Chinese alligator reveals wild-extinct genetic diversity and genomic consequences of their continuous decline. Mol Ecol Resour 23, 294–311.
Zhang, C., Dong, S., Xu, J., He, W., and Yang, T. (2019). PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35, 1786–1788.
Zhang, F., Wu, S., Zou, C., Wang, Q., Li, S., and Sun, R. (2016). A note on captive breeding and reproductive parameters of the Chinese pangolin, Manis pentadactyla Linnaeus, 1758. ZooKeys 618, 129–144.
Zhang, H., Miller, M.P., Yang, F., Chan, H.K., Gaubert, P., Ades, G., and Fischer, G.A. (2015). Molecular tracing of confiscated pangolin scales for conservation and illegal trade monitoring in Southeast Asia. Glob Ecol Conserv 4, 414–422.
Zhang, M., Cao, P., Dai, Q.Y., Wang, Y.Q., Feng, X.T., Wang, H.R., Wu, H., Min-Shan Ko, A., Mao, X.W., Liu, Y.C., et al. (2021). Comparative analysis of DNA extraction protocols for ancient soft tissue museum samples. Zoological Res 42, 280–286.
Zhao, S., Zheng, P., Dong, S., Zhan, X., Wu, Q., Guo, X., Hu, Y., He, W., Zhang, S., Fan, W., et al. (2013). Whole-genome sequencing of giant pandas provides insights into demographic history and local adaptation. Nat Genet 45, 67–71.
Zheng, B., Xu, Q., and Shen, Y. (2002). The relationship between climate change and Quaternary glacial cycles on the Qinghai-Tibetan Plateau: review and speculation. Quat Int 97–98, 93–101.
Acknowledgement
This work was supported by the Major Program of the Natural Science Foundation of Jiangxi Province of China (20233ACB209001), Guangdong Basic and Applied Basic Research Foundation (2022A1515111016), Science and Technology Department of Guangdong Province (2021QN02H103), Natural Resources Affairs Management-Ecological Forestry Construction Special Project of Forestry Administration of Guangdong Province (SLYJ2023B4002, SLYJ2023B4003, SLYJ2023B4005). We express sincere thanks to the Department of Wildlife Conservation of Forestry Administration of Guangdong Province for their support during the project implementation. We are also grateful to ** Huang & Fuwen Wei
Corresponding author
Ethics declarations
The author(s) declare that they have no conflict of interest. F.Wei. conceived and supervised the project. S.Wei., H.Fan., W.Zhou., G.Huang., Y.Hua., S.Wu., X.Wei. and Y.Chen. collected the samples. X.Tan. carried out the DNA extraction and library construction. S.Wei. performed the bioinformatics analysis. S.Wei. wrote the manuscript. F.Wei., H.Fan., W.Zhou., S.Wei. revised the manuscript. All authors contributed to interpreting the data and approved the final manuscript.
Rights and permissions
About this article
Cite this article
Wei, S., Fan, H., Zhou, W. et al. Conservation genomics of the critically endangered Chinese pangolin. Sci. China Life Sci. (2024). https://doi.org/10.1007/s11427-023-2540-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11427-023-2540-y