Abstract
The prevalence of age-related cognitive disorders/dementia is increasing, and effective prevention and treatment interventions are lacking due to an incomplete understanding of aging neuropathophysiology. Emerging evidence suggests that abnormalities in gut microbiome are linked with age-related cognitive decline and getting acceptance as one of the pillars of the Geroscience hypothesis. However, the potential clinical importance of gut microbiome abnormalities in predicting the risk of cognitive decline in older adults is unclear. Till now the majority of clinical studies were done using 16S rRNA sequencing which only accounts for analyzing bacterial abundance, while lacking an understanding of other crucial microbial kingdoms, such as viruses, fungi, archaea, and the functional profiling of the microbiome community. Utilizing data and samples of older adults with mild cognitive impairment (MCI; n = 23) and cognitively healthy controls (n = 25). Our whole-genome metagenomic sequencing revealed that the gut of older adults with MCI harbors a less diverse microbiome with a specific increase in total viruses and a decrease in bacterial abundance compared with controls. The virome, bacteriome, and microbial metabolic signatures were significantly distinct in subjects with MCI versus controls. Selected bacteriome signatures show high predictive potential of cognitive dysfunction than virome signatures while combining virome and metabolic signatures with bacteriome boosts the prediction power. Altogether, the results from our pilot study indicate that trans-kingdom microbiome signatures are significantly distinct in MCI gut compared with controls and may have utility for predicting the risk of develo** cognitive decline and dementia- debilitating public health problems in older adults.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With an aging world population, cognitive decline and dementia are debilitating public health problems in older adults [17]. Alzheimer’s disease (AD) is the most common age-related cognitive disorder [53]. Around 6 million older adults are living with Alzheimer’s disease and related dementia (ADRD) in the USA, and this number is expected to grow double by 2050 [41]. Currently, there are no clinically impactful prevention strategies and no treatments that can significantly alter the course of the illness. As a result, ADRD causes great strain on families, society, and the healthcare system [12, 39]. Drug clinical trials for treating AD [34, 57] lack a full understanding of ADRD pathophysiology as well as the right targets and time frame to introduce interventions. Prior studies indicate that specific diet and exercise regimens may slow the progression of ADRD in older adults [42, 49]. However, the early detection of cognitive decline and dementia risk is cumbersome, expensive, and not available for routine clinical use [29]. Therefore, development of inexpensive, safe, and easy-to-measure testing is direly needed for slowing or preventing the progression of dementia in older adults.
Emerging evidence suggests that abnormalities in gut microbiome may contribute to aging biology mechanisms [45]. A few studies also indicate that the gut microbiome signatures may be different in older adults with ADRD compared with their age-matched controls [14, 36, 37, 56]. Vogt, et al. showed that Blautia, Phascolarctobacterium, Gemella, Bacteroides, Bilophila, and Alistipes bacteria (many of them are commensal pathogens) were significantly increased, and SMB53 (family Clostridiaceae), Dialister, Clostridium, Turicibacter, Bifidobacterium, Adlercreutzia, and cc115 (family Erysipelotrichaceae) (many of them are beneficial/probiotics) were specifically decreased in gut of AD patients compared to controls [55]. In addition, Escherichia/Shigella, Ruminococcaceae, Enterococcaceae, and Lactobacillaceae bacteria were significantly increased and E. rectale, Lanchnospiraceae, Bacteroidaceae, and Veillonellaceae were significantly decreased in older adults with mild cognitive impairment (MCI) and were linked with AD markers in cerebrospinal fluid (CSF) [11, 36, 37, 62].
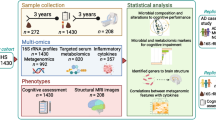
Gut microbiome signatures are greatly influenced by dietary habits, and impact of dietary manipulations on slowing cognitive decline or dementia progression may be through gut microbiome [2, 15]. We have shown that a modified Mediterranean ketogenic diet (MMKD) may change the gut microbiome composition and ameliorate AD pathology in MCI subjects [36]. However, these studies were aimed at describing the difference in microbiome signatures, but not testing their significance for predicting or differentiating cognitive dysfunctions in older adults. In addition, majority of previous studies have used 16S rRNA sequencing which only allows for analysis of the bacteria population (bacteriome) of gut microbiome. The role and significance of other microbial kingdoms (i.e., viruses, fungi, and archaea) that also coexist with bacteria in the human gut remain unstudied. Herein, we performed whole genome sequencing on fecal DNA samples of older adults (≥ 60 years of age) with MCI and normal cognition from the cohort of the MiaGB (Microbiome in aging Gut and Brain) consortium—a multi-site study focused on examining the relationship between the microbiome and aging [31]. The present study investigated the associations between shotgun metagenomics-based trans-kingdom microbiome signatures and cognitive health by comparing older adults with and without mild cognitive impairment.
Materials and methods
Human subjects
The data and samples used in this study were procured from the Microbiome in aging Gut and Brain (MiaGB) Consortium cohort as a pilot study. The MiaGB consortium is recruiting community dwelling older adults in Florida at five sites. All the participants (n = 48) included in this study were 60 years of age or older. Among them, 23 were with MCI, while 25 subjects were cognitively healthy controls. Cognitive function assessments were performed as described below. The demographic characteristics are depicted in Table 1. Exclusion criteria consisted of persons with (a) history of brain and gut-related surgeries in the past five years; (b) history of cancer diagnosis and/or treatment (except non-melanoma skin cancer) in the past five years; (c) neurological disorders of epilepsy, Parkinson’s disease, and amyotrophic lateral sclerosis; (d) antibiotic use in the preceding 30 days, (e) diarrhea, vomiting, or food poisoning in the past 30 days; and (f) a history of inflammatory bowel diseases. Informed consent was obtained from each participant. All recruitments, study protocols, and procedures were approved the Institutional Review Board of University of South Florida committee and were performed according to the approved guidelines.
Cognitive function assessments
The Montreal cognitive assessment (MoCA) [23], MiniCog [7], and Memory impairment screen (MIS) [30] were performed by trained staff and scores were calculated using standard protocols.
Stool sample collection
Fecal microbiome samples were collected using an in-house developed stool sample collection kit, which has been validated and accepted by older adults in several of our past [36, 37] and ongoing clinical studies. The use of this kit has increased the compliance and adherence in our studies. The stool collection kit is given to participants to take home, and samples transported to the lab within 24 h of stool passing and collection, and samples were immediately aliquoted and stored at − 80 °C until further analysis.
Metagenomic shotgun sequencing
Fecal DNA was extracted using 150 mg of the human stool samples using QIAamp PowerFecal Pro DNA Kit (Qiagen, USA) following the manufacturer’s instructions. The DNA was quantified using Qubit dsDNA HS assay kit (Thermo Fisher Scientific, USA). The extracted and quantified DNA (150 ng) was used for library preparation using Illumina® DNA Prep, (M) Tagmentation kit (Illumina, Inc, 5200 Illumina Way, San Diego CA, USA) by following the manufacturer’s instructions. Additionally, sample specific unique IDT for Illumina–Nextera DNA UD Indexes were used. The sequencing was done on Illumina NextSeq1000 machine using an Illumina NextSeq 1000/2000 P2 Reagents (300 Cycles) v3 reagent cartridge (Illumina, Inc, 5200 Illumina Way, San Diego CA, USA). All the data was captured and stored in the BaseSpace cloud and was analyzed further using bioinformatics pipelines, as described below.
Bioinformatics and statistical analysis
The analysis for the shotgun sequencing data was performed using the Yet Another Metagenomic Pipeline (YAMP) workflow [54]. The YAMP workflow uses tools from bbmap suite for de-duplication, trimming, and decontamination of metagenomics sequences [10]. It uses FastQC for the visualization of the raw and QC filtered metagenomic reads [1]. The additional tools used in the YAMP pipeline are MetaPhlAn [5] for taxonomic binning and profiling of microbes and their relative abundance in the samples, HUMAnN pipeline for the estimation of the functional capabilities of the microbiome community [5], and QIIME2 [21] for the evaluation of the multiple alpha diversity measures including observed OTUs, Shannon and Simpson alpha diversity. The MetaPhlAn database relies on ~ 1.1 M unique clade-specific marker genes identified from ~ 100,000 reference genomes (~ 99,500 bacterial and archaeal and ~ 500 eukaryotic), which allows unambiguous taxonomic assignments, an accurate estimation of organismal relative abundance, species-level resolution for bacteria, archaea, eukaryotes, and viruses. The HUMAnN pipeline which uses MetaPhlAn and ChocoPhlAn pangenome database to facilitate fast, accurate, and organism-specific functional profiling of Archaea, Bacteria, Eukaryotes, and Viruses considerably expanded databases of genomes, genes, and pathways by map** the metagenome reads on the reference databases. The β-diversity across the sample groups was estimated using Principal Component Analysis (PCA) based on Euclidean distances. Taxonomic abundance of microbial taxa at phylum and species level are represented. The shared and unique bacterial taxa were estimated using a web-based tool interactiveVenn [22]. Statistical analysis of the data was done using Graphpad Prism [6] and Stamp [40]. Various R-scripts including ggplot2 were used for the analysis and presentation of the data like corrplot for the correlation analysis of microbiome components and the cognitive scores of the study participants. The random forest analysis was performed using the web-based tool microbiome analysts [ The combination of bacteriome, virome, and microbial metabolic pathways improves the prediction of cognitive health in older adults. a–c) ROC analyses depicting prediction model 1 (a), 2 (b), and 3 (c) with a distinct combination of bacteriome, virome, and microbial metabolic pathways to predict the cognitive health of older adults
Discussion
Gut microbiome composition and functions are known to be significantly different between healthy older adults and young adults as well as between older adults with cognitive disorders like AD and cognitively healthy adults [8, 9, 14, 36, 37, 47, 56]. Similar observations are noted in animal models [26, 60]. However, the results are inconsistent and largely based on targeted 16S rRNA sequencing, which accounts only for bacterial abundance. It is evident that the microbiota is a complex community comprising bacteria, viruses, fungi, and archaea, which together, are interconnected to survive and function. However, the relationship of these microbiome communities in aging biology and cognitive health is poorly understood. Earlier studies show that the gut bacterial population (bacteriome) is variable, which limits its potential to be used as a biomarker to predict cognitive decline in older adults. Furthermore, it is unclear if combining bacteria, viruses, and their metabolic pathways using combinatorial approach can be used to predict cognitive decline. Herein, our results show that the gut of older adults with MCI harbors not only distinct bacteria, but also distinct viruses, with limited number of fungi and archaea detected, and we show that combining the bacteriome, virome, and their metabolic pathways boosts the predictive potential of microbiome to differentiate MCI from cognitively healthy older adults.
The microbiome α-diversity is an indicator of function of the microbiota (higher is better), and we show that the gut of older adults with MCI has lower microbiome α-diversity indices like Shannon and Simpson indexes, which were positively associated with reduced MoCA and MiniCog scores (lower scores indicate poor cognitive function). These results indicate that the gut of older adults with MCI harbors a significantly distinct microbiome compared with the gut of cognitively healthy older adults. Multiple emerging studies indicate associations between gut microbiota diversity and taxonomic signatures with neurological outcomes, including cognitive function and dementia [29, 42]. Preclinical studies using germ-free or antibiotic-treated rodents show no or reduced microbiome diversity, respectively with significant cognitive deficits such as reduced memory, impaired working memory, and changes in brain-derived neurotrophic factor in the hippocampus [19, 32, 35, 43]. Small-scale human studies including ours also showed a link between abnormalities in microbial features and cognition dysfunctions, or found significant improvements when comparing controls with persons who have been treated with probiotics or Mediterranean ketogenic diet to increase commensal microbiota [43]. Our findings are consistent with results from animal models and other clinical studies and advance the understanding of trans-kingdom differences of bacteria and viruses corresponding to the cognitive function in the gut of older adults.
The mechanisms by which the gut microbiome is associated with cognitive health are not fully established yet; however, growing data indicate that microbiota-produced beneficial metabolites such as short-chain fatty acids (SCFAs like acetate, propionate, and butyrate) significantly contribute in gut-brain communications [16, 24]. Herein, we observed that the abundance of butyrate producing bacteria such as Lachnospiraceae family, Subdoligranulum sp., Roseburia intestinalis, and Roseburia hominis were reduced in the gut of MCI participants compared to cognitively healthy controls. It has been previously reported that a decline in abundance of butyrate producing bacteria is associated with multiple disorders such inflammatory bowel disease, type 2 diabetes mellitus as well as poor intestinal barrier function, immune dysregulation, and gut dysbiosis [4, 50]. We showed that older adults with MCI had higher abundance of Lactococcus phage ul36 which specifically regulates the probiotic bacteria like Lactococcus lactis (a common yogurt culture) [44] and may diminish their abundance in the gut, which may ultimately be detrimental for gut-brain axis. On other hand, certain bacteriophages exhibit prophage-like properties, such as Stx2 converting phage 1717, which functions as a mobile genetic element in bacterial genome containing crucial genes associated with bacterial pathogenesis [61]. The virome signatures detected in the present study were highly variable, and their precise role in age-related cognitive decline remains to be determined.
The importance of microbiome signature differences is debatable due to the high variability in their diversity, taxonomic features, and functions. In this study, we tested the potential of bacteriome, virome, and their metabolic signatures to predict the cognitive health in older adults. Interestingly, we found that the virome showed the highest number of unique viral species in the gut of MCI versus cognitively healthy controls and the strongest predictive power. However, one limitation of using the virome is that viruses are not uniformly present in the human gut. Bacterial species chosen based on differential microbiome signatures and random forest analyses showed significant potential to differentiate (~ 76%) MCI from their cognitively healthy controls. We also observed that the one bacterial species representing a specific enterotype presents more predictive power to differentiate MCI from controls; however, single species or that particular enterotype was not present in all the samples, instead of present in limited samples. Thus, our approach of using combination of virome, bacteriome, and metabolic signatures presented broader application for all the individuals. Interestingly, the addition of selected signatures of virome and microbial metabolic pathways to bacteriome signatures boosted the power to predict the risk of cognitive decline in older adults.
Our study presents several strengths: our results are derived from community dwelling older adults rather than institutionalized patients; the microbiome sequencing was done using whole genome metagenomics which allowed us to identify bacteria, viruses, fungi, and archaea altogether, and we used the well-established cohort of MiaGB consortium which uses standardized protocols for all data collection and quality control, including participant surveys, cognitive functions assessments, stool collection, processing, and sequencing.
We acknowledge that our study also has few limitations. Our sample size is relatively small for comprehensive analysis of multiple microbial signatures, and we were also not able to determine the potential role of sex, race, and ethnicity. We attempted to understand the association of cognitive health and microbiome of the study participants. However, no statistically significant differences were recorded in this cross-sectional study. Nonetheless, the present study was an exploratory analysis of initial data collected in the MiaGB consortium, which is a cohort study aimed at recruiting approximately 400 older adults and following them on a yearly basis. Further analyses to use and validate the findings of this study will be performed in the future. One immediate use will be to use these results as a training cohort/dataset to test the efficacy and accuracy of these predictions. Lastly, while the current study is cross-sectional, which prevents the assessment of temporality in microbiome signatures as longitudinal data from participants in the MiaGB consortium become available, we will be able to define the temporal changes associated with trans-kingdom signatures so we can better understand the relationship between these and cognitive health.