Abstract
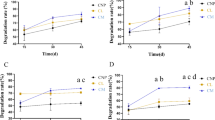
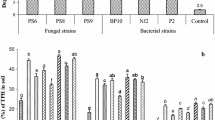
Hydrocarbonoclastic bacterial strains were isolated from rhizosphere of plants growing in crude oil-contaminated sites of Assam, India. These bacteria showed plant growth-promoting attributes, even when exposed to crude oil. Two independent pot trials were conducted to test the rhizodegradation ability of the bacterial consortium in combination of plants Azadirchta indica or Delonix regia in crude oil-contaminated soil. Field experiments were conducted at two crude oil-contaminated agricultural field at Assam (India), where plants (A. indica or D. regia) were grown with the selected bacterial consortium consisting of five hydrocarbonoclastic bacterial isolates (Gordonia amicalis BB-DAC, Pseudomonas aeruginosa BB-BE3, P. citronellolis BB-NA1, Rhodococcus ruber BB-VND, and Ochrobactrum anthropi BB-NM2), and NPK was added to the soil for biostimulation. The bacterial consortium-NPK biostimulation led to change in rhizosphere microbiome with enhanced degradation of petroleum hydrocarbons (PHs) in soils contaminated with crude oil. After 120 days of planting A. indica + consortium + NPK treatment, degradation of PHs was found to be up to 67%, which was 55% with D. regia with the same treatment. Significant changes in the activities of plant and soil enzymes were also noted. The shift is bacterial community was also apparent as with A. indica, the relative abundance of Proteobacteria, Actinobacteria, and Acidobacteria increased by 35.35%, 26.59%, and 20.98%, respectively. In the case of D. regia, the relative abundance of Proteobacteria, Actinobacteria, and Acidobacteria were increased by 39.28%, 35.79%, and 9.60%, respectively. The predicted gene functions shifted in favor of the breakdown of xenobiotic compounds. This study suggests that a combination of plant-bacterial consortium and NPK biostimulation could be a productive approach to bioengineering the rhizosphere microbiome for the purpose of commercial bioremediation of crude oil-contaminated sites, which is a major environmental issue faced globally.





Similar content being viewed by others
Data Availability
The 16S rRNA sequences were deposited to NCBI and obtained NCBI GenBank accession numbers MK967987 MN493647, MN508937, MN480535, and MN493755 and raw sequencing data of NGS-based community analysis has been deposited in the sequence read archive of the NCBI under SRA numbers SRR13994212, SRR22142642, SRR14292154, and SRR22163873 and BioProject numbers PRJNA715307, PRJNA896944, PRJNA723445, and PRJNA897097.
The identified and collected isolates were submitted to National Centre for Microbial Resource (NCMR), Pune, India under the accession nos. MCC4370, MCC 4426, MCC 4425, MCC4369, and MCC4942, respectively, and also available in Soil and Environmental Microbiology Laboratory (SEML), Department of Microbiology, Assam University Silchar, Assam, India.
References
Abbasian F, Lockington R, Mallavarapu M, Naidu R (2015) A comprehensive review of aliphatic hydrocarbon biodegradation by bacteria. Appl Biochem Biotechnol 176:670–699
Adams GO, Fufeyin PT, Okoro SE, Ehinomen I (2015) Bioremediation, biostimulation and bioaugmention: a review. Int J Environ Bioremediat Biodegrad 3(1):28–39
Aisien FA, Chiadikobi JC, Aisien ET (2009) Toxicity assessment of some crude oil contaminated soils in the Niger delta. In: Advanced Materials Research, vol 62. Trans Tech Publications Ltd., pp 451–455
Ali M, Song X, Ding D, Wang Q, Zhang Z, Tang Z (2022) Bioremediation of PAHs and heavy metals co-contaminated soils: challenges and enhancement strategies. Environ Pollut 295:118686
Ali MH, Khan MI, Naveed M, Tanvir MA (2023) Microbe-assisted rhizoremediation of hydrocarbons and growth promotion of chickpea plants in petroleum hydrocarbons-contaminated soil. Sustainability 15:6081. https://doi.org/10.3390/su15076081
Alrumman SA, Standing DB, Paton GI (2015) Effects of hydrocarbon contamination on soil microbial community and enzyme activity. J King Saud Univ Sci 27(1):31–41
Bhuyan B, Pandey P (2022) Remediation of petroleum hydrocarbon contaminated soil using hydrocarbonoclastic rhizobacteria, applied through Azadirachta indica rhizosphere. Int J Phytoremediation 24(13):1444–1454
Blokhina O, Virolainen E, Fagerstedt KV (2003) Antioxidants, oxidative damage and oxygen deprivation stress: a review. Ann Bot 91(2):179–194
Bourdel G, Roy-Bolduc A, St-Arnaud M et al (2016) Concentration of petroleum-hydrocarbon contamination shapes fungal endophytic community structure in plant roots. Front Microbiol 7:685
Brzeszcz J, Kaszycki P (2018) Aerobic bacteria degrading both n-alkanes and aromatic hydrocarbons: an undervalued strategy for metabolic diversity and flexibility. Biodegradation 29:359–407
Cavalcanti FR, Oliveira JTA, Martins-Miranda AS, Viégas RA, Silveira JAG (2004) Superoxide dismutase, catalase and peroxidase activities do not confer protection against oxidative damage in salt-stressed cowpea leaves. New Phytol 163(3):563–571
Chen W, Kong Y, Li J, Sun Y, Min J, Hu X (2020) Enhanced biodegradation of crude oil by constructed bacterial consortium comprising salt-tolerant petroleum degraders and biosurfactant producers. Int Biodeterior Biodegradation 154:105047
Chicca I, Becarelli S, Bernabei G, Siracusa G, Di Gregorio S (2022) Innovative culturomic approaches and predictive functional metagenomic analysis: the isolation of hydrocarbonoclastic bacteria with plant growth promoting capacity. Water 14(2):142
Curiel-Alegre S, Velasco-Arroyo B, Rumbo C, Khan AHA, Tamayo-Ramos JA, Rad C, Gallego JLR, Barros R (2022) Evaluation of biostimulation, bioaugmentation, and organic amendments application on the bioremediation of recalcitrant hydrocarbons of soil. Chemosphere 307:135638
Das N, Bhuyan B, Pandey P (2022) Correlation of soil microbiome with crude oil contamination drives detection of hydrocarbon degrading genes which are independent to quantity and type of contaminants. Environ Res 215:114185
Debnath S, Das A, Maheshwari DK, Pandey P (2023) Treatment with atypical rhizobia, Pararhizobium giardinii and Ochrobactrum sp. modulate the rhizospheric bacterial community, and enhances Lens culinaris growth in fallow-soil. Microbiol Res 267:127255
Ebadi A, Sima NAK, Olamaee M, Hashemi M, Nasrabadi RG (2017) Effective bioremediation of a petroleum-polluted saline soil by a surfactant-producing Pseudomonas aeruginosa consortium. J Adv Res 8(6):627–633
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461
Gao YC, Wang JN, Guo SH, Hu YL, Li TT, Mao R, Zeng DH (2015) Effects of salinization and crude oil contamination on soil bacterial community structure in the Yellow River Delta region, China. Appl Soil Ecol 86:165–173. https://doi.org/10.1016/j.apsoil.2014.10.011
Gaur VK, Gupta S, Pandey A (2021) Evolution in mitigation approaches for petroleum oil-polluted environment: recent advances and future directions. Environ Sci Pollut Res 29:61821–61837
Gelsomino A, Badalucco L, Landi L, Cacco G (2006) Soil carbon, nitrogen and phosphorus dynamics as affected by solarization alone or combined with organic amendment. Plant Soil 279:307–325
Glick BR (2010) Using soil bacteria to facilitate phytoremediation. Biotechnol Adv 28(3):367–374
Gong XB (2012) Remediation of weathered petroleum oil-contaminated soil using a combination of biostimulation and modified Fenton oxidation. Int Biodeterior Biodegradation 70:89–95
Grimm AC, Harwood CS (1997) Chemotaxis of Pseudomonas spp. to the polyaromatic hydrocarbon naphthalene. Appl Environ Microbiol 63(10):4111–4115
Hafiluddin (2011) Bioremediasi Tanah Tercemar Minyak dengan Teknik Bioaugmentasi dan Biostimulasi. Bioremediation of oil polluted soils by bioaugmentation technique and biostimulation. Jurnal Embryo 8:47–52
Hameed A, Sheikh MA (2007) Changes in catalase, peroxidase activities and soluble proteins in wheat leaves on thiourea and H2O2 treatments. Biosci Res 4(1):21–27
Haziev FH (2005) Methods of soil enzymology. Nauka, Moscow (in Russian)
Hussain F, Hussain I, Khan AHA, Muhammad YS, Iqbal M, Soja G, Reichenauer TG, Yousaf S (2018) Combined application of biochar, compost, and bacterial consortia with Italian ryegrass enhanced phytoremediation of petroleum hydrocarbon contaminated soil. Environ Exp Bot 153:80–88
Kaimi E, Mukaidani T, Tamaki M (2007) Screening of twelve plant species for phytoremediation of petroleum hydrocarbon-contaminated soil. Plant Prod Sci 10(2):211–218. https://doi.org/10.1626/pps.10.211
Kauppi S, Sinkkonen A, Romantschuk M (2011) Enhancing bioremediation of diesel-fuel-contaminated soil in a boreal climate: comparison of biostimulation and bioaugmentation. Int Biodeterior Biodegradation 65(2):359–368
Kim AW, Vane CH, Moss-Hayes V, Engelhart SE, Kemp AC (2018) PAH, PCB, TPH and mercury in surface sediments of the Delaware River Estuary and Delmarva Peninsula, USA. Mar Pollut Bull 129(2):835–845
Kotoky R, Das S, Singha LP, Pandey P, Singha KM (2017) Biodegradation of benzo(a)pyrene by biofilm-forming and plant growth promoting Acinetobacter sp. strain PDB4. Environ Technol Innov 8:256–268. https://doi.org/10.1016/j.eti.2017.07.007
Kotoky R, Nath S, Kumar Maheshwari D, Pandey P (2019) Cadmium resistant plant growth promoting rhizobacteria Serratia marcescens S2I7 associated with the growth promotion of rice plant. Environ Sustain 2:135–144
Kotoky R, Pandey P (2020) Difference in the rhizosphere microbiome of Melia azedarach during removal of benzo(a)pyrene from cadmium co-contaminated soil. Chemosphere 258:127175
Kwon JH, Ji MK, Kumar R, Islam MM, Khan MA, Park YK et al (2023) Recent advancement in enhanced soil flushing for remediation of petroleum hydrocarbon-contaminated soil: a state-of-the-art review. Rev Environ Sci Biotechnol:1–36
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821
Li Z, Lee K, King T, Niu H, Boufadel MC, Venosa AD (2011) Application of entropy analysis of in situ droplet-size spectra in evaluation of oil chemical dispersion efficacy. Mar Pollut Bull 62(10):2129–2136
Li J, Zhang J, Lu Y, Chen Y, Dong S, Shim H (2012) Determination of total petroleum hydrocarbons (TPH) in agricultural soils near a petrochemical complex in Guangzhou, China. Environ Monit Assess 184:281–287
Liu W, Zhang ZHE, Wan S (2009) Predominant role of water in regulating soil and microbial respiration and their responses to climate change in a semiarid grassland. Glob Chang Biol 15(1):184–195
Lu RK (ed) (1999) Analytical method of soil agricultural chemistry. China Agricultural Science and Technology Press, Bei**g
Lu J, Breitwieser FP, Thielen P, Salzberg SL (2017) Bracken: estimating species abundance in metagenomics data. PeerJ Comp Sci 3:104. https://doi.org/10.7717/peerj-cs.104
Lu C, Hong Y, Liu J, Gao Y, Ma Z, Yang B, Ling W, Waigi MG (2019) A PAH-degrading bacterial community enriched with contaminated agricultural soil and its utility for microbial bioremediation. Environ Pollut 251:773–782
Ma B, He Y, Chen HH, Xu JM, Rengel Z (2010) Dissipation of polycyclic aromatic hydrocarbons (PAHs) in the rhizosphere: synthesis through meta-analysis. Environ Pollut 158(3):855–861
McDonald D, Price MN, Goodrich J et al (2012) An improved Greengenes axonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6:610–618
Margesin R, Schinner F (2001) Bioremediation (natural attenuation and biostimulation) of diesel-oil-contaminated soil in an alpine glacier skiing area. Appl Environ Microbiol 67(7):3127–3133
Medvedeva N, Polyak Y, Zaytseva T, Zharikov G (2011) Biodegradation of mustard gas hydrolysis products in contaminated soils. In Contaminated Soils. In: Environmental Impact, Disposal and Treatment. elibrary.ru, pp 289–314
Njoku KL, Akinola MO, Oboh BO (2008) Does crude oil affect pH, moisture and organic matter content of soils. Ecol Environ Conserv 14:731–736 https://ir.unilag.edu.ng/handle/123456789/5656
Okerentugba PO, Ezeronye OU (2003) Petroleum degrading potentials of single and mixed microbial cultures isolated from rivers and refinery effluent in Nigeria. Afr J Biotechnol 2(9):288–292
Pacwa-Płociniczak M, Płaza GA, Piotrowska-Seget Z (2016) Monitoring the changes in a bacterial community in petroleum-polluted soil bioaugmented with hydrocarbon-degrading strains. Appl Soil Ecol 105:76–85
Pant R, Pandey P, Kotoky R (2016) Rhizosphere mediated biodegradation of 1,4-dichlorobenzene by plant growth promoting rhizobacteria of Jatropha curcas. Ecol Eng 94:50–56
Payne SM (1994) Detection, isolation and characterization of siderophores. Methods Enzymol 44:235–329
Polyak YM, Bakina LG, Chugunova MV, Mayachkina NV, Gerasimov AO, Bure VM (2018) Effect of remediation strategies on biological activity of oil-contaminated soil—a field study. Int Biodeterior Biodegradation 126:57–68
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5(3):9490. https://doi.org/10.1371/journal.pone.0009490
Panwar R, Mathur J (2023) Microbial-assisted phytodegradation for the amelioration of pyrene-contaminated soil using Pseudomonas aeruginosa and Aspergillus oryzae with alfalfa and sunflower. 3. Biotech 13(7):251
Patowary K, Patowary R, Kalita MC, Deka S (2016) Development of an efficient bacterial consortium for the potential remediation of hydrocarbons from contaminated sites. Front Microbiol 7:1092
Qu Y, Gong Y, Ma J, Wei H, Liu Q, Liu L, Wu H, Yang S, Chen Y (2020) Potential sources, influencing factors, and health risks of polycyclic aromatic hydrocarbons (PAHs) in the surface soil of urban parks in Bei**g, China. Environ Poll 260:114016
Rafique HM, Khan MY, Asghar HN, Ahmad Zahir Z, Nadeem SM, Sohaib M, Alotaibi F, Al-Barakah FN (2022) Converging alfalfa (Medicago sativa L.) and petroleum hydrocarbon acclimated ACC-deaminase containing bacteria for phytoremediation of petroleum hydrocarbon contaminated soil. Int J Phytoremediation 25(6):717–727
Rahman KSM, Thahira-Rahman J, Lakshmanaperumalsamy P, Banat IM (2002) Towards efficient crude oil degradation by a mixed bacterial consortium. Bioresour Technol 85(3):257–261
Rajkumari J, Choudhury Y, Bhattacharjee K, Pandey P (2021) Rhizodegradation of pyrene by a non-pathogenic Klebsiella pneumoniae isolate applied with Tagetes erecta L. and changes in the rhizobacterial community. Front Microbiol 12:593023
Readman JW, Fillmann G, Tolosa I, Bartocci J, Villeneuve JP, Catinni C, Mee LD (2002) Petroleum and PAH contamination of the Black Sea. Mar Pollut Bull 44(1):48–62. https://doi.org/10.1016/s0025-326x(01)00189-8
Samuel EA, Oladipupo OO (2012) Factorial designs application to study enhanced bioremediation of soil artificially contaminated with weathered bonny light crude oil through biostimulation and bioaugmentation strategy. J Environ Prot 3(8):21757
Sarkar P, Meghvanshi M, Singh R (2011) Microbial consortium: a new approach in effective degradation of organic kitchen wastes. Int J Environ Sci Technol 2(3):170
Saravanan A, Jeevanantham S, Narayanan VA, Kumar PS, Yaashikaa PR, Muthu CM (2020) Rhizoremediation—a promising tool for the removal of soil contaminants: a review. J Environ Chem Eng 8(2):103543
Schnrer J, Rosswall T (1982) Fluorescein diacetate hydrolysis as a measure of total microbial activity in soil and litter. Appl Environ Microbiol 43(6):1256–1261
Singha LP, Pandey P (2021) Rhizosphere assisted bioengineering approaches for the mitigation of petroleum hydrocarbons contamination in soil. Crit Rev Biotechnol 41(5):749–766
Singha LP, Sinha N, Pandey P (2018) Rhizoremediation prospects of polyaromatic hydrocarbon degrading rhizobacteria, that facilitate glutathione and glutathione-S-transferase mediated stress response, and enhance growth of rice plants in pyrene contaminated soil. Ecotoxicol Environ Saf 164:579–588
Singha LP, Pandey P (2017) Glutathione and glutathione-S-transferase activity in Jatropha curcas in association with pyrene degrader Pseudomonas aeruginosa PDB1 in rhizosphere, for alleviation of stress induced by polyaromatic hydrocarbon for effective rhizoremediation. Ecol Eng 102:422–432
Sun W, **ao E, Krumins V, Häggblom MM, Dong Y, Pu Z, Li B, Wang Q, **ao T, Li F (2018) Rhizosphere microbial response to multiple metal(loid)s in different contaminated arable soils indicates crop-specific metal-microbe interactions. Appl Environ Microbiol 84(24):e00701–e00718
Tabatabai MA (1994) Soil enzymes. Methods of soil analysis: Part 2. Microb Biochem Technol 5:775–833
Taccari M, Milanovic V, Comitini F, Casucci C, Ciani M (2012) Effects of biostimulation and bioaugmentation on diesel removal and bacterial community. Int Biodeterior Biodegradation 66(1):39–46
Taylor JP, Wilson B, Mills MS (2002) Comparison of microbial numbers and enzymatic activities in surface soils and subsoils using various techniques. Soil Biol Biochem 34:387–401
Tangahu BV, Vyatrawan L, Nurmalasari R, Pirade F (2017) Bioremediation of oil contaminated soil by biostimulation method using NPK fertilizer. Open Access Lib J 4(11):1–8
Topaç FO, Dindar E, Uçaroğlu S, Başkaya HS (2009) Effect of a sulfonated azo dye and sulfanilic acid on nitrogen transformation processes in soil. J Hazard Mater 170(2-3):1006–1013
Trofimov SY, Rozanova MS (2003) Transformation of soil properties under the impact of oil pollution. Eurasian Soil Sci 36(Suppl. 1)
Ummara U, Noreen S, Afzal M, Zafar ZU, Akhter MS, Iqbal S et al (2022) Induced systemic tolerance mediated by plant-microbe interaction in maize (Zea mays L.) plants under hydrocarbon contamination. Chemosphere 290:133327
Varjani SJ, Rana DP, Jain AK, Bateja S, Upasani VN (2015) Synergistic ex-situ biodegradation of crude oil by halotolerant bacterial consortium of indigenous strains isolated from on shore sites of Gujarat, India. Int Biodeterior Biodegradation 103:116–124
Walling DE, Zhang Y, He Q (2011) Models for deriving estimates of erosion and deposition rates from fallout radionuclide (caesium-137, excess lead-210, and beryllium-7) measurements and the development of user friendly software for model implementation. In: Impact of soil conservation measures on erosion control and soil quality. IAEA-TECDOC-1665, pp 11–33
Wang C, Wang G, Liu W, Wu P (2011) The effects of plant-soil-enzyme interactions on plant composition, biomass and diversity of alpine meadows in the Qinghai-Tibetan plateau. Int J Ecol 2011:180926
Wang S, Zhang C, Lu G, Li F, Guo G (2016) Screening of herbaceous plants for peat-enhanced rehabilitation of contaminated soil with oily sludge. Int J Phytoremediation 18(1):62–68. https://doi.org/10.1080/15226514.2015.1058332
Wu M, Dick WA, Li W, Wang X, Yang Q, Wang T, Xu L, Zhang M, Chen L (2016) Bioaugmentation and biostimulation of hydrocarbon degradation and the microbial community in a petroleum-contaminated soil. Int Biodeterior Biodegradation 107:158–164
Yaman C (2020) Performance and kinetics of bioaugmentation, biostimulation, and natural attenuation processes for bioremediation of crude oil-contaminated soils. Processes 8(8):883
Zhang S, Hu Z, Wang H (2019) Metagenomic analysis exhibited the cometabolism of polycyclic aromatic hydrocarbons by bacterial community from estuarine sediment. Environ Int 129:308–319
Disclaimer
The authors hereby confirm that this work is original and has not been published elsewhere nor is it currently under consideration for publication elsewhere. All the authors have agreed to have seen and approved the manuscript for submission.
Funding
Support for this study was provided by the Department of Biotechnology (DBT), Government of India.
Author information
Authors and Affiliations
Contributions
B. B. has done the acquisition, analysis, interpretation of the data, and written the draft manuscript. R. K. has done the PICRUSt analysis and interpretation of the data. P. P. has conceptualized and designed the work and also prepared, reviewed, and finalized the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
All the experiments were carried out in laboratory and field by the researchers as part of sponsored research project work.
Consent for publication
All the authors approved of the publication of this manuscript in Environmental Science and Pollution Research (ESPR).
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhuyan, ., Kotoky, R. & Pandey, P. Impacts of rhizoremediation and biostimulation on soil microbial community, for enhanced degradation of petroleum hydrocarbons in crude oil-contaminated agricultural soils. Environ Sci Pollut Res 30, 94649–94668 (2023). https://doi.org/10.1007/s11356-023-29033-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-29033-3