Abstract
The new compound 4-hydroxy-1-[(4-nitrophenyl)sulfonyl]pyrrolidine-2-carboxyllic acid was obtained by the reaction of 4-hydroxyproline with 4-nitrobenzenesulfonyl chloride. The compound was characterized using single crystal X-ray diffraction studies. Spectroscopic methods including NMR, FTIR, ES-MS, and UV were employed for further structural analysis of the synthesized compound. The title compound was found to have crystallized in an orthorhombic crystal system with space group P212121. The S1-N1 bond length of 1.628 (2) Å was a strong indication of the formation of the title compound. The absence of characteristic downfield 1H NMR peak of pyrrolidine ring and the presence of S–N stretching vibration at 857.82 cm−1 on the FTIR are strong indications for the formation of the sulfonamide. The experimental study was complemented with computations at the B3LYP/6-311G + + (d,p) level of theory to gain more understanding of interactions in the compound at the molecular level. Noncovalent interaction, Hirsfeld surface analysis and interaction energy calculations were employed in the analysis of the supramolecular architecture of the compound. Predicted ADMET parameters, awarded suitable bioavailability credentials, while the molecular docking study indicated that the compound enchants promising inhibition prospects against dihydropteroate synthase, DNA topoisomerase, and SARS-CoV-2 spike.
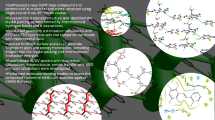
Graphical Abstract
Herein we present the solid state structure, noncovalent interaction and spectroscopic analysis of a prospective bioactive compound 4-hydroxy-1-[(4-nitrophenyl)sulphonyl]pyrrolidine-2-carboxyllic acid.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The relevance of sulfonamides or sulfa drugs in medicine can never be overemphasized [1] They have played significant roles in the medical revolution as they are very effective over a wide range of bacterial infections and thus were the first real success in the fight against bacteria [2, 3]. In asymmetric synthesis, the aryl sulfonyl group of aryl sulfonamides has been applied to protect nitrogen and oxygen functional groups as well as amino acids during conditions that involve the reaction of the carbonyl group with organometallic reagents [4, 5]. In addition, the presence of sulfonamide moiety in azo dyes has improved light stability, fixation to fiber, and water solubility [6, 7]. A lot of research on the activities of sulfonamides has uncovered their pharmacological applications not only as antibacterial [8, 9] but also as potent anticancer [10, 11], anti-influenza [12], anti-inflammatory[13], antitrypanosomal [14], antimalarial [15], anti-HIV [16], and antitumor [17] agents. The versatility of sulfonamides has led to the search for more active derivatives while applying different synthetic methodologies. Conventionally, sulfonamides are prepared by reacting either a primary or secondary amine with sulfonyl chloride in the presence of a base that assists to scavenge the HCl produced in the reaction [18]. Other methods also exist, some of which involve the use of transition metals as catalysts [19], N-chlorosulfonyl carbamate as a sulfonating agent at 0 °C [20,21,22], conversion of methyl sulfinates into sulfinamides, and subsequent oxidation to sulfonamides [23]. Milder synthetic conditions including the use of water as solvents for sulfonation of amines have been explored [24]. Recent reports on the beneficial properties of ionic liquids have also led to their use as an alternative solvent for sulfonamide synthesis [25,26,27]. These methods do however suffer some setbacks ranging from low reactivity, use of more catalysts to increase yield, metal contamination, and difficulty in catalyst recovery to complex purification steps. The use of amine as a nitrogen source and sulfonyl chloride to create the N–S bond is still preferred [28] due to its simplicity, good reactivity even at room temperature, and simple filtration process to obtain pure compounds [29]. Recently our group has been working on the synthesis and characterization of a number of sulfonyl and sulfonamide derivatives for various applications [30,31,32,33,34,35,36]. In continuation of this project, we report herein the synthesis and structural characterization of a novel sulfonamide derivative 4-hydroxy-1-[(4-nitrophenyl)sulfonyl]pyrrolidine-2-carboxyllic acid (HNPCA) from the coupling of 4-nitrobenzene sulfony chloride with the amino acid 4-hydroxyproline at room temperature using sodium carbonate in the presence of water, in-silico studies were used to understand the drug-likeness of the title compound.
Experimental
Materials
Na2CO3, NaCl, and 20% HCl were obtained from Fluka, while 4-nitrobenzene sulfonyl chloride and 4-hydroxyproline were obtained from Sigma Aldrich (Bristol Scientific Nigeria). IR spectra were recorded on Bruker FT-IR spectrophotometer. The NMR peaks were recorded on Bruker DPX 300 spectrophotometer with 1H at 400 MHz and 13 ℃ at 100 MHz. Chemical shifts δ are given in ppm and referenced to tetramethylsilane. Methanol was used as a solvent for crystallization. X-ray data of the single crystal were collected on an XtaLAB Synergy, Dualflex, Pilatus 200 K diffractometer using CuKα radiation at 105.4 (7)K. The pharmacokinetic properties were studied using SwissADME free online tool (https://www.swissadme.ch), while the Molecular docking was carried out using the Molecular operating environment (MOE) (AutoDock Vina, BIOVIA).
Synthesis of 4-Hydroxy-1-[(4-Nitrophenyl)Sulfonyl]Pyrrolidine-2-Carboxyllic Acid
Sodium carbonate (Na2CO3, 5 mmol) was added to a solution of 4-hydroxyproline (5 mmol) in water (15 mL) with continuous stirring until all the solutes dissolved. The solution was cooled to − 5 °C and 4-nitrobenzenesulfonyl chloride (5 mmol) was added in four portions over a period of 1 h. The reaction mixture was further stirred at room temperature for 4 h(Scheme 1). The mixture was acidified using 20% HCl until a pH of 2 was obtained. The white product obtained was filtered, washed, and dried in the open air. Yield (1.12 g, 71.34%), MP = 190–192 °(O), 1004 (C–N), 685 (S–N). 1H-NMR (DMSO, 400 MHz) δ: 8.40 (m, 2H, ArH), 8.10 (m, 2H, ArH), 4.87 (s, 1H, OH exchangeable with HDO), 4.31 (m, 2H, CH-CO2H and CH-OH), 3.59 (dd, 1H, CHa of CH2-CH-CO2H, J = 8.00, 4.00 Hz) 3.41 (dt, 1H, CHb of CH2-CH-CO2H, J = 8.00, 4.00 Hz), 2.19 (ddt, 1H, CH of CH2-N, J = 12.00, 8.00, 4.00 Hz), 2.06 (ddd, 1H, CHb of CH2-N, J = 12.00, 8.00,4.00 Hz).13C NMR (DMSO, 100 MHz) δ: 174.19 (C = O), 150.28, 143.40, 128.81, 123.79 (4 aromatic carbons), 69.21, 59.94, 56.51, 39.00 (4 aliphatic carbon). DEPT (100 MHz) δ: 174.19, 150.28, 143. 40 (tertiary C), 128.81, 123.79, 69.21, 59.94 (CH carbons), 56.51 and 39.00 (CH2 carbons).
X-Ray Determination of HNPCA
Dark green crystals of the titled compound were obtained from methanol under slow evaporation at room temperature. A single crystal suitable for X-ray diffraction measurement was mounted on a goniometer of an XtaLabSnergy, Dualflex Pilatus 200 K diffractometer. The crystal data were collected at 105.4 (7) K using graphite monochromated CuKα radiation (λ = 1,54,184) Table 1. Data reduction and empirical absorption correction were implemented in Crysallispro [37]. The structure was solved, with ShelXT [38], structure solution program using the intrinsic phasing solution method and Olex2 as the graphical interface. The structure was refined in ShelXL by full-matrix least-squares analysis of F2 against all reflections [39]. All non-hydrogen atoms were refined with anisotropic atomic displacement parameters. Unless otherwise noted, hydrogens were included in calculated positions. Atomic displacement parameters for the hydrogens were tied to the Ueq parameter of the atom to which they are bonded (1.5), for methyl, 1.2. for all others). N–H protons refined freely without any restraints. The molecular structural diagram was drawn in Olex2 Software Package [40]
Computational Method
The compound HNPCA was fully optimized in the gas phase using the DFT method. The functional used was B3LYP [41, 42] with the 6–311 + + G(d,p) basis set. HNPCA was also optimized in methanol using the same theoretical method within the PCM solvation model [43, 44] The ground-state structure was confirmed by frequency computations with the absence of imaginary frequency in both the gas phase and methanol. The optimized structure of HNPCA in methanol was used for computing chemical shifts with the Gauge-Including Atomic Orbital method [45] using isotropic shieldings of tetramethylsilane (TMS) computed using the same method. The non-covalent interaction (NCI), based on the reports of Johnson et al. [46], was explored by the noncovalent interaction-reduced density gradient (NCI-RDG) analysis using the Multiwfn program [47]. The isosurfaces were plotted using the Visual Molecular Dynamic (VMD) software [48] and The Gnuplot 4.2 program [49] and Ghostscript interpreter were employed to generate the 2D plots. All computations were carried out at 1 atm and 298.15 K using Gaussian 16 [50]. Visualization of the output files was done using GaussView 6 [51] and Chemcraft [52]. The coordinates of the optimized structure of HNPCA in the gas phase and methanol are provided in Table S1 of the supplementary information.
The interactions in the crystal structure of HNPCA were investigated by the Hirshfeld surface analysis [53] along with their 2D fingerprint plots [54] which were generated using the CrystalExplorer17.5 software [55]. Hirshfeld surface is represented by de and di, denoting the distance from the nearest atom outside and inside of the surface, respectively, and both are used to define the normalized contact distance (dnorm) with respect to the Van der Waals (vdW) radii as per Eq. 1.
For the visualization of dnorm, a red-blue-white color scale was selected. The red color denotes a negative value of the dnorm whereas the blue color denotes a positive value of the dnorm. The positive and negative values of the dnorm denote whether intermolecular interactions are larger or smaller than the vdW separation respectively. Therefore, the map** of dnorm on the Hirshfeld surface illustrates the donor and acceptor properties and helps in the analysis of the intermolecular interactions.
ADMET Prediction
The drug-likeness of HNPCA was predicted using Swiss free online ADMET tool (SwissADME; https://www.swissadme.ch). Calculated parameters include molecular weight, H-bond donor, H-bond acceptor, number of rotatable bonds, water partition coefficient (MlogP), and total polar surface area (TPSA).
Ligand and Protein Preparation
The compound was designed using ChembioDraw Professional 13.0 [56] and was converted to 2D by using BIOVIA Discover Studio Visualizer 17.2.0.16349 [57]. Structure optimization was achieved by applying the Hahn forcefield [58]. Optimized structures were used for the docking study. Crystal structure of dihydropteroate synthase, 5uoy [59], DNA topoisomerase II gyrase; 5mmn [60], and SARS-CoV-2 spike; 6vsb [61] were retrieved from protein data bank with resolutions 1.82 Å, 1.90 Å and 3.46 Å respectively. Protein editing was done by means of Discovery Studio[57] which included the deletion of co-crystallized ligands, multiple chains, hetero atoms, the water of crystallization, the addition of polar hydrogens, energy minimization, and structure optimization [58]. Enhanced proteins were used for molecular docking.
Ligand–Protein Docking Protocol Validation
The reliability of the docking protocol was corroborated by re-docking. Initially, the native ligand was withdrawn from protein binding site cavity and the binding site was defined by applying current ligand selection to obtain the ligand attributes; X = − 26.1303, Y = 11.7748, Z = − 17.6060, radius = 9.1985 for 5uoy, X = − 47.8198, Y = 3.2066, Z = 4.8274, radius = 8.6011 for 5mmn, and X = 190.2069, Y = 233.0658, Z = 210.6196, radius = 5.2447 for 6vsb. Docking was performed using Autodock Vina PyRx [62]. The native ligand was re-docked into the defined binding site and desired site attributes were carefully adjusted to fit the obtained Grid center. PyRx nonspecific algorithm was used to generate protein–ligand docked pose by applying Vina Wizard script. The lowest binding energy was used to select the best-docked model, while a comparison of H-bond and other key interactions selected the best-docked pose. The lower the binding energy, the higher the binding affinity. The RMSD of > 2 Å obtained for the superimposing of native crystal with the best-docked pose confirms the acceptability of the docking.
Results and Discussions
X-Ray Crystallography
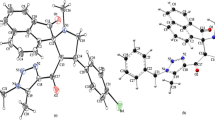
The molecular structure of the titled compound is shown in Fig. 1 The compound consists of a sulfonamide moiety sandwiched between a para-nitro phenyl moiety and hydroxy pyrrolidone carboxylic acid in a slightly distorted tetrahedral. The S1-N1 [1.628 (2) Å] bond length indicates the formation of the titled compound. This is similar to the reported value of 1.636 in the work of Murthy et al. [63]. The phenyl ring is planar to a rmsd 0.013 Å and disubstituted at para positions with nitro and sulfonyl groups. The N2-C4 and S1–C1 bonds are 1.472 (4) and 1.774 (3) Å respectively. These values are similar to the values reported for a related sulfonyl compound [64]. The pyrrolidine ring on the other hand is puckered to a rmsd of 0.182 Å and trisubstituted by carboxylic acid, hydroxyl, and sulfonyl groups. The dihedral angles between the sulfonyl groups and the paranitrophenyl and the five membered pyrrolidone rings respectively are 47.16 and 27.90°. The bond lengths of O4-C11 and O3-C11 are 1.331 and 1.200 Å respectively. The sulfonyl bonds S1-O1 and S1-O2 with lenghts 1.431(5) and 1.427(3) are similar to the values reported for related benzene sulfonyl compounds.[64, 65] As expected, the length of C = O (O3-C11) is shorter than that of C–OH (O4-C11). The angles around the slightly distorted tetrahedron range from a narrower O1-S1-N1 (106.21 (13)) to a slightly larger O1-S1-C1 [107.57 (13)]. The sulfonamide is stabilized by the torsion angles of O2-S1-N1-C10 [− 51.5 (2)˚] and O1-S1-NI-C7 [39.9 (2)˚]. The molecular aggregation structure of the solid-state structure of the compound in Fig. 2 consists of a 9-molecule aggregate synthon linked by a chain of O–H \(\cdots\) O hydrogen bonding interactions and stabilized by both intramolecular and bifurcated intermolecular C-H \(\cdots\) H short contacts in a 2-D supramolecular architecture. There are predominantly two types of O–H \(\cdots\) O intermolecular hydrogen bonding interactions, the carboxyl-O4-H4-O5(hydroxyl) and hydroxyl-O5-H5-O7(nitrophenyl). The hydrogen bond geometries are presented in Table 2.
Non Covalent Index NCI
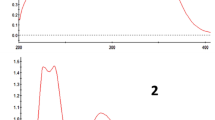
The stability of HNPCA can be determined based on its molecular interactions which can be analyzed using the RDG based on NCI. The NCI isosurface was generated using the B3LYP optimized geometry and this is illustrated in Fig. 3. The blue-green–red isosurfaces color are classified according to the strength and type of interaction whereby blue indicates strong attractive interactions, green indicates Van der Waals interaction and red indicates a strong repulsion[66]. Therefore, the presence of the green and red isosurfaces in Fig. 3 indicates the presence of Van der Waals interaction as well as steric interactions [67]. In addition to the NCI isosurface, the 2D NCI scatter plot is displayed in Fig. 4. The spikes in the RDG region where λ2 ≈ 0 and ρ ≈ 0, suggest that HPNCA consists of Van der Waals interactions (− 0.02 to 0 a. u.) mainly corresponding to the intramolecular O⋯H interactions. Spikes are also observed in the positive region (0 to 0.04 a. u.) indicating steric hindrance arising from the aromatic ring as well as the pyrrolidine ring. The absence of blue coloured spikes confirms that HNPCA does not exhibit any conventional intramolecular hydrogen bond within the structure but engages in intermolecular hydrogen bond.
Hirshfeld Surface Analysis
The Hirshfeld surface contacts were mapped over dnorm using the method described in [68,69,70] and presented in Fig. 5a. The red regions are located mainly on the oxygen atoms and one of the H atoms of the aromatic ring of the compound, indicating stronger intermolecular hydrogen bonds with neighboring atoms. However, the light-colored regions indicate weaker and longer contacts other than hydrogen bonds [34, 71, 72]. Some of the significant intermolecular interactions within the crystal structure of HNPCA are shown in Fig. 5b and these include phenyl-C-H⋯O(sulfonyl) in green dotted lines, pyrrolidone-C-H⋯O(nitro) in red dotted lines and phenyl-C⋯O interactions in magenta[73]. The fingerprint plots in Fig. 6, have been decomposed to analyze individual contributions and corroborate the results of the Hirshfeld surface analysis [54, 74]. The fingerprint plots indicate that the most dominant interaction is the O⋯H interactions (58.9%). The two sharp spikes correspond to the -OH group (O4-H) of the carboxylic group and the hydroxyl group (O5–H) of the pyrrolidone ring. The H⋯H interaction contributes to 19.9% of the surface area of the fingerprint plot. All other interactions observed are less than 8.0% [C⋯H (7.1%) > S⋯H (0.2%) > O⋯C (5.6%) > O⋯O (4.1%) > O⋯N (2.1%) > N⋯H (2.0%) > C⋯C (0.1%)].
Interaction Energy Calculations
The strength of intermolecular interactions can be estimated by calculating the interaction energies between pairs of molecules within the crystal of HNPCA and adding up the resulting four energy components comprising of the electrostatic (Eele), polarization (Epol), dispersion(Edis), and exchange repulsion (Erep) energies [75]. The energies were obtained by calculating the wave function of each pair of molecules or atoms at the B3LYP/6-31G(d,p) level of theory [69, 75]. Quantitative estimation of the strength and nature of the intermolecular interactions in HNPCA crystal with individual energy components (Eele, Epol, Edis, and Erep) as well as the sum of the energy components, Etot are presented in Table 3. The energy values show that the electrostatic component of the energy makes the most significant contribution to the total interaction energy profile in the crystal structure probably due to the intermolecular Van der Waals interactions resulting from the high level of H \(\cdots\) O/H \(\cdots\) O and H \(\cdots\) H intermolecular interactions. The dispersive interaction energy component of the total interaction energy is the second highest contributor to the total interaction energy and these probably result from the π \(\cdots\) π interaction of successive pairwise phenyl and pyrrolidone rings in the crystal lattice [70, 76]. A graphical representation of the magnitude of the interaction energies is presented in Fig. 7a–d in the form of energy frameworks to show the supramolecular architecture using cylindrical poles joining the centroids of molecular pairs. The red, green, and blue color-coded frameworks in 7a, 7b, and 7c respectively represent Eele, Edis, and Etot, energy components for intermolecular interactions HNPCA crystal, while 7d is the annotated Etot energy. The magnitude of the cylindrical poles corroborates the results from the fingerprint plots indicating that the Eele energy component to the total interaction energy is significant and contributes substantially to the molecular packing in the crystal lattice.
Perspective views of the energy frameworks of the title compound showing a electrostatic, b dispersion, c total energy, and d annotated total energy. The cylindrical radius is proportional to the relative strength of the corresponding energies and they were adjusted to the same scale factor of 100 with a cut-off value of 5 kJmol−1 within a 2 × 2 × 2 unit cells
Spectroscopic Studies
IR Spectra
The Infrared spectrum of the compound being investigated was recorded in the 4000–600 region using Bruker FT-IR spectrophotometer. The spectra of the compound (Figure S1) shows an S–N stretching prevalent for benzenesulfonamide derivatives at 857 cm−1(usually in the range of 947–836 cm-1) [77], and the band at 1355 cm−1 was assigned to S = O indicating the formation of the benzenesulfonamide. Other characteristics bands present are; the C = O stretching of a carboxylic at 1745.19 cm−1, N–O stretching for a nitro compound at 1524.70 cm−1, O–H stretching at 3450 and 2700 cm−1assigned to the OH of the alcohol and carboxylic acid respectively, C–H stretching at 2950 and 3100 cm−1 for the aliphatic and aromatic carbons respectively. The simulated IR spectrum is provided in Figure S2 and describes a detailed vibrational analysis of HNPCA in the gas phase as performed based on potential energy distribution using VEDA 4 program.
1H NMR,13C NMR, and DEPT
The 1H and 13C NMR of the named compound are presented in Figures S4 and S5 of the supplementary information. 1H NMR chemical shift at 8.38–8.41 were assigned to the protons present in the aromatic ring. The chemical shift at 8.08–8.11 were assigned to the protons of the pyrrolidine ring. Also, the absence of a chemical shift for N–H at the upper field confirms the formation of the desired compounds. The 13C NMR like the 1H NMR agrees with the crystal structure of the compound gotten via X-ray diffraction studies of the compound’s crystal. The signal at 174.19 was assigned to C = O, signals between 150.28 and 123.79 were assigned to the six aromatic carbon atoms, while the signals from 69.21 to 39.00 were assigned to the four aliphatic carbons of the compound. The NMR values are corroborated by the calculated NMR values presented in Table S1 of the supplementary information. The DEPT analysis (Figure S6) showed that the compound had three tertiary carbons, six CH carbons, and two CH2 carbons. Although only four carbons appeared among the CH carbons, this was expected due to chemical equivalence in the para-substituted benzene ring. There was also no appearance of methyl carbon in the DEPT results. These observations are in agreement with the structure obtained from XRD analysis.
ADMET
Due to poor bioavailability, most orally administered drugs are unable to reach the target therapeutic site, resulting in failed clinical trials despite promising in vitro and in vivo efficacy [78]. lipniski rule of five (MW ≤ 500, HBD ≤ 5, HBA ≤ 10, log p ≤ 5, RBC ≤ 10) is a valid assessment of the oral bioavailability of a drug candidate [79]. A molecule that violates more than two categories is adjudged to elicit poor bioavailability. HNPCA was studied for lipinski rule compliance and other properties that influence drug absorption, and the result is presented in Table 4. HNPCA showed total compliance in all calculated parameters except for TPSA (Veber rule: TPSA ≤ 140 Å2)[80], signaling a good bioavailability potential.
Molecular Docking Study
Molecular docking permits the prediction of molecular interaction between ligand–protein and has become a reliable tool in drug discovery. A docking study on HNPCA was done with selected protein targets; dihydropteroate synthase (5uoy), topoisomerase II DNA gyrase (5mmn), and a 2019-nCoV spike (6vsb) Fig 8. Dihydropteroate synthase (DHPS) catalyzes the transformation of 6-hydroxymethyl-7,8-dihydropterin pyrophosphate (DHPP) plus p-aminobenzoic acid (PABA) to 7,8-dihydropteroate, important folate intermediate in the folate biosynthetic pathway [81]. As a result, DHPS is an important antibacterial drug target and research on new inhibitors has intensified in recent times [82,83,84]. DNA gyrase is a type II topoisomerase that functions in DNA supercoils and replication, hence is investigated as a drug target for many agents, including antitumor and antibacterial [85]. The CoV spike (S) glycoprotein is a vital target for vaccines, and therapeutic antibodies [61]. HNPCA was docked in the active site of augmented protein targets using the intrinsic PyRx algorithm and obtained ligand–protein docked poses were visualized with Discovery Studio.
Binding Affinity of HNPCA with the Protein Target
The binding energy of HNPCA with protein targets had been shown in Table 5. In the case of 5uoy and 5mmn, HNPCA interacted significantly with receptor site residues, thereby eliciting higher or comparable binding affinity with the native ligand, ciprofloxacin or gentamicin. This suggests that the compound could intercalate well with the studied antibacterial targets to demonstrate promising efficacy. A lower binding affinity was recorded with 6vsb, compared to the crystal ligand, remdesivir or dexamethasone, which have been indicated to reduce mortality in people with severe COVID-19 [86].
Bonding Model HNPCA with Protein Target
H-bond is an important interaction stabilizing ligands with receptors. In the binding pocket of 5uoy, Fig 9, HNPCA demonstrated extensive H-bond between the COOH, OH, NO2, SO2-NR2 moieties and Thr62, Arg63, Glu60, Asn22, Arg255, His257, and Lys221 residues, Fig. 8. In addition, a closer look showed a pi-bonding between the O = S = O group with His257. The extensive H-bonding may have conferred a higher binding affinity, indicating a reliable stability with protein target active site residues to impact promising antibacterial activity.
In the case of 5mmn, the NO2 group of HNPCA engaged in H-bond with Arg76, Gly77, Thr165, while the COOH group approached Asn46 in H-bond. Additional stability was supported by the phenyl ring, which had invested in Ile78 and Glu50 via pi-alkyl and pi-anion interactions. These key interactions providing stability encourage potential strong inhibitory efficacy against microbial strains, Fig. 8.
On the other hand, the COOH, SO2, and NO2 groups established H-bond with Glu619, Thr618, Ser591, Thr553, and Asn536 amino acid residues in the binding cavity of 6vsb, while the phenyl group added stability by engaging in hydrophobic interaction with Val558, Fig. 8.
Conclusion
A novel compound 4-hydroxy-1-[(4-nitrophenyl)sulfonyl]pyrrolidine-2-carboxyllic has been synthesized and characterized using XRD and spectroscopic techniques. Intermolecular O4-H4 \(\cdots\) O5 interaction in the crystal structure of the compound connects different molecules of the compound in a continuous chain and is stabilized by N4-H4 \(\cdots\) O3 intermolecular hydrogen bonding interaction. Hirshfeld surface analysis of the intermolecular interactions suggests that O \(\cdots\) H/H \(\cdots\) O interactions contributed more than half of the total intermolecular contacts in the compound. The NCI analysis of the compound did not reveal any intermolecular hydrogen bonding interactions in the compounds. There was however evidence of the presence of some Van der waals and steric interactions, corroborated by the results of the interaction energy calculations which showed higher electrostatic interactions than dispersive forces. The results of the molecular docking studies showed that the compound had a high binding affinity and is well stabilized in the active pocket of dihydropteroate synthase, DNAtopoisomerase II gyrase, and SARS-CoV-2 spike via extensive H-bond network, which suggests that the compound may be a starting point for a rational drug design.
Code Availability
All softwares and codes used are open source and can be obtained freely online.
References
Supuran CT (2017) Sulfonamides. Molecules 22(10):1642
Davenport D (2012) The war against bacteria: how were sulphonamide drugs used by Britain during World War II? Med Humanit 38(1):55–58
Igwe CN, Okoro UC (2014) Synthesis, characterization, and evaluation for antibacterial and antifungal activities of n-heteroaryl substituted benzene sulphonamides. Org Chem Int. https://doi.org/10.1155/2014/419518
Harmata M, Zheng P, Huang C, Gomes MG, Ying W, Ranyanil K-O, Balan G, Calkins NL (2007) Expedient synthesis of sulfinamides from sulfonyl chlorides. J Org Chem 72(2):683–685
O’Connell JF, Rapoport H (1992) 1-Benzenesulfonyl-and 1-p-toluenesulfonyl-3-methylimidazolium triflates: efficient reagents for the preparation of arylsulfonamides and arylsulfonates. J Org Chem 57(17):4775–4777
Cui Z, Yang J, Chen W, Zhang S (2010) Dyeing fine denier polypropylene fibers with phenylazo-β-naphthol-containing sulfonamide disperse dyes. Front Chem Eng China 4(3):328–335
Hansch C, Sammes PG, Taylor JB (1990) Comprehensive medicinal chemistry: the rational design, mechanistic study & therapeutic applications of chemical compounds, vol 5. Pergamon Press, Robert Maxwell
Basanagouda M, Shivashankar K, Kulkarni MV, Rasal VP, Patel H, Mutha SS, Mohite AA (2010) Synthesis and antimicrobial studies on novel sulfonamides containing 4-azidomethyl coumarin. Eur J Med Chem 45(3):1151–1157
Eze FU, Okoro UC, Ukoha PO, Ugwu DI, Okafor SN (2021) New antioxidant agents bearing carboxamide moiety: synthesis, molecular docking and in vitro studies of new benzenesulfonamide derivatives. Iran J Chem Chem Eng 40(3):853–865
Altoparlak U, Kadanali A, Celebi S (2004) Slime factor positivity in coagulase negative staphylococci isolated from nasal samples of haemodialysis patients. Int J Clin Pract 58(12):1112–1114
Ghorab MM, Ragab FA, Heiba HI, Arafa RK, El-Hossary EM (2010) In vitro anticancer screening and radiosensitizing evaluation of some new quinolines and pyrimido [4, 5-b] quinolines bearing a sulfonamide moiety. Eur J Med Chem 45(9):3677–3684
Tang G, Lin X, Qiu Z, Li W, Zhu L, Wang L, Li S, Li H, Lin W, Yang M (2011) Design and synthesis of benzenesulfonamide derivatives as potent anti-influenza hemagglutinin inhibitors. ACS Med Chem Lett 2(8):603–607
Lima LdM, Castro P, Machado AL, Fraga CAM, Lugnier C, De Moraes VLG, Barreiro EJ (2002) Synthesis and anti-inflammatory activity of phthalimide derivatives, designed as new thalidomide analogues. Bioorg Med Chem 10(9):3067–3073
Ugwu DI, Okoro UC, Mishra NK (2018) Synthesis of proline derived benzenesulfonamides: a potent anti-Trypanosoma brucei gambiense agent. Eur J Med Chem 154:110–116
Agrawal V, Sinha S, Bano S, Khadikar P (2001) QSAR studies on antimalarial 2, 4-diamino-6-quinazoline sulfonamides. Acta Microbiol Immunol Hung 48(1):17–26
Selvam P, Murugesh N, Chandramohan M, Debyser Z, Witvrouw M (2008) Design, synthesis and antiHIV activity of novel isatine-sulphonamides. Indian J Pharm Sci 70(6):779
Owa T, Yoshino H, Okauchi T, Yoshimatsu K, Ozawa Y, Sugi NH, Nagasu T, Koyanagi N, Kitoh K (1999) Discovery of novel antitumor sulfonamides targeting G1 phase of the cell cycle. J Med Chem 42(19):3789–3799
Yasuhara A, Kameda M, Sakamoto T (1999) Selective monodesulfonylation of N, N-disulfonylarylamines with tetrabutylammonium fluoride. Chem Pharm Bull 47(6):809–812
Flegeau EF, Harrison JM, Willis MC (2016) One-Pot Sulfonamide synthesis exploiting the palladium-catalyzed sulfination of aryl iodides. Synlett 27(01):101–105
Barbey C, Bouasla R, Berredjem M, Dupont N, Retailleau P, Aouf N-E, Lecouvey M (2012) Synthesis and structural study of new substituted chiral sulfamoyl oxazolidin-2-ones. Tetrahedron 68(44):9125–9130
Berredjem M, Bouasla R, Aouf N-E, Barbey C (2010) Crystal structure of 4-phenyl-piperazine-1-sulfonamide. X-Ray Struct Anal Online 26:13–14
Bouasla R, Berredjem M, Aouf N-E, Barbey C (2008) 1, 2, 3, 4-Tetrahydroisoquinoline-2-sulfonamide. Acta Crystallogr Sect E 64(2):o432–o432
Ruano JLG, Parra A, Yuste F, Mastranzo VM (2008) Mild and general method for the synthesis of sulfonamides. Synthesis 2008(02):311–319
Eid N, Karamé I, Andrioletti B (2018) Straightforward and sustainable synthesis of sulfonamides in water under mild conditions. Eur J Org Chem 2018(36):5016–5022
Hall YD, Uzoewulu CP, Nizam ZM, Ishizawa S, El-Shaffey HM, Ohata J (2022) Phosphine-mediated three-component bioconjugation of amino- and azidosaccharides in ionic liquids. Chem Commun 58:10568–10571
Muthayala MK, Chhikara BS, Parang K, Kumar A (2012) Ionic liquid-supported synthesis of sulfonamides and carboxamides. ACS Comb Sci 14(1):60–65
Uzoewulu CP, Ohata J (2022) Translation of a phosphine- and azide-based reaction to chemical modification of biomolecules in ionic liquid. Synlett 33(19):1879–1883
Pandya R, Murashima T, Tedeschi L, Barrett AG (2003) Facile one-pot synthesis of aromatic and heteroaromatic sulfonamides. J Org Chem 68(21):8274–8276
Gioiello A, Rosatelli E, Teofrasti M, Filipponi P, Pellicciari R (2013) Building a sulfonamide library by eco-friendly flow synthesis. ACS Comb Sci 15(5):235–239
Aronimo BS, Okoro UC, Ali R, Ibeji CU, Ezugwu JA, Ugwu DI (2021) Synthesis, molecular docking and antimalarial activity of phenylalanine-glycine dipeptide bearing sulphonamide moiety. J Mol Struct 1246:131201
Eze FU, Okoro UC, Ugwu DI, Okafor SN (2019) Biological activity evaluation of some new benzenesulphonamide derivatives. Front Chem 7:634
Ezugwu JA, Okoro UC, Ezeokonkwo MA, Hariprasad KS, Rudrapal M, Ugwu DI, Gogoi N, Chetia D, Celik I, Ekoh OC (2022) Design, synthesis, molecular docking, molecular dynamics and in vivo antimalarial activity of new dipeptide-sulfonamides. ChemistrySelect 7(5):e202103908
Ekoh OC, Okoro U, Ugwu D, Ali R, Okafor S, Ugwuja D, Attah S (2022) Novel dipeptides bearing sulfonamide as antimalarial and antitrypanosomal agents: synthesis and molecular docking. Med Chem 18(3):394–405
Ozochukwu IS, Okpareke OC, Izuogu DC, Ibezim A, Ujam OT, Asegbeloyin JN (2021) N’-(Pyridin-3-ylmethylene) benzenesulfonohydrazide: crystal structure, dft, hirshfeld surface and in silico anticancer studies. Eur J Chem 12(3):256–264
Nwokelo MO, Izuogu DC, Okpareke OC, Ibeji CU, Oyeka EE, Lane JR, Asegbeloyin JN (2021) Structural, computational and antimicrobial studies of 2–[(E)–[2–(2, 4, 6-trimethylbenzenesulfonyl)-hydrazinylidene] methyl] benzoic acid and its Cu (II), Zn (II) and Co (II) complexes. J Mol Struct 1225:129019
Asegbeloyin JN, Izuogu DC, Oyeka EE, Okpareke OC, Ibezim A (2019) Crystal structure, non-covalent interaction and molecular docking studies of 2-{[2-phenylsulfonyl) hydrazinylidene] methyl} benzoic acid and its dysprosium catalysed cyclized product: 2-(phenyl-sulfonyl) phthalazin-1 (2H)-one. J Mol Struct 1175:219–229
Diffraction, R. O. ((2015)). CrysAlisPro Software System, version 1.171. 38.41 l, .
Sheldrick GM (2015) SHELXT–Integrated space-group and crystal-structure determination. Acta Crystallogr Sec A 71(1):3–8
Sheldrick GM (2015) Crystal structure refinement with SHELXL. Acta Crystallogr Sec C 71(1):3–8
Dolomanov OV, Bourhis LJ, Gildea RJ, Howard JAK, Puschmann H (2009) OLEX2: a complete structure solution, refinement and analysis program. J Appl Crystallogr 42(2):339–341
Becke AD (1993) A new mixing of Hartree-Fock and local density-functional theories. J Chem Phys 98(2):1372–1377
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37(2):785
Simkin BIAk, Sheĭkhet IIi (1995) Quantum chemical and statistical theory of solutions: a computational approach. Ellis Horwood, Newyork
Tomasi J, Persico M (1994) Molecular interactions in solution: an overview of methods based on continuous distributions of the solvent. Chem Rev 94(7):2027–2094
Wolinski K, Hinton JF, Pulay P (1990) Efficient implementation of the gauge-independent atomic orbital method for NMR chemical shift calculations. J Am Chem Soc 112(23):8251–8260
Johnson ER, Keinan S, Mori-Sánchez P, Contreras-García J, Cohen AJ, Yang W (2010) Revealing noncovalent interactions. J Am Chem Soc 132(18):6498–6506
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33(5):580–592
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Williams, T., Kelley, C., Bröker, HB., Campbell, J., Cunningham, R., Denholm, D., Zellner, J. (2012). gnuplot 4.6. An Interactive Plotting Program.
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Petersson GA, Nakatsuji H, Li X, Caricato M, Marenich AV, Bloino J, Janesko BG, Gomperts R, Mennucci B, Hratchian HP, Ortiz JV, Izmaylov AF, Sonnenberg JL, Williams-Young D, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski VG, Gao J, Rega N, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark MJ, Heyd JJ, Brothers EN, Kudin KN, Staroverov VN, Keith TA, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Adamo C, Cammi R, Ochterski JW, Martin RL, Morokuma K, Farkas O, Foresman JB, Fox DJ (2016) Gaussian 16 Revision C. 01. 2016. Gaussian Inc, Wallingford CT
Dennington R, Keith TA, Millam JM (2016) GaussView, version 6.0. 16. Semichem Inc, Shawnee Mission KS
Zhurko, G., & Zhurko, D. (2013). Chemcraft, Version 1.7 (build 365). URL:http://www. chemcraftprog. com (accessed March 27, 2016).
McKinnon JJ, Jayatilaka D, Spackman MA (2007) Towards quantitative analysis of intermolecular interactions with Hirshfeld surfaces. Chem Commun 5(37):3814–3816
Spackman MA, McKinnon JJ (2002) Fingerprinting intermolecular interactions in molecular crystals. CrystEngComm 4(66):378–392
Turner, M., McKinnon, J., Wolff, S., Grimwood, D., Spackman, P., Jayatilaka, D., & Spackman, M. (2017). CrystalExplorer17. University of Western Australia.
Narayanaswamy VK, Rissdörfer M, Odhav B (2013) Review on CambridgeSoft ChemBioDraw Ultra 13.0 v. Int J Theor Appl Sci 5:45–49
BIOVIA, D. S. (2017) BIOVIA discovery studio visualizer. Software version 20:779
Hahn M (1995) Receptor surface models. 1. Definition and construction. J Med Chem 38(12):2080–2090
Dyck B, Branstetter B, Gharbaoui T, Hudson AR, Breitenbucher JG, Gomez L, Botrous I, Marrone T, Barido R, Allerston CK (2017) Discovery of selective phosphodiesterase 1 inhibitors with memory enhancing properties. J Med Chem 60(8):3472–3483
Panchaud P, Bruyere T, Blumstein A-C, Bur D, Chambovey A, Ertel EA, Gude M, Hubschwerlen C, Jacob L, Kimmerlin T (2017) Discovery and optimization of isoquinoline ethyl ureas as antibacterial agents. J Med Chem 60(9):3755–3775
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh C-L, Abiona O, Graham BS, McLellan JS (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367(6483):1260–1263
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461
Murthy PK, Suneetha V, Armaković S, Armaković SJ, Suchetan P, Giri L, Rao RS (2018) Synthesis, characterization and computational study of the newly synthetized sulfonamide molecule. J Mol Struct 1153:212–229
Khan BA, Hamdani SS, Ahmed MN, Hameed S, Ashfaq M, Shawky AM, Ibrahim MA, Sidhom PA (2022) Synthesis, X-ray diffraction analysis, quantum chemical studies and α-amylase inhibition of probenecid derived S-alkylphthalimide-oxadiazole-benzenesulfonamide hybrids. J Enzyme Inhib Med Chem 37(1):1464–1478
Ashfaq M, Tahir MN, Munawar KS, Behjatmanesh-Ardakani R, Kargar H (2022) Single crystal exploration, supramolecular behaviour, Hirshfeld surface analysis, linear and non-linear theoretical optical properties of Schiff bases derived from Benzene sulfonamides. J Mol Struct 1261:132952
Zhang Y, Huang J, Qiao L, Zhang X, Cao W, Ding Z, Hang X, Qin B, Song J (2018) Investigations based on non-covalent interactions in 1-(4-chloromethylbenzoyl)-3-(4, 6-di-substituted pyrimidine-2-yl) thioureas: Synthesis, characterizations and quantum chemical calculations. J Mol Struct 1169:85–95
Okpareke OC, Henderson W, Lane JR, Okafor SN (2020) Synthesis, structure, computational and molecular docking studies of asymmetrically di-substituted ureas containing carboxyl and phosphoryl hydrogen bond acceptor functional groups. J Mol Struct 1203:127360
Spackman MA, Jayatilaka D (2009) Hirshfeld surface analysis. CrystEngComm 11(1):19–32
Tan SL, Jotani MM, Tiekink ER (2019) Utilizing Hirshfeld surface calculations, non-covalent interaction (NCI) plots and the calculation of interaction energies in the analysis of molecular packing. Acta Crystallogr Sec E 75(3):308–318
Ayiya BB, Okpareke OC (2021) N, N′-Di (pyridine-4-yl)-pyridine-3, 5-dicarboxamide, a pincer-type tricationic compound; synthesis, crystal structure, hirshfeld surface analysis, and computational chemistry studies. J Chem Crystallogr 52:1–11
Ifeanyieze KJ, Ayiya BB, Okpareke OC, Groutso TV, Asegbeloyin JN (2022) Crystal structure, Hirshfeld surface and computational study of 1-(9, 10-dioxo-9, 10-dihydroanthracen-1-yl)-3-propanoylthiourea. Acta Crystallogr Sec E 78(4):439
Ekowo LC, Eze SI, Ezeorah JC, Groutso T, Atiga S, Lane JR, Okafor S, Akpomie KG, Okparaeke OC (2020) Synthesis, structure, Hirshfeld surface, DFT and in silico studies of 4-[(E)-(2, 5-dimethoxybenzylidene) amino]-1, 5-dimethyl-2-phenyl-1, 2-dihydro-3H-pyrazol-3-one (DMAP) and its metal complexes. J Mol Struct 1210:127994
Ali A, Khalid M, Rehman MFu, Haq S, Ali A, Tahir MN, Ashfaq M, Rasool F, Braga AAC (2020) Efficient synthesis, SC-XRD, and theoretical studies of o-benzenesulfonylated pyrimidines: role of noncovalent interaction influence in their supramolecular network. ACS Omega 5(25):15115–15128
Khalid M, Ali A, Haq S, Tahir MN, Iqbal J, Braga AA, Ashfaq M, Akhtar SUH (2021) O-4-Acetylamino-benzenesulfonylated pyrimidine derivatives: synthesis, SC-XRD, DFT analysis and electronic behaviour investigation. J Mol Struct 1224:129308
Mackenzie CF, Spackman PR, Jayatilaka D, Spackman MA (2017) CrystalExplorer model energies and energy frameworks: extension to metal coordination compounds, organic salts, solvates and open-shell systems. IUCrJ 4(5):575–587
Tan SL, Tiekink ER (2020) The 1: 2 co-crystal formed between N, N′-bis (pyridin-4-ylmethyl) ethanediamide and benzoic acid: crystal structure, Hirshfeld surface analysis and computational study. Acta Crystallogr Sec E 76(1):102–110
Tanaka Y, Tanaka Y (1965) Infrared absorption spectra of organic sulfur compounds. II. Studies on SN stretching bands of methanesulfonamide derivatives. Chem Pharm Bull 13(7):858–861
Buxton, I. (2006). Goodman & Gilman’s The Pharmacological Basis of Therapeutics; Pharmacokinetics and Pharmacodynamics: The Dynamics of Drug Absorption, Distribution, Action and Elimination: Introduction. by Brunton LL, Lazo JS, Parker KL, McGraw-Hill, New York, 1–39.
Lipinski CA (2000) Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol Toxicol Method 44(1):235–249
Veber DF, Johnson SR, Cheng H-Y, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45(12):2615–2623
Wolff ME (1996) Burger’s medicinal chemistry and drug discovery. Am J Ther 3(8):608
Babaoglu K, Qi J, Lee RE, White SW (2004) Crystal structure of 7, 8-dihydropteroate synthase from Bacillus anthracis: mechanism and novel inhibitor design. Structure 12(9):1705–1717
Hevener KE, Yun M-K, Qi J, Kerr ID, Babaoglu K, Hurdle JG, Balakrishna K, White SW, Lee RE (2010) Structural studies of pterin-based inhibitors of dihydropteroate synthase. J Med Chem 53(1):166–177
Pemble CW IV, Mehta PK, Mehra S, Li Z, Nourse A, Lee RE, White SW (2010) Crystal structure of the 6-hydroxymethyl-7, 8-dihydropterin pyrophosphokinase• dihydropteroate synthase bifunctional enzyme from Francisella tularensis. PLoS ONE 5(11):e14165
Maxwell A (1997) DNA gyrase as a drug target. Trends Microbiol 5(3):102–109
Siemieniuk RA, Bartoszko JJ, Ge L, Zeraatkar D, Izcovich A, Kum E, Pardo-Hernandez H, Qasim A, Martinez JPD, Rochwerg B (2020) Drug treatments for covid-19: living systematic review and network meta-analysis. BMJ 370:m2980
Author information
Authors and Affiliations
Contributions
Category 1 Conception and design of the study: DIU, FUE, CJE Acquisition of data: Blessing CO, TG, OCO, CCE Analysis and interpretation of data: OCO, PR, LR, Supervision of research: DIU, FUE, Revision of final manuscript. CCE, CPU, OCO, Category 2 Drafting the manuscript: CPU, OCO, Revising the manuscript critically for important intellectual content: DIU, FUE, CJE. BCO, OCO, CCE, PR, CPU, Category 3 Approval of the version of the manuscript to be published; DIU, FUE, CJE. BCO, OCO, CCE, PR, CPU.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10870_2023_978_MOESM1_ESM.docx
Supplementary file1 (DOCX 929 KB)—CCDC 2216313 contains the supplementary crystallographic data for HNPCA. These data can be obtained free of charge via http://www.ccdc.cam.ac.uk/conts/retrieving.html (or from the Cambridge Crystallographic Data Centre, 12, Union Road, Cambridge CB2 1EZ, UK; fax: +44 1223 336033)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ugwu, D.I., Eze, F.U., Ezeorah, C.J. et al. Synthesis, Structure, Hirshfeld Surface Analysis, Non-Covalent Interaction, and In Silico Studies of 4-Hydroxy-1-[(4-Nitrophenyl)Sulfonyl]Pyrrolidine-2-Carboxyllic Acid. J Chem Crystallogr 53, 386–399 (2023). https://doi.org/10.1007/s10870-023-00978-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10870-023-00978-0