Abstract
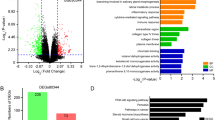
Lung cancer is one of the deadliest malignant tumors with non-small cell lung cancer (NSCLC) being the most prevalent type. Patients with NSCLC usually were diagnosed at the advance clinical stages, and these patients often had high rate of tumor-recurrence, thus leading to poor prognosis. Yet, the molecular mechanisms underlying NSCLC progression and recurrence are largely unknown. This study aimed to identify potential hub genes associated with the pathophysiology of NSCLC by bioinformatics analysis and laboratory validation. The GSE51852, GSE52248 and GSE75037 datasets were downloaded from the Gene Expression Omnibus database. The overlap** differentially expressed genes (DEGs) were analyzed by GEO2R tool. Gene Ontology (GO) and KEGG pathway enrichment analysis were performed on these overlap** DEGs. The protein-protein interaction network was constructed to identify hub genes from DEGs. The expression and survival analysis of these hub genes were performed by using the integrated bioinformatics tools. Finally, the effects of GOLM1 on the proliferation and chemo-sensitivity of NSCLC cells were determined by in vitro functional assays. A total of 197 overlap** DEGs (37 up-regulated and 160 down-regulated) were identified from the microarray datasets. Furthermore, the PPI network with 89 nodes and 768 edges was constructed and 17 hub genes were identified from PPI network by using MCODE analysis. The survival analysis revealed that the expression of 5 hub genes (FGF2, GOLM1, GPC3, IL6 and SPP1) were significantly correlated with overall survival of patients with lung cancer. Furthermore, the in vitro functional studies showed that GOLM1 overexpression promoted the NSCLC cell proliferation and colony formation; while GOLM1 knockdown exerted the opposite effects. Importantly, GOLM1 overexpression reduced the chemo-sensitivity of cisplatin in NSCLC cells by attenuating the inhibitory effects of cisplatin on the cell proliferation and colony formation. In conclusion, the present study showed that 5 hub genes including FGF2, GOLM1, GPC3, IL6 and SPP1 were deregulated in NSCLC tissues and may predict the prognosis of patients with NSCLC. GOLM1 may play an important role in regulating the cell proliferation and chemo-sensitivity of cisplatin in NSCLC.









Similar content being viewed by others
Data availability
All the data are available upon request from the corresponding author.
References
Aruna, Li LM (2018) Overexpression of golgi membrane protein 1 promotes non-small-cell carcinoma aggressiveness by regulating the matrix metallopeptidase 13. Am J Cancer Res 8(3):551–565
Boldrini L, Donati V, Dell’Omodarme M, Prati MC, Faviana P, Camacci T et al (2005) Prognostic significance of osteopontin expression in early-stage non-small-cell lung cancer. Br J Cancer 93(4):453–457. https://doi.org/10.1038/sj.bjc.6602715
Ding X, Deng G, Liu J, Liu B, Yuan F, Yang X et al (2019) GOLM1 silencing inhibits the proliferation and motility of human glioblastoma cells via the Wnt/β-catenin signaling pathway. Brain Res 1717:117–126. https://doi.org/10.1016/j.brainres.2019.03.035
Duma N, Santana-Davila R, Molina JR (2019) Non-small cell lung cancer: epidemiology, screening, diagnosis, and treatment. Mayo Clin Proc 94(8):1623–1640. https://doi.org/10.1016/j.mayocp.2019.01.013
Gao SP, Mark KG, Leslie K, Pao W, Motoi N, Gerald WL et al (2007) Mutations in the EGFR kinase domain mediate STAT3 activation via IL-6 production in human lung adenocarcinomas. J Clin Invest 117(12):3846–3856. https://doi.org/10.1172/jci31871
Hirsch FR, Scagliotti GV, Mulshine JL, Kwon R, Curran WJ Jr, Wu YL et al (2017) Lung cancer: current therapies and new targeted treatments. Lancet 389(10066):299–311. https://doi.org/10.1016/s0140-6736(16)30958-8
Hossain MN, Sakemura R, Fujii M, Ayusawa D (2006) G-protein gamma subunit GNG11 strongly regulates cellular senescence. Biochem Biophys Res Commun 351(3):645–650. https://doi.org/10.1016/j.bbrc.2006.10.112
Hou XL, Ji CD, Tang J, Wang YX, **ang DF, Li HQ et al (2017) FPR2 promotes invasion and metastasis of gastric cancer cells and predicts the prognosis of patients. Sci Rep 7(1):3153. https://doi.org/10.1038/s41598-017-03368-7
Hsu YL, Hung JY, Lee YL, Chen FW, Chang KF, Chang WA et al (2017) Identification of novel gene expression signature in lung adenocarcinoma by using next-generation sequencing data and bioinformatics analysis. Oncotarget 8(62):104831–104854. https://doi.org/10.18632/oncotarget.21022
Ishiguro T, Ohata H, Sato A, Yamawaki K, Enomoto T, Okamoto K (2017) Tumor-derived spheroids: Relevance to cancer stem cells and clinical applications. Cancer Sci 108(3):283–289. https://doi.org/10.1111/cas.13155
**g B, Wang T, Sun B, Xu J, Xu D, Liao Y et al (2020) IL6/STAT3 signaling orchestrates premetastatic niche formation and immunosuppressive traits in lung. Cancer Res 80(4):784–797. https://doi.org/10.1158/0008-5472.can-19-2013
Karasaki T, Nagayama K, Kuwano H, Nitadori JI, Sato M, Anraku M et al (2017) An immunogram for the cancer-immunity cycle: towards personalized immunotherapy of lung cancer. J Thorac Oncol 12(5):791–803. https://doi.org/10.1016/j.jtho.2017.01.005
Ko EC, Raben D, Formenti SC (2018) The integration of radiotherapy with immunotherapy for the treatment of non-small cell lung cancer. Clin Cancer Res 24(23):5792–5806. https://doi.org/10.1158/1078-0432.ccr-17-3620
Li Y, Sun N, Lu Z, Sun S, Huang J, Chen Z et al (2017) Prognostic alternative mRNA splicing signature in non-small cell lung cancer. Cancer Lett 393:40–51. https://doi.org/10.1016/j.canlet.2017.02.016
Li RM, Nai MM, Duan SJ, Li SX, Yin BN, An F et al (2018) Down-expression of GOLM1 enhances the chemo-sensitivity of cervical cancer to methotrexate through modulation of the MMP13/EMT axis. Am J Cancer Res 8(6):964–980
Liu Y, Chen K, Wang C, Gong W, Yoshimura T, Liu M et al (2013) Cell surface receptor FPR2 promotes antitumor host defense by limiting M2 polarization of macrophages. Cancer Res 73(2):550–560. https://doi.org/10.1158/0008-5472.can-12-2290
Liu X, Chen L, Zhang T (2018) Increased GOLM1 expression independently predicts unfavorable overall survival and recurrence-free survival in lung adenocarcinoma. Cancer Control 25(1):1073274818778001. https://doi.org/10.1177/1073274818778001
Lu J, Zhao J, Jia C, Zhou L, Cai Y, Ni J et al (2019) FPR2 enhances colorectal cancer progression by promoting EMT process. Neoplasma 66(5):785–791. https://doi.org/10.4149/neo_2018_181123N890
Mayo-de-Las-Casas C, Garzón Ibáñez M, Jordana-Ariza N, García-Peláez B, Balada-Bel A, Villatoro S et al (2018) An update on liquid biopsy analysis for diagnostic and monitoring applications in non-small cell lung cancer. Expert Rev Mol Diagn 18(1):35–45. https://doi.org/10.1080/14737159.2018.1407243
Migeotte I, Communi D, Parmentier M (2006) Formyl peptide receptors: a promiscuous subfamily of G protein-coupled receptors controlling immune responses. Cytokine Growth Factor Rev 17(6):501–519. https://doi.org/10.1016/j.cytogfr.2006.09.009
Mlakar V, Strazisar M, Sok M, Glavac D (2010) Oligonucleotide DNA microarray profiling of lung adenocarcinoma revealed significant downregulation and deletions of vasoactive intestinal peptide receptor 1. Cancer Invest 28(5):487–494. https://doi.org/10.3109/07357900903476752
Nakagawa H, Fujita M (2018) Whole genome sequencing analysis for cancer genomics and precision medicine. Cancer Sci 109(3):513–522. https://doi.org/10.1111/cas.13505
Niemira M, Collin F, Szalkowska A, Bielska A, Chwialkowska K, Reszec J et al (2019) Molecular signature of subtypes of non-small-cell lung cancer by large-scale transcriptional profiling: identification of key modules and genes by Weighted Gene Co-Expression Network Analysis (WGCNA). Cancers (Basel) 12(1). https://doi.org/10.3390/cancers12010037
Reuter JA, Spacek DV, Snyder MP (2015) High-throughput sequencing technologies. Mol Cell 58(4):586–597. https://doi.org/10.1016/j.molcel.2015.05.004
Riess JW, Gandara DR, Frampton GM, Madison R, Peled N, Bufill JA et al (2018) Diverse EGFR exon 20 insertions and co-occurring molecular alterations identified by comprehensive genomic profiling of NSCLC. J Thorac Oncol 13(10):1560–1568. https://doi.org/10.1016/j.jtho.2018.06.019
Shi K, Li N, Yang M, Li W (2019) Identification of key genes and pathways in female lung cancer patients who never smoked by a bioinformatics analysis. J Cancer 10(1):51–60. https://doi.org/10.7150/jca.26908
Shojaei F, Scott N, Kang X, Lappin PB, Fitzgerald AA, Karlicek S et al (2012) Osteopontin induces growth of metastatic tumors in a preclinical model of non-small lung cancer. J Exp Clin Cancer Res 31(1):26. https://doi.org/10.1186/1756-9966-31-26
Sriram KB, Larsen JE, Yang IA, Bowman RV, Fong KM (2011) Genomic medicine in non-small cell lung cancer: paving the path to personalized care. Respirology 16(2):257–263. https://doi.org/10.1111/j.1440-1843.2010.01892.x
Takauji Y, Kudo I, En A, Matsuo R, Hossain MN, Nakabayashi K et al (2017) GNG11 (G-protein subunit γ 11) suppresses cell growth with induction of reactive oxygen species and abnormal nuclear morphology in human SUSM-1 cells. Biochem Cell Biol 95(4):517–523. https://doi.org/10.1139/bcb-2016-0248
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98–w102. https://doi.org/10.1093/nar/gkx247
Varambally S, Laxman B, Mehra R, Cao Q, Dhanasekaran SM, Tomlins SA et al (2008) Golgi protein GOLM1 is a tissue and urine biomarker of prostate cancer. Neoplasia 10(11):1285–1294. https://doi.org/10.1593/neo.08922
Wang C, Tan S, Liu WR, Lei Q, Qiao W, Wu Y et al (2019) RNA-Seq profiling of circular RNA in human lung adenocarcinoma and squamous cell carcinoma. Mol Cancer 18(1):134. https://doi.org/10.1186/s12943-019-1061-8
Yan G, Ru Y, Wu K, Yan F, Wang Q, Wang J et al (2018) GOLM1 promotes prostate cancer progression through activating PI3K-AKT-mTOR signaling. Prostate 78(3):166–177. https://doi.org/10.1002/pros.23461
Yanagawa H, Sone S, Takahashi Y, Haku T, Yano S, Shinohara T et al (1995) Serum levels of interleukin 6 in patients with lung cancer. Br J Cancer 71(5):1095–1098. https://doi.org/10.1038/bjc.1995.212
Yang HJ, Liu GL, Liu B, Liu T (2018) GP73 promotes invasion and metastasis of bladder cancer by regulating the epithelial-mesenchymal transition through the TGF-β1/Smad2 signalling pathway. J Cell Mol Med 22(3):1650–1665. https://doi.org/10.1111/jcmm.13442
Ye QH, Zhu WW, Zhang JB, Qin Y, Lu M, Lin GL et al (2016) GOLM1 modulates EGFR/RTK cell-surface recycling to drive hepatocellular carcinoma metastasis. Cancer Cell 30(3):444–458. https://doi.org/10.1016/j.ccell.2016.07.017
Zhang X, Liu Y, Huang WC, Zheng LC (2018) MiR-125b-1-3p exerts antitumor functions in lung carcinoma cells by targeting S1PR1. Chin Med J (Engl) 131(16):1909–1916. https://doi.org/10.4103/0366-6999.238135
Zhang R, Zhu Z, Shen W, Li X, Dhoomun DK, Tian Y (2019) Golgi Membrane Protein 1 (GOLM1) promotes growth and metastasis of breast cancer cells via Regulating Matrix Metalloproteinase-13 (MMP13). Med Sci Monit 25:847–855. https://doi.org/10.12659/msm.911667
Zhao L, Yu Z, Zhao B (2019) Mechanism of VIPR1 gene regulating human lung adenocarcinoma H1299 cells. Med Oncol 36(11):91. https://doi.org/10.1007/s12032-019-1312-y
Zhu Y, Luo G, Jiang B, Yu M, Feng Y, Wang M et al (2018) Apolipoprotein M promotes proliferation and invasion in non-small cell lung cancers via upregulating S1PR1 and activating the ERK1/2 and PI3K/AKT signaling pathways. Biochem Biophys Res Commun 501(2):520–526. https://doi.org/10.1016/j.bbrc.2018.05.029
Funding
This study was supported by Hainan Province Health and Family Planning Industry Research Project (18A200148) and Sanya Medical and Health Project (2019YW09).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Figure S1
PPI network construction of overlap** DEGs derived from STRING database. (PNG 631 kb)
Supplementary Figure S2
The correlation of hub gene expression levels and the overall survival of patients with NSCLC was analyzed by KM Plotter. The analyzed hub genes include (A) FGF2, (B) GPBAR1, (C) GPC3, (D) PTH1R, (E) RAMP2, (F) RAMP3 and (G) SSTR1. The red lines represent patients with high gene expression, and black lines represent patients with a low gene expression. HR = hazard ratio. (PNG 445 kb)
Supplementary Figure S3
The correlation of hub gene expression levels and the overall survival of patients with NSCLC was analyzed by Human Protein Atlas database. The analyzed hub genes include (A) CALCRL, (B) CXC13, (C) GNG11, (D) VIPR1, (E) S1PR1, (F) FPR2, (G) GOLM1, (H) IL6 and (I) SPP1. The pink lines represent patients with high gene expression, and blue lines represent patients with a low gene expression. (PNG 681 kb)
Supplementary Table S1
(XLSX 11 kb)
Supplementary Table S2
(XLSX 19 kb)
Rights and permissions
About this article
Cite this article
Zhao, M., Li, X. & Chen, X. GOLM1 predicts poor prognosis of patients with NSCLC and is associated with the proliferation and chemo‐sensitivity of cisplatin in NSCLC cells: bioinformatics analysis and laboratory validation. J Bioenerg Biomembr 53, 177–189 (2021). https://doi.org/10.1007/s10863-021-09875-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-021-09875-7