Abstract
The aim of this study is to systematically appraise the evidence on available full thickness 3D gingival and mucosal models (3D culture in scaffold base system) and their application in periodontal and peri-implant research. This study involved a systematic review of twenty-two studies obtained from searching from five electronic databases: MEDLINE-OVID, EMBASE, EBSCOhost, Web of Science Core Collection and LILACS, as well as a hand search of eligible articles up to September 2022. A total of 2338 studies were initially identified, after removal of duplicates (573), abstracts/title selection (1765), and full text screening (95), twenty-two studies were included, thirty-seven models were identified. Several cellular markers were reported by the studies included. The expression of keratinocytes differentiation markers (K4, K5, K10, K13, K14, K16, K17, K18, K19, involucrin, laminin5), proliferation marker (Ki67, CD90), and vimentin, Type I, II and IV collagen produced by fibroblasts were investigated in thirty models. No quantitative analyses were performed, and results of the review confirmed a substantial level of heterogeneity across experiments. In conclusion, there is currently insufficient evidence to conclude that the available 3D gingival and mucosal models can entirely recapitulate the human gingival tissue/mucosa and provide a useful research tool for periodontal and peri-implant research. This review also highlighted the lack of a standardized protocol to construct and characterize 3D gingival models. A new protocol is proposed for the characterization of in vitro gingival models for future research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
For several years, two-dimensional (2D) cell cultures as an in-vitro tool, and animal models have been commonly used in periodontal research to study: disease patho-mechanisms, test new therapeutics and evaluate new regenerative strategies [1, 2]. 2D cell culture and animal models, however, are not free from limitations. For example, a 2D cell culture of gingival cells cannot fully replicate the architecture, physiological, and pathological microenvironment of living human gingival tissue, plus ethical and financial concerns are associated with animal experiments [3].
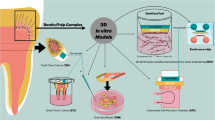
Three-dimensional (3D) gingival models provide researchers with an alternative to animal experimentation and 2D cell culture. Studies have reported the construction of 3D gingival models since 1997 with modified cell sources, scaffolds, and culture media. Initially, partial thickness models were constructed including epithelial tissue in absence of underlying connective tissue or connective tissue including gingival fibroblast cells without epithelial components [4, 5]. To date, full thickness 3D gingival models using human gingival-derived cellular sources including keratinocytes to assemble the epithelial layer and human gingival fibroblasts to establish the connective tissue layer are available. The advantage of full thickness 3D gingival model is their closer recapitulation to the complex structures and functions of native human gingival tissue [6, 7]. Several studies have demonstrated the application of these models in periodontal research. For instance, Dabija-Wolter et al. demonstrated the using of 3D gingival model to study host-microbial interaction. In this study, they examined the extent destruction of epithelial layer due to invation of F. nucleatum. They concluded invation of this pathogenic bacteria will trigger elimination of bacterial infection through epithelial shredding without causing a permanent damage of the tissue in 3D gingival model [8]. Razali et al. used 3D peri-implant model to understand the effect of photofunctionalization on three different types of implant abutment materials (yttriastabilized zirconia, alumina-toughened zirconia, and grade 2 commercially pure titanium). They and concluded that photofunctionalization of implant abutment materials improved the biological seal of the surrounding soft tissue peri-implant interface [9].
Although growing evidence have shown the promising outcome of 3D gingival model in periodontal research, there’s no consensus on fabrication method and material neither ideal characteristics for 3D gingival model. Studies have suggested that to recapitulate native gingival tissue, 3D model should be consisted of epithelial and connective tissue layers, which were separated by well define basement membrane. In addition, differentiation markers of each cell component, and functional assessment of the layers are also crucial [10, 11]. However, a critical evaluation of all these different types of models is missing. Indeed, all these different types of gingival models have not been reviewed with regards to their representation of human gingival tissue. The aim of this study was therefore to appraise current available 3D in vitro gingival models constructed using organoid cell culture system and provide answers to the following questions:
-
1.
Are any of the current 3D gingival models better replicate the native human gingival tissue in terms of their structure, differentiation characteristics, and barrier function.
-
2.
What are the available substrates that are used to reconstruct 3D gingival models?
2 Materials and methods
2.1 Focused questions
In view of the lack of specific tools to define the specific research questions we adapted the PICOS tool to search systematically for available evidence.
(P)Participant: 3D cell culture gingival model that is constructed by seeding gingival fibroblasts cells in the substrate and co-cultured with oral epithelial cells.
(I/E) Type of intervention/Exposure: N/A.
(C) Comparison: native human gingival tissue.
(O) Outcomes:
-
1-Resemblance of native human gingival tissue (3D structural layers evaluated by
-
histological analysis)
-
2- Differentiation markers of each cell component.
-
3- Functional assessment of the layers
(S) Studies type: In vitro experiments.
2.2 Protocol registration and reporting format
A systematic review protocol was developed and registered with the Open Science Framework (OSF) database, hosted by the Center for Open Science(COS) (https://archive.org/details/osf-registrations-6mzw2-v1 - License: http://www.gnu.org/licenses/lgpl-3.0.txt). Further when possible the systematic review was conducting according to the PRISMA guidelines [12].
2.3 Search strategy
Five electronic databases: MEDLINE (OVID), EMBASE, Dentistry and Oral Science Source (EBSCOhost), Web of Science Core Collection and LILACS (Latin American & Carribbean Health Sciences Literature) were included and updated up to the 12th of September 2022.
Hand searching process was performed by 2 independent reviewers (ZA and MH) and in case of any dispute further discussion with a third reviewer occurred (FDA). Only studies in the English language were included.
2.4 Study selection
All articles retrieved were exported and de-duplicated using the Reference Management Software “EndNote X9.3.3 (Bld 13966)”.
2.4.1 Study eligibility assessment
Screening and assessment of study eligibility were performed by 2 reviewers independently (ZM & MH) according to the inclusion and exclusion criteria. Agreement between the 2 reviewers was determined by kappa statistics.
Inclusion/exclusion criteria
Inclusion Criteria:
-
Studies of 3D cell culture gingival models constructed with a substrate seeded by human gingival fibroblasts or human periodontal ligament cells and human gingival/oral epithelial cells
-
3D cell culture gingival model construct with scaffold base system
-
Including histological analysis
-
Published in the English language.
Exclusion Criteria:
-
3D cell culture gingival model which was constructed without substrate base system
-
Studies of 3D cell culture gingival model which constructed with a substrate that seeded by non-human sources of fibroblast or epithelial cells.
-
Studies of 3D cell culture gingival model which was constructed with a substrate that seeded by human gingival fibroblasts or human periodontal ligament cells without human gingival/oral epithelial cells.
-
Studies of 3D cell culture gingival model which was constructed with a substrate that seeded by human gingival/oral epithelial cells without human gingival fibroblasts or human periodontal ligament cells.
-
Animal studies.
-
Studies without clear histological analysis.
-
Abstracts without full papers.
2.4.2 Data extraction strategy
Piloting of data extraction was conducted before starting with the full search strategy, further as some articles had a different methodology to prepare 3D models other than human cell sources two reviewers (ZM & MH) performed pilot runs using a specially designed data extraction spreadsheet. Any disagreements were resolved by discussion and if this was not possible, arbitration with an experienced reviewer was considered (FDA). Main categories of data were extracted as listed below: Study Characteristics Data: “Study authors, Year of publication and title, Study design, Conclusions”, “Participant/ 3D cell culture gingival model with inclusion/exclusion criteria, Human gingival fibroblasts cells, Specific substrate for cells seeding, Human epithelial cells “.
2.5 Study bias protection assessment
Quality assessment of included trials undertaken independently and in duplicate by two reviewers (ZM & MH) as part of the data extraction process. There are no established criteria for evaluating in vitro studies. Two tools of risk of bias were used in this review. The first one was the modified ARRIVE guidelines (Supplemental Data 2) to assess the quality of each study [13]. A second tool ‘Systematic Review Centre for Laboratory Animal Experimentation (SYRCLE)’s risk of bias tool’ was also used to analyze data and adapted by ruling out the blind intervention section [14].
3 Results
3.1 Study selection
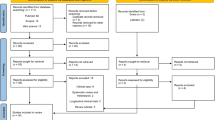
A total of 2338 articles were identified through database searching and Midline OVID n = 743; EMBASE n = 697; Web of Science n = 639; EBSCO n = 250; LILAC n = 9. The final number retrieved after completing the selection process was 22 (Fig. 1). Due to the absence of relevant quantitative measures to evaluate gingival models, quantitative models, and meta-analysis were not possible. Qualitative analyses of the evidence retrieved was conducted to summarize the characteristics of 3D gingival models.
3.2 Quality of studies
3.2.1 Modified ARRIVE guidelines (Supplemental Data 2)
Most of the selected studies were of high quality based on modified ARRIVE guidelines. Only seven studies discussed the scientific implications and limitations [9, 15,16,17,18,19,20].
Five studies did not give the statement of potential conflicts and funding disclosure [21,22,23,24,25] while one article was not published in a peer reviewed journal [26].
3.2.2 SYRCLE bias assessment
Well-balanced results in terms of low, unclear, and high risk of selection bias across studies were identified. All studies presented with high risk of bias in the random sequence generation and baseline variable characteristics. On analyzing allocation concealment, most selected articles had an unclear risk of bias, and only two articles had a low risk of bias [25, 27]. The randomization parameter was at high risk of bias. On analyzing random outcome assessment, all studies had an unclear risk of bias. In addition, all articles presented a low risk of bias in the results of incomplete outcome data, selective outcome reporting and other sources of bias (Tables 1 and 2).
3.3 3D Gingival model characteristics
Up to thirty-seven gingival and peri-implant models were described in the included twenty-two studies. Thirty-six models were constructed using the organotypic culture technique in a static cell culture condition. Only one study used a dynamic perfusion bioreactor system, where disc shape collagen sponge scaffolds were fitted in a perfusion bioreactor [22].
Regarding the cellular source, different types of cells were used including primary cells from gingival tissue biopsies or immortalized cell lines or a combination of both (Table 3).
Among these twenty-two studies, only six studies examined human gingival biopsy as a control [16, 25, 26, 28,29,30].
3.3.1 Macroscopical model appearance
In this review, one study by Koskinen Holm, C., & Qu, C. investigated macroscopical appearance of three gingival models constructed by using collagen type I (rat tail) that crosslinked with genipin, cytochalasin D, and genipin/ cytochalasin D, respectively [16]. Genipin is a chemical crosslinking agent, while cytochalasin D, is used to inhibit the rapid actin polymerization [31, 32]. This study showed that the crosslinked models using genipin or genipin/cytochalasin D were larger size in compared with non-crosslinked model.
3.4 Histological analysis
The included studies performed histological structure analysis to evaluate the successful construction of 3D model by using different types of staining techniques such as hematoxylin (H), hematoxylin and eosin (H&E), (H&E) and Periodic acid-Schiff (PAS), Masson’s trichrome, and van Gieson.
3.4.1 Epithelium layer
The number of epithelial cell layers was reported in nine studies with thirteen models and it ranged between 4 and 16 layers [8,9,10, 15, 17, 19, 23, 28, 33] (Table 4).
Dabija-Wolter et al. reported the number and thickness of epithelial layers. The thickness of epithelium at day 3 of development was 37.73 µm, and 49.79 µm, 130.93 µm, and 190.83 µm were at days 5, 7, and 9 respectively [28]. The study by Jennings et al. reported 120 µm thickness of well stratified epithelium. Chai et al. reported a pre-implant gingival model with a thickness of 50–100 µm [19] while Kriegebaum et al. demonstrated the formation of gingival model with an epithelium layer with 111.6 µm and 31 µm in thick when (TFE) and (DRT) were used respectively [23].
3.4.2 Connective tissue layer
With regards to the characteristic of connective tissue layer formation, eleven studies with nineteen models confirmed fibroblasts embedded in well-structured collagen fibrils [16, 17, 21, 23,24,25, 29, 30, 33,34,35]
Only one study reported the thickness of connective tissue layer, this study showed that by using TFE and DRT as substrates for gingival model construction the formation of connective tissue layers was 249.3 µm and 420.9 µm respectively [23].
3.5 Differentiation of gingival model
Thirty models from sixteen studies reported several expression markers to evaluate biological structures included in the constructed models. The reported markers were K4, K5, K10, K13, K14, K16, K17, K18, K19, involucrin, laminin5, proliferation marker Ki67, CD90, and vimentin, Type I, II and IV collagen [8, 16,17,18, 20, 21, 23,24,25,26, 28,29,30, 33,34,35].
3.5.1 Keratinocytes proliferation marker
The expression of keratinocytes proliferation marker Ki67 was investigated in eleven models [8, 16,17,18, 20, 26, 27, 33]. In addition to Ki67, one study analyzed the expression of PCNA as a marker for cell proliferation, which also confirmed the proliferation potential of keratinocytes in the model [20]. In contrast, apoptotic p53 marker was not detected in models prepared by using collagen type I hydrogel [16].
3.5.2 Keratinocytes differentiation markers
Cytokeratins
Cytokeratins (CKs) are the main intermediate filaments of gingival epithelia. Within the gingiva, the expression patterns of various CKs have been used as molecular indicators for different oral gingival epithelium regions [36, 37].
CK4 is predominantly found in the suprabasal compartment of non-keratinized epithelia including the buccal mucosa of the sulcular gingival epithelium. Tomakidi et al. analyzed the expression of CK4 in models constructed using primary non-keratinized gingival cells where the positive expression of CK4 in suprabasal layer was observed [24]. Roffel et al. reported a peri-implant gingival model, and the expression of CK4 was observed in the free gingival epithelia and sulcular epithelium but not in the junctional epithelium [17]. Sakulpaptong et al. reported the expressions of CK4 were observed in peri-implant gingival models prepared from human primary gingival cells. In addition, the expression of CK4 in the human native gingival tissue was also reported in this study [30].
CK13, a marker for non-stratified epithelial, was investigated in eight studies [8, 15, 24,25,26, 28, 33, 34]. Buskermolen et al. showed the expression pattern of CK13 in the gingival model, constructed with both primary and immortalized gingival keratinocytes, was similar to native gingiva. The gingival model established with KC-HPV showed a very low expression of CK13 [25]. However, the study by Jennings et al. reported that the abnormal expression of CK13 was observed in the gingival model using OSCC cells [26].
CK14, a basal cell specific marker, was evaluated in four studies. Tomakidi et al. showed the expression of CK14 was only limited to the basal layer [24]. In contrast, de Carvalho Diasa et al. and Koskinen Holm, C., & Qu, C., reported the expression of CK14 in both basal and suprabasal layer [16, 33]. Jennings et al. observed the expression CK14 throughout the entire epithelium [26]. And Bao et al. reported lower levels of CK14 expressions in gingival models in comparison to the human gingiva tissue [34].
CK5 is generally found in the basal cell compartment in all stratified epithelia. Two studies investigated the expression of CK5 [24, 30] and reported its expression limited to the basal cell compartment as revealed by gene expression study as well as immunolocalisation study. In a study by Sakulpaptong et al. [30], CK5 was expressed in peri-implant gingival models as well as in human native gingival tissue.
CK10 is known to be largely expressed in cornifying stratified and proliferating epithelia. Six studies analyzed the expression of CK10 in gingival models [8, 15, 16, 24, 25, 34]. Buskermolen et al. and Koskinen Holm, C., & Qu, C., showed the expression pattern of CK10 in the gingival models were similar to native human gingiva. However, the expression of CK10 was at a very low level in the model made with immortalized cell KC-HPV [16, 25].
Other cytokeratins such as CK8, CK16, CK18, CK19 and CK17 were investigated only in three studies [21, 28, 34].
The expression levels of CK18 and CK19 were similar between 3D and native human gingival tissue [34]. The expression of CK17 and CK19 was confirmed to be expressed in keratinocytes at multilayer in the 3D model by Ferra-Cancellas [21]. Dabija-Wolter et al. reported the expressions of CK 16 were observed in the suprabasal layer of the gingival model, and in both parabasal and suprabasal layers in native human gingival tissue. In the same study, the expression of CK19 and CK8 was observed in all cell layers l. However, both markers were expressed in few patterns in the basal layer of human native gingival tissue [28].
Other keratinocytes differentiation markers
Two studies showed the expression pattern of involucrin in the 3D gingival model was similar to native human gingival tissue [16, 25]. Other markers such as ODAM, FDC-SP, transglutaminase, and filaggrin were reported as junctional epithelial-specific markers [28].
3.5.3 E-cadherin (epithelial cadherin)
E-cadherin is a major protein involved in cell-to-cell adhesion. The expression of E-cadherin was reported in three models [8, 15, 26], confirming the tight epithelial barrier.
3.5.4 Basement membrane markers
Collagen IV and laminin are important proteins within the basement membrane. Six studies investigated and confirmed the expression of these two proteins in the basement membrane in the models [17, 23,24,25, 28, 30].
3.5.5 ECM components collagen type I and collagen type II
In this review the expression pattern of collagen type I (Col I) and collagen type II (Col II) was reported in two studies, and the levels of expression were found not significantly differed from native human gingival tissue [30, 34]. However, one study reported expression of both collagen 1, and CD90 by using qRT-PCR technique [16].
3.5.6 Vimentin
Vimentin is a differentiation marker for fibroblast. Buskermolen et al. and Koskinen Holm, C., & Qu, C., showed the expression of this marker in gingival model to be similar to native gingival tissue. Similarly, Ferrà -Cañellas et al. reported the expression of vimentin in the gingival model, which confirms the development of fibroblast in the gingival model [16, 21, 25].
3.6 Gingival model for periodontal research
With regards to the application of these gingival models, studies demonstrated the utilization of these models in several periodontal research applications as well as eight peri-implant models used in five studies were found (Fig. 2).
3.6.1 Host and microbial interaction study
In total, nine studies demonstrated the applicability of gingival models in host-microbial interaction studies. Within these nine studies, seven studies reported the response of gingival models to different bacterial challenges [8, 18, 20,21,22, 26, 27]. Four studies [8, 20, 21, 27] demonstrated the alteration of the epithelial layer upon the host-microbial interaction.
Apart from host-microbial interaction, the gingival model was used to investigate candida infection and it showed alteration of the structure by prominent degradation of the cornified layer of epithelial cells [10].
3.6.2 Mucosal model for dental implant research
The peri-implant mucosal models were used either for comparing different types of titanium and dental material posts surfaces [17, 19, 30], or for photofunctionalized effect on the biological seal of different types of abutment materials [9].
3.6.3 Gingival model for periodontal wound healing and regeneration
Potential application of 3D gingival models to study wound healing processes of the gingiva either following cold injury [27], micro-immunotherapy medicine (low dose of bone morphogenic protein (LD BMP4)) [21], or for the exposed model to sensitizers (Lin [18]).
3.7 Substrate biomaterials for construction of gingival model
In this review, 10 different substrate types were identified among all 37 models reported. The most used substrate was type I collagen sourced from rat tail, which was used in twenty-three models [8, 10, 17, 18, 20, 21, 24,25,26,27,28, 33,34,35]). Acellular human cadaveric dermis substrate (Alloderm) was used in three models [9, 19, 35], and decellularized dermis (purose dermis allograft) used in another model [30]. The other substrate including porcine collagen type I [22], porcine acellular dermal matrices (Strattice) [35], collagen/elastin matrix substrate (Matriderm) (bovine collagen type I with elastin) [29], dermal regeneration template (DRT) Single Layer substrate, Vicryl substrate, Tissu Foil E (TFE) [23], bovine type I collagen substrate were used to prepare four models [15, 30], (Table 4) and (Fig. 2).
4 Discussion
This review comprehensively described thirty-seven different 3D gingival and peri-implant models from twenty-two research studies. Twelve of these models confirmed good cell proliferation (marker Ki67) in both basal and suprabasal layers and most of the models confirmed good differentiation of epithelial cells (reporting different CKs markers). This was the first attempt to collectively appraise the available evidence resulting in not a single better model to study and test 3D gingival or peri-implant tissues.
Several studies have constructed gingival models from different cell origins, including primary cells, immortalized cell lines or a mixture of both. The highest number of epithelial layers was reported from the model using the primary cells origin [8]. In this review, two models were prepared from immortalized cell lines, H357, and OSCC, and demonstrated to be deficient in a well-defined differentiated epithelium [25, 26]. In contrast, one study reported that established Immortalized cell lines from primary human gingival cell induced by E6 and E7 oncoproteins of human papillomavirus, and resulted in a successful formation of gingival model with multi-layered epithelia [34]. These observations confirmed that these two types of immortalized human gingival cells (H357 and OSCC) are not suitable sources for gingival model construction. Further this review highlighted that when using cell lines in 3D gingival model construction, greater clarity in the presentation of the results is needed, this is because cell lines generally inherit the characteristics of their parental primary tissue cells hence when used these cells may not accurately reproduce properties or responses of normal epithelial cells [10, 34].
A crucial element in the construction of a gingival model is the substrate that provides scaffolding for the cells. The ideal substrate should have a high level of biocompatibility, porosity, biostability, and mechanical properties. In this review ten different substrates demonstrated to be applicable as matrices to mimic native gingival ECM and most of them were of animal origin. Rat tail collagen type I isolated from rat tail tendon was the most used and confirmed to allow the formation of the highest number of epithelial layers [8, 10, 28, 33]. The stratification of epithelial layers indicates the development of a gingival model, at the same time, a high level of stratification of keratinocytes has been demonstrated when there is an underlying homogenous distribution of fibroblasts among substrates. Rat tail collagen is considered the major type of collagen that is used as a substrate to mimic human ECM. Unfortunately, shrinkage is considered a disadvantage of models prepared by using collagen type I. This shrinkage can lead to a drastic decrease in the size of cell population in the hydrogel. However, it was reported that using genipin and genipin/cytochalasin D to crosslink collagen type I hydrogel allowed the construction of a model with more resistance to shrinkage facilitating in turn high cells survival and function [16]. Lastly, additional drawbacks for this collagen include its cost and its differences with human ECM´s collagen (where type I and III collagens are present as major constituents) plus isolated rat tail collagen is invariably fragmented [38]. All these drawbacks prevent considering rat tail collagen hydrogel to be ideal for gingival model construction.
Two more animal type of substrates were identified. A bovine collagen type I [15, 30] which demonstrated stratification and differentiation of epithelial layers with underlying connective tissue containing fibroblasts and a porcine substrate as a source of collagen type I to mimic human ECM as 3D collagen sponge scaffolds in a perfusion bioreactor system for easy manipulation [22]. However, these two substrates were not counted as a promising type for model construction due to lacking resemblance to native gingival human connective tissue.
In addition to collagen, dermal substrates were also widely used for tissue engineering and cell culture experiments. In this review, four dermal substrates were used for reconstructing gingival models including acellular cadaveric dermis and decellularized dermis (porous dermis allograft) as well as human [9, 19, 30, 35], porcine (strattice matrix) [35], and bovine (Matriderm) [29] sources. All of these dermal substrates showed good proliferation, differentiation and stratification of keratinocytes with a high distribution of fibroblasts. However, these types of substrates suffer from limited availability.
Lastly DRT was used as substrate for gingival model construction, a porous matrix of fibers of crosslinked bovine tendon collagen. High thickness tissue layers of gingival model with higher cells proliferation when compared to equine (TissuFoil E) and synthetic materials (Vicryl) substrate [23]. Electrospun type I crosslinked bovine collagen was used in one study to recreate a peri-implant gingival model [30] resulting in less tissue contraction and promising results. Size changes and contraction that occurred after model construction are attributed to the slow remodelling activity of the used substrates compared with native gingival tissue. This drawback is added to others mentioned above to take into account for proper selection of substrate to construct a developed gingival model.
It is worth mentioning that all the evidence on the use of different substrates collectively confirmed a high level of heterogeneity and the lack of a clear superior substrate to use for constructing the best 3D gingival model.
5 Limitations and future research
This review highlighted high heterogeneity, and lack of standardized fabrication and characterization protocols for the creation of a valid 3D gingival or peri-implant model. As such, a new framework for future characterization and construction of a 3D gingival model should be proposed that accounts for the uncertainty identified within this study.
The first step should include histological confirmation that the new model results in well-defined stratified epithelium layers with equal or more than four cell layers, and fibroblasts embedded and distributed homogenously in a well-structured substrate. Secondly well differentiated tissue layers should be confirmed via specific markers expression for each cell or layer regions, as following:
-
Ki67 for cell proliferation near basal epithelial layer
-
CK14 and CK5 for early differentiation in the basal layer and CK4 or CK13 in the suprabasal layer.
-
CK16, CK18, CK19 and CK17 in different epithelial layers as late differentiation markers
-
Involucrin as terminal differentiation marker for keratinocytes within the upper two third of the epithelium
-
CK10 marker to confirm the presence of cornifying stratified epithelia as well as in proliferating epithelia
-
Collagen IV and Laminin expression for the basement membrane
-
CD90 and Collagen (I and II) in ECM
-
Vimentin expression to confirm development of fibroblasts.
Thirdly an ideal 3D gingival model to use for different dental applications will need a well-developed vascular structure including capillary vessels, epithelial and stromal cells as well as immune, neural and bone cells (Fig. 3).
6 Conclusions
There is insufficient evidence to suggest whether the available 3D gingival models can entirely recapitulate the human gingival tissue and be valuable when performing experimental periodontal research. This review highlighted the lack of specific cell origin or substrate for constructing gingival models to reproduce physiologic properties of native human gingival tissue structures. Future research should aim at resolving the current challenges of construction a developed vascularized 3D gingival model mimic native human gingival tissue by engineering a new substrate with a high remodeling activity and suitable microenvironment for seeding human gingival cells.
Data availability
All data used to support the findings of this study are included within the article. Specifically, the registered systematic review protocol can be found at the Open Science Framework (OSF) database, hosted by the Center for Open Science(COS) (https://archive.org/details/osf-registrations-6mzw2-v1.
References
Sancilio S, di Giacomo V, Di Giulio M, Gallorini M, Marsich E, Travan A, et al. Biological responses of human gingival fibroblasts (HGFs) in an innovative co-culture model with Streptococcus mitis to thermosets coated with a silver polysaccharide antimicrobial system. Plos One. 2014;9:e96520.
Bates AM, Fischer CL, Abhyankar VP, Johnson GK, Guthmiller JM, Progulske-Fox A, et al. Matrix Metalloproteinase Response of Dendritic Cell, Gingival Epithelial Keratinocyte, and T-Cell Transwell Co-Cultures Treated with Porphyromonas gingivalis Hemagglutinin-B. Int J Mol Sci. 2018;19:07.
Su X, Fang D, Liu Y, Ramamoorthi M, Zeitouni A, Chen W, et al. Three‐dimensional organotypic culture of human salivary glands: the slice culture model. Oral Dis. 2016;22:639–48.
Morin M-P, Grenier D. Regulation of matrix metalloproteinase secretion by green tea catechins in a three-dimensional co-culture model of macrophages and gingival fibroblasts. Arch Oral Biol. 2017;75:89–99. https://doi.org/10.1016/j.archoralbio.2016.10.035
Basso FG, Pansani TN, Soares DG, Hebling J, de Souza Costa CA. LLLT Effects on Oral Keratinocytes in an Organotypic 3D Model. Photochem Photobiol. 2018;94:190–4.
Huh D, Hamilton GA, Ingber DE. From 3D cell culture to organs-on-chips. Trends Cell Biol. 2011;21:745–54.
Shamir ER, Ewald AJ. Three-dimensional organotypic culture: experimental models of mammalian biology and disease. Nat Rev Mol Cell Biol. 2014;15:647–64.
Dabija-Wolter G, Sapkota D, Cimpan MR, Neppelberg E, Bakken V, Costea DE. Limited in-depth invasion of Fusobacterium nucleatum into in vitro reconstructed human gingiva. Arch Oral Biol. 2012;57:344–51. https://doi.org/10.1016/j.archoralbio.2011.09.015
Razali M, Ngeow, WC, Omar, RA, & Chai, WL (2021). An In-Vitro Analysis of Peri-Implant Mucosal Seal Following Photofunctionalization of Zirconia Abutment Materials. Biomedicines. 9. https://doi.org/10.3390/biomedicines9010078
Dongari-Bagtzoglou A, Kashleva H. Development of a novel three-dimensional in vitro model of oral Candida infection. Microb Pathogenesis. 2006;40:271–8. https://doi.org/10.1016/j.micpath.2006.02.004
Klausner M, Handa Y, Aizawa S. In vitro three-dimensional organotypic culture models of the oral mucosa. Vitr Cell Dev Biol-Anim. 2021;57:148–59.
Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. J Clin Epidemiol. 2009;62:1006–12. https://doi.org/10.1016/j.jclinepi.2009.06.005
Ramamoorthi M, Bakkar M, Jordan J, Tran SD. Osteogenic potential of dental mesenchymal stem cells in preclinical studies: a systematic review using modified ARRIVE and CONSORT guidelines. Stem Cells Int. 2015;2015:378368. https://doi.org/10.1155/2015/378368
Hooijmans CR, Rovers MM, de Vries R, Leenaars M, Ritskes-Hoitinga M, Langendam MW. SYRCLE’s risk of bias tool for animal studies. BMC Med Res Methodol. 2014;14:1–9.
Ingendoh-Tsakmakidis A, Mikolai C, Winkel A, Szafranski SP, Flak CS, Rossi A, et al. Commensal and pathogenic biofilms differently modulate peri-implant oral mucosa in an organotypic model. Cell Microbiol. 2019;21. https://doi.org/10.1111/cmi.13078
Koskinen Holm C, Qu C. Engineering a 3D In Vitro Model of Human Gingival Tissue Equivalent with Genipin/Cytochalasin D. Int J Mol Sci. 2022;23:03.
Roffel S, Wu G, Nedeljkovic I, Meyer M, Razafiarison T, Gibbs S. Evaluation of a novel oral mucosa in vitro implantation model for analysis of molecular interactions with dental abutment surfaces. Clin Implant Dent Relat Res. 2019;21:25–33. https://doi.org/10.1111/cid.12750
Shang L, Deng D, Roffel S, Gibbs S. Differential influence of Streptococcus mitis on host response to metals in reconstructed human skin and oral mucosa. Contact Dermat. 2020;83:347–60.
Chai WL, Brook IM, Palmquist A, van Noort R, Moharamzadeh K. The biological seal of the implant-soft tissue interface evaluated in a tissue-engineered oral mucosal model. J R Soc Interface. 2012;9:3528–38.
Shang L, Deng DM, Buskermolen JK, Janus MM, Krom BP, Roffel S, et al. Multi-species oral biofilm promotes reconstructed human gingiva epithelial barrier function. Sci Rep. 2018;8. https://doi.org/10.1038/s41598-018-34390-y
Ferrà‐Cañellas MDM, Munar‐Bestard M, Garcia‐Sureda L, Lejeune B, Ramis JM, Monjo M, et al. BMP4 micro-immunotherapy increases collagen deposition and reduces PGE2 release in human gingival fibroblasts and increases tissue viability of engineered 3D gingiva under inflammatory conditions. J Periodontol. 2021;92:1448–59. https://doi.org/10.1002/JPER.20-0552
Bao K, Papadimitropoulos A, Akgul B, Belibasakis GN, Bostanci N. Establishment of an oral infection model resembling the periodontal pocket in a perfusion bioreactor system. Virulence. 2015;6:265–73.
Kriegebaum U, Mildenberger M, Mueller-Richter UD, Klammert U, Kuebler AC, Reuther T. Tissue engineering of human oral mucosa on different scaffolds: in vitro experiments as a basis for clinical applications. Oral Surg, Oral Med, Oral Pathol Oral Radiol. 2012;114:S190–198.
Tomakidi P, Fusenig NE, Kohl A, Komposch G. Histomorphological and biochemical differentiation capacity in organotypic co-cultures of primary gingival cells. J Periodontal Res. 1997;32:388–400.
Buskermolen JK, Reijnders CM, Spiekstra SW, Steinberg T, Kleverlaan CJ, Feilzer AJ, et al. Development of a Full-Thickness Human Gingiva Equivalent Constructed from Immortalized Keratinocytes and Fibroblasts. Tissue Eng - Part C: Methods. 2016;22:781–91.
Jennings LR, Colley HE, Ong J, Panagakos F, Masters JG, Trivedi HM, et al. Development and characterization of in vitro human oral mucosal equivalents derived from immortalized oral keratinocytes. Tissue Eng Part C: Methods. 2016;22:1108–17.
Buskermolen JK, Janus MM, Roffel S, Krom BP, Gibbs S. Saliva-Derived Commensal and Pathogenic Biofilms in a Human Gingiva Model. J Dent Res. 2018;97:201–8.
Dabija-Wolter G, Bakken V, Cimpan MR, Johannessen AC, Costea DE. In vitro reconstruction of human junctional and sulcular epithelium. J Oral Pathol Med. 2013;42:396–404.
Golinski PA, Gröger S, Herrmann JM, Bernd A, Meyle J. Oral mucosa model based on a collagen-elastin matrix. J Periodontal Res. 2011;46:704–11. https://doi.org/10.1111/j.1600-0765.2011.01393.x
Sakulpaptong W, Clairmonte IA, Blackstone BN, Leblebicioglu B, Powell HM. 3D engineered human gingiva fabricated with electrospun collagen scaffolds provides a platform for in vitro analysis of gingival seal to abutment materials. Plos One. 2022;17:e0263083.
Hwang YJ, Larsen J, Krasieva TB, Lyubovitsky JG. Effect of genipin crosslinking on the optical spectral properties and structures of collagen hydrogels. ACS Appl Mater Interfaces. 2011;3:2579–84. https://doi.org/10.1021/am200416h
Casella JF, Flanagan MD, Lin S. Cytochalasin D inhibits actin polymerization and induces depolymerization of actin filaments formed during platelet shape change. Nature. 1981;293:302–5. https://doi.org/10.1038/293302a0
de Carvalho Dias K, de Sousa DL, Barbugli PA, Cerri PS, Salih VM, Vergani CE. Development and characterization of a 3D oral mucosa model as a tool for host-pathogen interactions. J Microbiol Methods. 2018;152:52–60.
Bao K, Akguel B, Bostanci N. Establishment and characterization of immortalized gingival epithelial and fibroblastic cell lines for the development of organotypic cultures. Cells Tissues Organs. 2014;199:228–37.
Basso FG, Pansani TN, Marcelo CL, de Souza Costa CA, Hebling J, Feinberg SE. Phenotypic markers of oral keratinocytes seeded on two distinct 3D oral mucosa models. Toxicol Vitr. 2018;51:34–9.
Li S, Ge S, Yang P. Expression of cytokeratins in enamel organ, junctional epithelium and epithelial cell rests of Malassez. J Periodontal Res. 2015;50:846–54.
Newman MG Takei HH Klokkevold PR Carranza FA. Newman and Carranza’s Clinical Periodontology. Thirteenth ed. Philadelphia PA: Elsevier; 2018.
Ng Y-Z, South AP. Tissue engineering of tumor stromal microenvironment with application to cancer cell invasion. JoVE (J Visualized Exp). 2014;85:e51321.
Acknowledgements
Figure 3 was created with BioRender.com.
Funding
The authors are grateful for the support of Republic of Iraq/Ministry of Higher Education and Scientific Research, (Grant/Award Number: 177365) who provided a grant to promote this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
AlFatlawi, Z., Huang, M., Chau, D. et al. Three dimensional (3D) gingival models in periodontal research: a systematic review. J Mater Sci: Mater Med 34, 58 (2023). https://doi.org/10.1007/s10856-023-06761-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10856-023-06761-z