Abstract
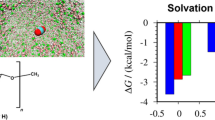
Cellulose I crystals swell on exposure to ethylenediamine (EDA) molecules to form a cellulose I–EDA complex, and successive extraction of EDA molecules converts the complex crystalline phase to either original cellulose I or cellulose IIII, depending on the treatment procedure. The present study reports the extended ensemble molecular dynamics (MD) simulation of the cellulose I–EDA complex models. An accelerated MD simulation allows most of the EDA molecules to desorb from the crystal model through a hydrophilic channel between the piles of cellulose chains, one at a time. Migration of a single EDA molecule along the channel is simulated by the adopted steered MD method combined with the umbrella sampling method to evaluate the potential of mean force (PMF) or free energy change on its movement. The PMF continues to increase during the migration of an EDA molecule to give a final PMF value of more than 30 kcal/mol. The PMF profiles are largely lowered by the removal of EDA molecules in the neighboring channels and by the widening of the channel. The former suggests that the EDA desorption cooperates with that in the neighboring channels, and, in the latter case, an EDA migration is efficiently promoted by solvation with water molecules in the expanded channel. We conclude that the atomistic picture of the EDA desorption behaviors observed in the crystal models is applicable to the real crystalline phase.
Graphic abstract





Similar content being viewed by others
References
Beckham GT, Crowley MF (2011) Examination of the α-chitin structure and decrystallization thermodynamics at the nanoscale. J Phys Chem B 115:4516–4522. https://doi.org/10.1021/jp200912q
Beckham GT, Matthews JF, Peters B, Bomble YJ, Himmel ME, Crowley MF (2011) Molecular-level origins of biomass recalcitrance: decrystallization free energies for four common cellulose polymorphs. J Phys Chem B 115:4118–4127. https://doi.org/10.1021/jp1106394
Bellesia G, Chundawat SP, Langan P, Dale BE, Gnanakaran S (2011) Probing the early events associated with liquid ammonia pretreatment of native crystalline cellulose. J Phys Chem B 115:9782–9788. https://doi.org/10.1021/jp2048844
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690. https://doi.org/10.1063/1.448118
Case DA, Ben-Shalom IY, Brozell SR, Cerutti DS, Cheatham III TE, Cruzeiro VWD, Darden TA, Duke RE, Ghoreishi D, Gilson MK, Gohlke H, Goetz AW, Greene D, Harris R, Homeyer N, Huang Y, Izadi S, Kovalenko A, Kurtzman T, Lee TS, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Mermelstein DJ, Merz KM, Miao Y, Monard G, Nguyen C, Nguyen H, Omelyan I, Onufriev A, Pan F, Qi R, Roe DR, Roitberg A, Sagui C, Schott-Verdugo S, Shen J, Simmerling CL, Smith J, Salomon-Ferrer R, Swails J, Walker RC, Wang J, Wei H, Wolf RM, Wu X, **ao L, York DM, Kollman PA (2018) Amber 18, https://ambermd.org: University of California, San Francisco, CA
Chanzy H, Henrissat B, Vincendon M, Tanner SF, Belton PS (1987) Solid-state 13C-N.M.R. and electron microscopy study on the reversible cellulose I→cellulose IIII transformation in Valonia. Carbohydr Res 160:1–11. https://doi.org/10.1016/0008-6215(87)80299-9
Chundawat SP, Bellesia G, Uppugundla N, da Costa SL, Gao D, Cheh AM, Agarwal UP, Bianchetti CM, Phillips GN Jr, Langan P, Balan V, Gnanakaran S, Dale BE (2011) Restructuring the crystalline cellulose hydrogen bond network enhances its depolymerization rate. J Am Chem Soc 133:11163–11174. https://doi.org/10.1021/ja2011115
Clark GL, Parker EA (1937) An X-ray diffraction study of the action of liquid ammonia on cellulose and its derivatives. J Phys Chem 41:777–786. https://doi.org/10.1021/j150384a001
Davis WE, Barry AJ, Peterson FC, King AJ (1943) X-ray studies of reactions of cellulose in non-aqueous systems. II. Interaction of cellulose and primary amines. J Am Chem Soc 65:1294–1299. https://doi.org/10.1021/ja01247a012
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593. https://doi.org/10.1063/1.470117
Evans DJ (1983) Computer “experiment” for nonlinear thermodynamics of Couette flow. J Chem Phys 78:3297–3302. https://doi.org/10.1063/1.445195
French AD, Dowd MK (1993) Exploration of disaccharide conformations by molecular mechanics. J Mol Struct (theochem) 286:183–201. https://doi.org/10.1016/0166-1280(93)87162-7
French AD, Miller DP, Aabloo A (1993) Miniature crystal models of cellulose polymorphs and other carbohydrates. Int J Biol Macromol 15:30–36. https://doi.org/10.1016/s0141-8130(05)80085-6
Grossfield A (2018) WHAM: an implementation of the weighted histogram analysis method, ver. 2.0.10, http://membrane.urmc.rochester.edu/content/wham: University of Rochester Medical Center: Rochester, NY
Hamelberg D, de Oliveira CA, McCammon JA (2007) Sampling of slow diffusive conformational transitions with accelerated molecular dynamics. J Chem Phys 127:155102. https://doi.org/10.1063/1.2789432
Hamelberg D, Mongan J, McCammon JA (2004) Accelerated molecular dynamics: a promising and efficient simulation method for biomolecules. J Chem Phys 120:11919–11929. https://doi.org/10.1063/1.1755656
Hoover WG, Ladd AJC, Moran B (1982) High-strain-rate plastic flow studied via nonequilibrium molecular dynamics. Phys Rev Lett 48:1818–1820. https://doi.org/10.1103/PhysRevLett.48.1818
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Igarashi K, Uchihashi T, Koivula A, Wada M, Kimura S, Okamoto T, Penttila M, Ando T, Samejima M (2011) Traffic jams reduce hydrolytic efficiency of cellulase on cellulose surface. Science 333:1279–1282. https://doi.org/10.1126/science.1208386
Igarashi K, Wada M, Samejima M (2007) Activation of crystalline cellulose to cellulose IIII results in efficient hydrolysis by cellobiohydrolase. FEBS J 274:1785–1792. https://doi.org/10.1111/j.1742-4658.2007.05727.x
Jarzynski C (1997) Nonequilibrium equality for free energy differences. Phys Rev Lett 78:2690–2693. https://doi.org/10.1103/PhysRevLett.78.2690
Jorgensen WL (1981) Quantum and statistical mechanical studies of liquids. 10. Transferable intermolecular potential functions for water, alcohols, and ethers. Application to liquid water. J Am Chem Soc 103:335–340. https://doi.org/10.1021/ja00392a016
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935. https://doi.org/10.1063/1.445869
Kirschner KN, Yongye AB, Tschampel SM, Gonzalez-Outeirino J, Daniels CR, Foley BL, Woods RJ (2008) GLYCAM06: a generalizable biomolecular force field. Carbohydrates J Comput Chem 29:622–655. https://doi.org/10.1002/jcc.20820
Kumar S, Rosenberg JM, Bouzida D, Swendsen RH, Kollman PA (1992) THE weighted histogram analysis method for free-energy calculations on biomolecules. I the Method J Comput Chem 13:1011–1021. https://doi.org/10.1002/jcc.540130812
Kumar S, Rosenberg JM, Bouzida D, Swendsen RH, Kollman PA (1995) Multidimensional free-energy calculations using the weighted histogram analysis method. J Comput Chem 16:1339–1350. https://doi.org/10.1002/jcc.540161104
Lau MW, Dale BE (2009) Cellulosic ethanol production from AFEX-treated corn stover using Saccharomyces cerevisiae 424A(LNH-ST). Proc Natl Acad Sci USA 106:1368–1373. https://doi.org/10.1073/pnas.0812364106
Le Grand S, Götz AW, Walker RC (2013) SPFP: Speed without compromise—A mixed precision model for GPU accelerated molecular dynamics simulations. Comput Phys Commun 184:374–380. https://doi.org/10.1016/j.cpc.2012.09.022
Loeb L, Segal L (1955) Studies of the ethylenediamine-cellulose complex. I. Decomposition of the complex by solvents. J Polym Sci 15:343–354. https://doi.org/10.1002/pol.1955.120158003
Lokhande HT, Shukla SR, Chidambareswaran PK, Patil NB (1977) Ethylenediamine-induced conversion of cellulose I to cellulose III. J Polymer Sci Polymer Lett Edition 15:97–99. https://doi.org/10.1002/pol.1977.130150207
Matthews JF, Skopec CE, Mason PE, Zuccato P, Torget RW, Sugiyama J, Himmel ME, Brady JW (2006) Computer simulation studies of microcrystalline cellulose Iβ. Carbohydr Res 341:138–152. https://doi.org/10.1016/j.carres.2005.09.028
Miao Y, Sinko W, Pierce L, Bucher D, Walker RC, McCammon JA (2014) Improved reweighting of accelerated molecular dynamics simulations for free energy calculation. J Chem Theory Comput 10:2677–2689. https://doi.org/10.1021/ct500090q
Nishiyama Y, Langan P, Chanzy H (2002) Crystal structure and hydrogen-bonding system in cellulose Iβ from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 124:9074–9082. https://doi.org/10.1021/ja0257319
Nishiyama Y, Wada M, Hanson BL, Langan P (2010) Time-resolved X-ray diffraction microprobe studies of the conversion of cellulose I to ethylenediamine-cellulose I. Cellulose 17:735–745. https://doi.org/10.1007/s10570-010-9415-9
Ozer G, Quirk S, Hernandez R (2012) Adaptive steered molecular dynamics: validation of the selection criterion and benchmarking energetics in vacuum. J Chem Phys 136:215104. https://doi.org/10.1063/1.4725183
Ozer G, Valeev EF, Quirk S, Hernandez R (2010) Adaptive steered molecular dynamics of the long-distance unfolding of neuropeptide Y. J Chem Theory Comput 6:3026–3038. https://doi.org/10.1021/ct100320g
Park S, Khalili-Araghi F, Tajkhorshid E, Schulten K (2003) Free energy calculation from steered molecular dynamics simulations using Jarzynski’s equality. J Chem Phys 119:3559–3566. https://doi.org/10.1063/1.1590311
Park S, Schulten K (2004) Calculating potentials of mean force from steered molecular dynamics simulations. J Chem Phys 120:5946–5961. https://doi.org/10.1063/1.1651473
Pentoney RE (1966) Liquid ammonia-solvent combinations in wood plasticization. Properties of treated wood. Ind Eng Chem Prod Res Dev 5:105–110. https://doi.org/10.1021/i360018a003
Perez DDS, Montanari S, Vignon MR (2003) TEMPO-mediated oxidation of cellulose III. Biomacromol 4:1417–1425. https://doi.org/10.1021/bm034144s
Pierce LCT, Salomon-Ferrer R, de Oliveira CAF, McCammon JA, Walker RC (2012) Routine access to millisecond time scale events with accelerated molecular dynamics. J Chem Theory Comput 8:2997–3002. https://doi.org/10.1021/ct300284c
Roche E, Chanzy H (1981) Electron microscopy study of the transformation of cellulose I into cellulose IIII in Valonia. Int J Biol Macromol 3:201–206. https://doi.org/10.1016/0141-8130(81)90064-7
Roe DR, Cheatham TE III (2013) PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9:3084–3095. https://doi.org/10.1021/ct400341p
Roux B (1995) The calculation of the potential of mean force using computer simulations. Comput Phys Commun 91:275–282. https://doi.org/10.1016/0010-4655(95)00053-i
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J Comput Phys 23:327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Salminen R, Baccile N, Reza M, Kontturi E (2017) Surface-induced frustration in solid state polymorphic transition of native cellulose nanocrystals. Biomacromol 18:1975–1982. https://doi.org/10.1021/acs.biomac.7b00463
Salomon-Ferrer R, Gotz AW, Poole D, Le Grand S, Walker RC (2013a) Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh ewald. J Chem Theory Comput 9:3878–3888. https://doi.org/10.1021/ct400314y
Salomon-Ferrer R, Pierce L, Walker R (2013b) Accelerated molecular dynamics in AMBER example analysis of all-atom enhanced sampling method accelerated molecular dynamics (aMD) to investigate conformational changes in proteins that typically occur on the millisecond time scale. https://ambermd.org/tutorials/advanced/tutorial22. Accessed June 1, 2021
Sawada D, Nishiyama Y, Petridis L, Parthasarathi R, Gnanakaran S, Forsyth VT, Wada M, Langan P (2013) Structure and dynamics of a complex of cellulose with EDA: insights into the action of amines on cellulose. Cellulose 20:1563–1571. https://doi.org/10.1007/s10570-013-9974-7
Schrödinger LLC. (2021) The PyMOL molecular graphics system, ver. 2.5.0, https://pymol.org: New York, NY
Sugiyama J, Harada H, Saiki H (1987) Crystalline morphology of Valonia macrophysa cellulose IIII revealed by direct lattice imaging. Int J Biol Macromol 9:122–130. https://doi.org/10.1016/0141-8130(87)90039-0
Torrie GM, Valleau JP (1974) Monte Carlo free energy estimates using non-Boltzmann sampling: Application to the sub-critical Lennard-Jones fluid. Chem Phys Lett 28:578–581. https://doi.org/10.1016/0009-2614(74)80109-0
Torrie GM, Valleau JP (1977) Nonphysical sampling distributions in Monte Carlo free-energy estimation: umbrella sampling. J Comput Phys 23:187–199. https://doi.org/10.1016/0021-9991(77)90121-8
Uto T, Hosoya T, Hayashi S, Yui T (2013) Partial crystalline transformation of solvated cellulose IIII crystals, reproduced by theoretical calculations. Cellulose 20:605–612. https://doi.org/10.1007/s10570-013-9861-2
Uto T, Mawatari S, Yui T (2014) Theoretical study of the structural stability of molecular chain sheet models of cellulose crystal allomorphs. J Phys Chem B118: 9313–9321
Uto T, Minamizaki M, Yui T (2019) Molecular dynamics simulation of cellulose I-ethylenediamine complex crystal models. J Phys Chem B 124:134–143. https://doi.org/10.1021/acs.jpcb.9b08153
Uto T, Yui T (2018) DFT optimization of isolated molecular chain sheet models constituting native cellulose crystal structures. ACS Omega 3:8050–8058. https://doi.org/10.1021/acsomega.8b00834
Wada M (2001) In situ observation of the crystalline transformation from cellulose IIII to Iβ. Macromolecules 34:3271–3275. https://doi.org/10.1021/ma0013354
Wada M, Chanzy H, Nishiyama Y, Langan P (2004) Cellulose IIII crystal structure and hydrogen bonding by synchrotron X-ray and neutron fiber diffraction. Macromolecules 37:8548–8555. https://doi.org/10.1021/ma0485585
Wada M, Heux L, Isogai A, Nishiyama Y, Chanzy H, Sugiyama J (2001) Improved structural data of cellulose IIII prepared in supercritical ammonia. Macromolecules 34:1237–1243. https://doi.org/10.1021/ma001406z
Wada M, Heux L, Nishiyama Y, Langan P (2009a) The structure of the complex of cellulose I with ethylenediamine by X-ray crystallography and cross-polarization/magic angle spinning 13C nuclear magnetic resonance. Cellulose 16:943–957. https://doi.org/10.1007/s10570-009-9338-5
Wada M, Heux L, Nishiyama Y, Langan P (2009b) X-ray crystallographic, scanning microprobe X-ray diffraction, and cross-polarized/magic angle spinning 13C NMR studies of the structure of cellulose IIIII. Biomacromol 10:302–309. https://doi.org/10.1021/bm8010227
Wada M, Kwon GJ, Nishiyama Y (2008) Structure and thermal behavior of a cellulose I-ethylenediamine complex. Biomacromol 9:2898–2904. https://doi.org/10.1021/bm8006709
Wada M, Nishiyama Y, Bellesia G, Forsyth T, Gnanakaran S, Langan P (2011) Neutron crystallographic and molecular dynamics studies of the structure of ammonia-cellulose I: rearrangement of hydrogen bonding during the treatment of cellulose with ammonia. Cellulose 18:191–206. https://doi.org/10.1007/s10570-010-9488-5
Wada M, Nishiyama Y, Langan P (2006) X-ray structure of ammonia−cellulose I: new insights into the conversion of cellulose I to cellulose IIII. Macromolecules 39:2947–2952. https://doi.org/10.1021/ma060228s
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general Amber force field. J Comput Chem 25:1157–1174. https://doi.org/10.1002/jcc.20035
Yui T, Hayashi S (2008) Structural stability of the solvated cellulose IIII crystal models: a molecular dynamics study. Cellulose 16:151–165. https://doi.org/10.1007/s10570-008-9265-x
Yui T, Nishimura S, Akiba S, Hayashi S (2006) Swelling behavior of the cellulose Iβ crystal models by molecular dynamics. Carbohydr Res 341:2521–2530. https://doi.org/10.1016/j.carres.2006.04.051
Yui T, Okayama N, Hayashi S (2010) Structure conversions of cellulose IIII crystal models in solution state: a molecular dynamics study. Cellulose 17:679–691. https://doi.org/10.1007/s10570-010-9422-x
Yui T, Taki N, Sugiyama J, Hayashi S (2007) Exhaustive crystal structure search and crystal modeling of β-chitin. Int J Biol Macromol 40:336–344. https://doi.org/10.1016/j.ijbiomac.2006.08.017
Zheng Y, Pan Z, Zhang R (2009) Overview of biomass pretreatment for cellulosic ethanol production. Int J Agric & Biol Eng 2:51–68. https://doi.org/10.3965/j.issn.1934-6344.2009.03.051-068
Acknowledgments
MD calculations were partly performed by using the computer cluster systems at Research Center for Computational Science (RCCS), Okazaki Research Facilities, and National Institutes of Natural Sciences (NINS), Japan. We thank Edanz (https://www.jp.edanz.com/ac) for editing a draft of this manuscript.
Funding
The authors did not receive support from any organization for the submitted work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The authors have no statement about ethical approval as this research does not involve humans and animals.
Informed consent
The authors have no statements about informed consent as this research does not involve human participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary file2 (MP4 280855 kb)
Supplementary file3 (MP4 118615 kb)
Rights and permissions
About this article
Cite this article
Yui, T., Uto, T. Extended ensemble molecular dynamics study of cellulose I–ethylenediamine complex crystal models: atomistic picture of desorption behaviors of ethylenediamine. Cellulose 29, 2855–2867 (2022). https://doi.org/10.1007/s10570-021-04136-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-021-04136-7