Abstract
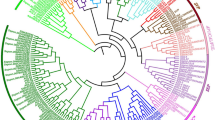
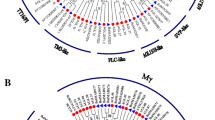
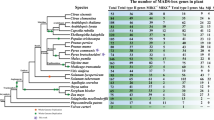
MADS-box genes encode a family of transcription factors that regulate diverse growth and developmental processes in plants, including flowering. In this study, comprehensive characterization and expression profiling analyses of seven sugar apple (Annona squamosa L.) MADS-box genes were performed using rapid amplification of cDNA ends method. Domain and phylogenetic analyses grouped these seven MADS-box genes into six different clades and they showed high similarity with orthologs in Arabidopsis. Expression patterns of these MADS-box genes were investigated during different flower developmental stages and in various reproductive organs, including petal, stamen, sepal, and pistil. Most of the MADS-box genes studied were least expressed in the sepal and AsAGL67 and AsAGL80 expression was weak in all tissues. AsSEP1 and AsAGAMOUS showed highest expressions in the stamen and pistil, and AsAGL12 showed stamen-specific expression. Dynamic expression patterns of MADS-box genes in different reproductive stages suggest involvement in flower development. Interestingly, a number of these MADS-box genes showed responses to gibberellin, abscisic acid, and salicylic acid treatments, suggesting control of their expression by phytohormones.
Similar content being viewed by others
Abbreviations
- ABA:
-
abscisic acid
- AG:
-
AGAMOUS
- CaMV:
-
cauliflower mosaic virus
- GA:
-
gibberellin
- GFP:
-
green fluorescent protein
- NJ:
-
neighbour-joining
- RACE:
-
rapid amplification of cDNA ends
- SA:
-
salicylic acid
References
Almeida, A.M., Yockteng, R., Otoni, W.C., Specht, C.D.: Positive selection on the K domain of the AGAMOUS protein in the Zingiberales suggests a mechanism for the evolution of androecial morphology. - EvoDevo 6: 7, 2015.
Alvarez-Buylla, E.R., Liljegren, S.J., Pelaz, S., Gold, S.E., Burgeff, C., Ditta, G.S., Vergara-Silva, F., Yanofsky, M.F.: MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes. - Plant J. 24: 457–466, 2000.
Bezerra, I.C., Michaels, S.D., Schomburg, F.M., Amasino, R.M.: Lesions in the mRNA cap-binding gene ABA HYPERSENSITIVE 1 suppress FRIGIDA-mediated delayed flowering in Arabidopsis. - Plant J. 40: 112–119, 2004.
Causier, B., Schwarz-Sommer, Z., Davies, B.: Floral organ identity: 20 years of ABCs. - Semin. Cell. Dev. Biol. 21: 73–79, 2010.
Cho, S., Jang, S., Chae, S., Chung, K.M., Moon, Y.H., An, G., Jang, S.K.: Analysis of the C-terminal region of Arabidopsis thaliana APETALA1 as a transcription activation domain. - Plant mol. Biol. 40: 419–429, 1999.
Cui, Z., Zhou, B., Zhang, Z., Hu, Z.: Abscisic acid promotes flowering and enhances LcAP1 expression in Litchi chinensis Sonn. - S. Afr. J. Bot. 88: 76–79, 2013.
De Bodt, S., Raes, J., Van de Peer, Y., Theißen, G.: And then there were many: MADS goes genomic. - Trends Plant Sci. 8: 475–483, 2003.
Diaz-Riquelme, J., Lijavetzky, D., Martinez-Zapater, J.M., Carmona, M.J.: Genome-wide analysis of MIKCC-type MADS box genes in grapevine. - Plant Physiol. 149: 354–369, 2009.
Dornelas, M.C., Patreze, C.M., Angenent, G.C., Immink, R.G.H.: MADS: the missing link between identity and growth? - Trends Plant Sci. 16: 89–97, 2010.
Duan, W., Song, X., Liu, T., Huang, Z., Ren, J., Hou, X., Li, Y.: Genome-wide analysis of the MADS-box gene family in Brassica rapa (Chinese cabbage). - Mol. Genet. Genomics 290: 239–255, 2015.
Eisen, M.B., Spellman, P.T., Brown, P.O., Botstein, D.: Cluster analysis and display of genome-wide expression patterns. - Proc. nat. Acad. Sci. USA 95: 14863–14868, 1998.
Fernandez, D.E., Heck, G.R., Perry, S.E., Patterson, S.E., Bleecker, A.B., Fang, S.C.: The embryo MADS domain factor AGL15 acts postembryonically. Inhibition of perianth senescence and abscission via constitutive expression. - Plant Cel. 12: 183–198, 2000.
Groth, E., Tandre, K., Engstrom, P., Vergara-Silva, F.: AGAMOUS subfamily MADS-box genes and the evolution of seed cone morphology in Cupressaceae and Taxodiaceae. - Evol. Dev. 13: 159–170, 2011.
Huang, F., Chi, Y., Gai, J., Yu, D.: Identification of transcription factors predominantly expressed in soybean flowers and characterization of GmSEP1 encoding a SEPALLATA1-like protein. - Gene 438: 40–48, 2009.
Kaufmann, K., Melzer, R., Theiß en, G.: MIKC-type MADSdomain proteins: structural modularity, protein interactions and network evolution in land plants. - Gene 347: 183–198, 2005.
Kim, S., Koh, J., Yoo, M.J., Kong, H., Hu, Y., Ma, H., Soltis, P.S., Soltis, D.E.: Expression of floral MADS-box genes in basal angiosperms: implications for the evolution of floral regulators. - Plant J. 43: 724–744, 2005.
Kramer, E.M., Irish, V.F.: Evolution of genetic mechanisms controlling petal development. - Nature 399: 144–148, 1999.
Lee, S., Woo, Y.M., Ryu, S.I., Shin, Y.D., Kim, W.T., Park, K.Y., Lee, I.J., An, G.: Further characterization of a rice AGL12 group MADS-box gene, OsMADS26. - Plant physiol. 147: 156–168, 2008.
Li, H.Y., Liu, F.F., Liu, G.F., Wang, S., Guo, X.H., **g, J.: Molecular cloning and expression analysis of 13 MADSbox genes in Betula platyphylla. - Plant mol. Biol. Rep. 30: 149–157, 2012a.
Li, Z., Liu, G., Zhang, J., Lu, S., Yi, S., Bao, M.: Cloning and characterization of paleoAP3-like MADS-box gene in London plane tree. - Biol. Plant. 56: 585–589, 2012b.
Little, C.H., MacDonald, J.E.: Effects of exogenous gibberellin and auxin on shoot elongation and vegetative bud development in seedlings of Pinus sylvestris and Picea glauca. - Tree Physiol. 23: 73–83, 2003.
Liu, K.D., Li, H.L., Yuan, C.C., Huang, Y.L., Chen, Y., Liu, J.X.: Identification of phenological growth stages of sugar apple (Annona squamosa L.) using the extended BBCHscale. - Sci. Hort. 181: 76–80, 2015.
Lu, S.J., Wei, H., Wang, Y., Wang, H.M., Yang, R.F., Zhang, X.B., Tu, J.M.: Overexpression of a transcription factor OsMADS15 modifies plant architecture and flowering time in rice (Oryza sativa L.). - Plant mol. Biol. Rep. 30: 1461–1469, 2012.
Martínez, C., Pons, E., Prats, G., León, J.: Salicylic acid regulates flowering time and links defence responses and reproductive development. - Plant J. 37: 209–217, 2004.
Masiero, S., Colombo, L., Grini, P.E., Schnittger, A., Kater, M.M.: The emerging importance of type I MADS box transcription factors for plant reproduction. - Plant Cell 23: 865–872, 2011.
Moon, J., Suh, S.S., Lee, H., Choi, K.R., Hong, C.B., Paek, N.C., Kim, S.G., Lee, I.: The SOC1 MADS-box gene integrates vernalization and gibberellin signals for flowering in Arabidopsis. - Plant J. 35: 613–623, 2003.
Mutasa-Göttgens, E., Hedden, P.: Gibberellin as a factor in floral regulatory networks. - J. exp. Bot. 60: 1979–1989, 2009.
Ng, M., Yanofsky, M.F.: Function and evolution of the plant MADS-box gene family. - Nat. Rev. Genet. 2: 186–195, 2001.
Norman, C., Runswick, M., Pollock, R., Treisman, R.: Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. - Cell 55: 989–1003, 1988.
Passmore, S., Maine, G.T., Elble, R., Christ, C., Tye, B.-K.: Saccharomyces cerevisiae protein involved in plasmid maintenance is necessary for mating of MATa cells. - J. mol. Biol. 204: 593–606, 1988.
Patharkar, O.R., Walker, J.C.: Floral organ abscission is regulated by a positive feedback loop. - Proc. nat. Acad. Sci. USA 112: 2906–2911, 2015.
Pelaz, S., Ditta, G.S., Baumann, E., Wisman, E., Yanofsky, M.F.: B and C floral organ identity functions require SEPALLATA MADS-box genes. - Nature 405: 200–203, 2000.
Pelaz, S., Gustafson-Brown, C., Kohalmi, S.E., Crosby, W.L., Yanofsky, M.F.: APETALA1 and SEPALLATA3 interact to promote flower development. - Plant J. 26: 385–394, 2001.
Robles, P., Pelaz, S.: Flower and fruit development in Arabidopsis thaliana. - Int. J. dev. Biol. 49: 633–643, 2005.
Puig, J., Meynard, D., Khong, G.N., Pauluzzi, G., Guiderdoni, E., Gantet, P.: Analysis of the expression of the AGL17-like clade of MADS-box transcription factors in rice. - Gene Expression Patterns 13: 160–170, 2013.
Remay, A., Lalanne, D., Thouroude, T., Le Couviour, F., Hibrand-Saint Oyant, L., Foucher, F.: A survey of flowering genes reveals the role of gibberellins in floral control in rose. - Theor. appl. Genet. 119: 767–781, 2009.
Sablowski, R.: Flowering and determinacy in Arabidopsis. - J. exp. Bot. 58: 899–907, 2007.
Sakamoto, T., Kobayashi, M., Itoh, H., Tagiri, A., Kayano, T., Tanaka, H., Iwahori, S., Matsuoka, M.: Expression of a gibberellin 2-oxidase gene around the shoot apex is related to phase transitionin rice. - Plant Physiol. 125: 1508–1516, 2001.
Schwarz-Sommer, Z., Huijser, P., Nacken, W., Saedler, H., Sommer, H.: Genetic control of flower development by homeotic genes in Antirrhinum majus. - Science 250: 931–936, 1990.
Seymour, G.B., Ryder, C.D., Cevik, V., Hammond, J.P., Popovich, A., King, G.J., Vrebalov, J., Giovannoni, J.J., Manning, K.: A SEPALLATA gene is involved in the development and ripening of strawberry (Fragaria × ananassa Duch.) fruit, a non-climacteric tissue. - J. exp. Bot. 62: 1179–1188, 2011.
Shu, Y., Yu, D., Wang, D., Guo, D., Guo, C.: Genome-wide survey and expression analysis of the MADS-box gene family in soybean. - Mol. Biol. Rep. 40: 3901–3911, 2013.
Smaczniak, C., Immink, R.G., Angenent, G.C., Kaufmann, K.: Developmental and evolutionary diversity of plant MADSdomain factors: insights from recent studies. - Development 139: 3081–3098, 2012.
Sommer, H., Beltran, J.P., Huijser, P., Pape, H., Lonnig, W.E., Saedler, H., Schwarz-Sommer, Z.: Deficiens, a homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: the protein shows homology to transcription factors. - EMBO J. 9: 605–613, 1990.
Sun, B., Xu, Y., Ng, K.H., Ito, T.: A timing mechanism for stem cell maintenance and differentiation in the Arabidopsis floral meristem. - Genes Dev. 23: 1791–1804, 2009.
Tanaka, Y., Oshima, Y., Yamamura, T., Sugiyama, M., Mitsuda, N., Ohtsubo, N., Ohme-Takagi, M., Terakawa, T.: Multi-petal cyclamen flowers produced by AGAMOUS chimeric repressor expression. - Sci. Rep. 3: 2641, 2013.
Tapia-Lopez, R., Garcia-Ponce, B., Dubrovsky, J.G., Garay- Arroyo, A., Perez-Ruiz, R.V., Kim, S.H., Acevedo, F., Pelaz, S., Alvarez-Buylla, E.R.: An AGAMOUS-related MADS-box gene, XAL1 (AGL12), regulates root meristem cell proliferation and flowering transition in Arabidopsis. - Plant Physiol. 146: 1182–1192, 2008.
Theissen, G., Becker, A., Di Rosa, A., Kanno, A., Kim, J.T., Munster, T., Winter, K.U., Saedler, H.: A short history of MADS-box genes in plants. - Plant mol. Biol. 42: 115–149, 2000.
Theissen, G., Kim, J.T., Saedler, H.: Classification and phylogeny of the MADS-box multigene family suggest defined roles of MADS-box gene subfamilies in the morphological evolution of eukaryotes. - J. mol. Evol. 43: 484–516, 1996.
Theissen, G., Melzer, R.: Molecular mechanisms underlying origin and diversification of the angiosperm flower. - Ann. Bot. 100: 603–619, 2007.
Theissen, G., Saedler, H.: Plant biology. Floral quartets. - Nature 409: 469–471, 2001.
Weigel, D., Meyerowitz, E.M.: The ABCs of floral homeotic genes. - Cell 78: 203–209, 1994.
Wells, C.E., Vendramin, E., Jimenez Tarodo, S., Verde, I., Bielenberg, D.G.: A genome-wide analysis of MADS-box genes in peach [Prunus persica (L.) Batsch]. - BMC Plant Biol. 15: 41, 2015.
Wilmowicz, E., Kesy, J., Kopcewicz, J.: Ethylene and ABA interactions in the regulation of flower induction in Pharbitis nil. - J. Plant Physiol. 165: 1917–1928, 2008.
Xu, Z., Zhang, Q., Sun, L., Du, D., Cheng, T., Pan, H., Yang, W., Wang, J.: Genome-wide identification, characterisation and expression analysis of the MADS-box gene family in Prunus mume. - Mol. Genet. Genomics 289: 903–920, 2014.
Yamada, M., Takeno, K.: Stress and salicylic acid induce the expression of PnFT2 in the regulation of the stress-induced flowering of Pharbitis nil. - J. Plant Physiol. 171: 205–212, 2014.
Yang, Y., Fanning, L., Jack, T.: The K domain mediates heterodimerization of the Arabidopsis floral organ identity proteins, APETALA3 and PISTILLATA. - Plant J. 33: 47–59, 2003.
Yanofsky, M.F., Ma, H., Bowman, J.L., Drews, G.N., Feldmann, K.A., Meyerowitz, E.M.: The protein encoded by the Arabidopsis homeotic gene AGAMOUS resembles transcription factors. - Nature 346: 35–39, 1990.
Yu, S., Galvã o, V.C., Zhang, Y.C., Horrer, D., Zhang, T.Q., Hao, Y.H., Feng, Y.Q., Wang, S., Schmid, M., Wang, J.W.: Gibberellin regulates the Arabidopsis floral transition through miR156-targeted SQUAMOSA PROMOTER BINDING-LIKE transcription factors. - Plant Cell 24: 3320–3332, 2012.
Zahn, L.M., Kong, H., Leebens-Mack, J.H., Kim, S., Soltis, P.S., Landherr, L.L., Soltis, D.E., Depamphilis, C.W., Ma, H.: The evolution of the SEPALLATA subfamily of MADSbox genes: a preangiosperm origin with multiple duplications throughout angiosperm history. - Genetics 169: 2209–2223, 2005.
Zhang, Z., Li, H., Zhang, D., Liu, Y., Fu, J., Shi, Y., Song, Y., Wang, T., Li, Y.: Characterization and expression analysis of six MADS-box genes in maize (Zea mays L.). - J. Plant Physiol. 169: 797–806, 2012.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Acknowledgements: This work was supported by the National Natural Science Foundation of China (grant No. 31201586); the Science and Technology Program of Guangdong, China (grant Nos. 2013B020304008 and 2014A020208138); the Key Project of Department of Education of Guangdong Province (grant No. 2013KJCX0124); the Key Laboratory of Zhanjiang Tropical Characteristic Plant Resources Technology Development (grant No. 2014A06008). The first two authors contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Liu, K., Feng, S., Jiang, Y. et al. Identification and expression analysis of seven MADS-box genes from Annona squamosa . Biol Plant 61, 24–34 (2017). https://doi.org/10.1007/s10535-016-0688-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-016-0688-1