Abstract
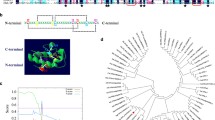
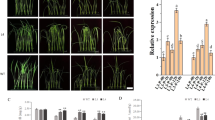
Rice (Oryza sativa L.) is a salt-sensitive species. Salt stress can cause injury to the plant cellular membrane. Plant lipid transfer proteins (LTPs) are abundant lipid binding proteins that are important in membrane vesicle biogenesis and trafficking, however, the biological importance of LTPs on salt-stress response in rice remains unclear. Therefore, salt-responsive rice LTPs were identified and characterized in this study. Microarray analysis showed seven genes positively regulated by salinity, including five Ltp genes (LtpII.3, LtpII.5, LtpII.6, LtpV.1, and LtpV.2) and two Ltp-like (LtpL; LtpL1, and LtpL2) genes. Amino acid alignment revealed that all these Ltp and LtpL genes contained the N-terminal signal peptide. Apart from LtpL1, all salt-inducible Ltp genes had the conserved eight cysteine residue motifs backbone. Verification of gene expression to different stimuli in rice seedlings revealed that salt-regulated Ltp genes differentially responded to drought, cold, H2O2, abscisic acid (ABA) and CaCl2. Furthermore, the expression of Ltp and LtpL genes was tissue-specifically regulated by ABA-dependent and independent pathway. In silico analysis of a 1.5-kb 5’-upstream region of these genes showed regulatory cis-elements associated with ABA, calcium, and cold/drought responses. Three LtpII subfamily genes, including LtpII.3, LtpII.5, and LtpII.6, were strictly expressed in flowers and seeds, and LtpIII.1 mRNA strongly accumulated in stem tissue. Subcellular localization analysis of LTP-DsRed fusion proteins revealed that the five LTPs and two LTPLs localized at the endoplasmic reticulum. The results provide new clues to further understanding the biological functions of Ltp genes.
Similar content being viewed by others
Abbreviations
- ABA:
-
abscisic acid
- ABRE:
-
ABA-responsive element
- 8-CM:
-
eight-cysteine motif
- DRE/CRT:
-
dehydration-responsive/C-repeat element
- ER:
-
endoplasmic reticulum
- LTPs:
-
lipid transfer proteins
- LtpL:
-
Ltp-like
- MYB:
-
MYB transcription factor recognition sequence
- MYC:
-
MYC transcription factor recognition sequence
- ROS:
-
reactive oxygen species
- ROSE:
-
ROS/oxidative stress-responsive element
- RT:
-
reverse transcription
- TNG67:
-
Tainung 67
References
Ahmad, P., Hakeem, K.R., Kumar, A., Ashraf, M., Akram, N.A.: Salt-induced changes in photosynthetic activity and oxidative defense system of three cultivars of mustard (Brassica juncea L.). — Afr. J. Biotechnol. 11: 2694–2703, 2012.
Beisson, F., Koo, A.J., Ruuska, S., Schwender, J., Pollard, M., Thelen, J.J., Paddock, T., Salas, J.J., Savage, L., Milcamps, A., Mhaske, V.B., Cho, Y., Ohlrogge, J.B.: Arabidopsis genes involved in acyl lipid metabolism. A 2003 census of the candidates, a study of the distribution of expressed sequence tags in organs, and a web-based database. — Plant Physiol. 132: 681–697, 2003.
Boutrot, F., Chantret, N., Gautier, M.F.: Genome-wide analysis of the rice and Arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining. — BMC Genomics 9: 86–104, 2008.
Charvolin, D., Douliez, J.P., Marion, D., Cohen-Addad, C., Pebay-Peyroula, E.: The crystal structure of a wheat nonspecific lipid transfer protein (ns-LTP1) complexed with two molecules of phospholipid at 2.1 Å resolution. — Eur. J. Biochem. 264: 562–568, 1999.
Christensen, A.H., Quail, P.H.: Ubiquitin promoter-based vectors for high-level expression of selectable and/or screenable marker genes in monocotyledonous plants. — Transgen. Res. 5: 213–218, 1996.
Crooks, G.E., Hon, G., Chandonia, J.M., Brenner, S.E.: WebLogo: a sequence logo generator. — Genome Res. 14:1188–1190, 2004.
Dionisio-Sese, M.L., Tobita, S.: Antioxidant responses of rice seedlings to salinity stress. — Plant Sci. 135:1–9, 1998.
Diz, M.S., Carvalho, A.O., Ribeiro, S.F., Da Cunha, M., Beltramini, L., Rodrigues, R., Nascimento, V.V., Machado, O.L., Gomes, V.M.: Characterisation, immunolocalisation and antifungal activity of a lipid transfer protein from chili pepper (Capsicum annuum) seeds with novel α-amylase inhibitory properties. — Physiol. Plant. 142: 233–246, 2011.
Edstam, M.M., Laurila, M., Höglund, A., Raman, A., Dahlström, K.M., Salminen, T.A., Edqvist, J., Blomqvist, K: Characterization of the GPI-anchored lipid transfer proteins in the moss Physcomitrella patens. - Plant Physiol. Biochem. 75: 55–69, 2014.
Edstam, M.M., Viitanen, L., Salminen, T.A., Edqvist, J.: Evolutionary history of the non-specific lipid transfer proteins. — Mol. Plant 4: 947–964, 2011.
Emanuelsson, O., Nielsen, H., Von Heijne, G.: ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. — Protein Sci. 8: 978–984, 1999.
Gao, G., **, L.P., **e, K.Y., Qu, D.Y.: The potato StLTPa7 gene displays a complex Ca-associated pattern of expression during the early stage of potato-Ralstonia solanacearum interaction. — Mol. Plant Pathol. 10: 15–27, 2009.
Garcı́a-Garrido, J.M., Menossi, M., Puigdoménech, P., Martı́nez-Izquierdo, J.A., Delseny, M. Characterization of a gene encoding an abscisic acid-inducible type-2 lipid transfer protein from rice. - FEBS Lett. 428: 193–199, 1998.
Gill, S.S., Tuteja, N.: Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. — Plant Physiol. Biochem. 48: 909–930, 2010.
Grattan, S.R., Grieve, C.M.: Mineral nutrient acquisition and response by plants grown in saline environments. - In: Pessarakli, M. (ed.): Handbook of Plant and Crop Stress. Pp. 203–229. Marcel Dekker, New York 1999.
Gueta-Dahan, Y., Yaniv, Z., Zilinskas, B.A., Ben-Hayyim, G.: Salt and oxidative stress: similar and specific responses and their relation to salt tolerance in citrus. — Planta 203: 460–469, 1997.
Guiderdoni, E., Cordero, M.J., Vignols, F., García-Garrido, J.M., Lescot, M., Tharreau, D., Meynard, D., Ferrière, N., Notteghem, J.L., Delseny, M.: Inducibility by pathogen attack and developmental regulation of the rice Ltp1 gene. — Plant mol. Biol. 49: 683–699, 2002.
Guo, L., Yang, H., Zhang, X., Yang S.: Lipid transfer protein 3 as a target of MYB96 mediates freezing and drought stress in Arabidopsis. — J. exp. Bot. 64: 1755–1767, 2013.
Higo, K., Ugawa, Y., Iwamoto, M., Korenaga, T.: Plant cis-acting regulatory DNA elements (PLACE) database. — Nucl. Acids Res. 27: 297–300, 1999.
Heinemann, B., Andersen, K.V., Nielsen, P.R., Bech, L.M., Poulsen, F.M.: Structure in solution of a four-helix lipid binding protein. — Protein Sci. 5: 13–23, 1996.
Holmberg, N., Bülow, L.: Improving stress tolerance in plants by gene transfer. — Trends Plant Sci. 3: 61–66, 1998.
Jang, C.S., Lee, H.J., Chang, S.J., Seo, Y.W.: Expression and promoter analysis of the TaLTP1 gene induced by drought and salt stress in wheat (Triticum aestivum L.). — Plant Sci. 167: 995–1001, 2004.
Jose-Estanyol, M., Gomis-Ruth, F.X., Puigdomenech, P.: The eight-cysteine motif, a versatile structure in plant proteins. — Plant Physiol. Biochem. 42: 355–365, 2004.
Kader, J.C.: Lipid-transfer proteins in plants. — Annu. Rev. Plant Physiol. Plant mol. Biol. 47: 627–654, 1996.
Kader, J.C.: Lipid transfer proteins: a puzzling family of plant proteins. — Trends Plant Sci. 2: 66–70, 1997.
Kaplan, B., Davydov, O., Knight, H., Galon, Y., Knight, M.R., Fluhr, R., Fromm, H.: Rapid transcriptome changes induced by cytosolic Ca2+ transients reveal ABRE-related sequences as Ca2+-responsive cis elements in Arabidopsis. — Plant Cell 18: 2733–2748, 2006.
Kawahara, Y., De la Bastide, M., Hamilton, J.P., Kanamori, H., McCombie, W.R., Ouyang, S., Schwartz, D.C., Tanaka, T., Wu, J., Zhou, S., Childs, K.L., Davidson, R.M., Lin, H., Quesada-Ocampo, L., Vaillancourt, B., Sakai, H., Lee, S.S., Kim, J., Numa, H., Itoh, T., Buell, C.R., Matsumoto, T.: Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. — Rice 6: 4, 2013.
Kim, T.H., Park, J.H., Kim, M.C., Cho, SH: cutin monomer induces expression of the rice OsLTP5 lipid transfer protein gene. - J. Plant Physiol. 165: 345–349, 2008.
Larkin, M.A., Blackshields, G., Brown, N.P., Chenna, R., McGettigan, P.A., McWilliam, H., Valentin, F., Wallace, I.M., Wilm, A., Lopez, R., Thompson, J.D., Gibson, T.J., Higgins, D.G.: Clustal W and Clustal X version 2.0. — Bioinformatics 23: 2947–2948, 2007.
Liu, F., Zhang, X., Lu, C., Zeng, X., Li, Y., Fu, D., Wu, G.: Non-specific lipid transfer proteins in plants: presenting new advances and an integrated functional analysis. — J. exp. Bot. 66: 5663–5681, 2015.
Liu, X., Shangguan, Y., Zhu, J., Lu, Y., Han, B.: The rice OsLTP6 gene promoter directs anther-specific expression by a combination of positive and negative regulatory elements. — Planta 238: 845–857, 2013.
Madrid, S.M.: The barley lipid transfer protein is targeted into the lumen of the endoplasmic reticulum. — Plant Physiol. Biochem. 29: 659–703, 1991.
Matz, M.V., Fradkov, A.F., Labas, Y.A., Savitsky, A.P., Zaraisky, A.G., Markelov, M.L., Lukyanov S, A.: Fluorescent proteins from nonbioluminescent Anthozoa species. — Nat. Biotechnol. 17: 969–973, 1999.
Mittler, R.: Oxidative stress, antioxidants and stress tolerance. — Trends Plant Sci. 7: 405–410, 2002.
Mittova, V., Guy, M., Tal, M., Volokita, M.: Salinity upregulates the antioxidative system in root mitochondria and peroxisomes of the wild salt-tolerant tomato species Lycopersicon pennellii. — J. exp. Bot. 55: 1105–1113, 2004.
Munns, R., Tester, M.: Mechanisms of salinity tolerance. — Annu. Rev. Plant Biol. 59: 651–681, 2008.
Pagnussat, L., Burbach, C., Baluska, F., De la Canal, L.: An extracellular lipid transfer protein is relocalized intracellularly during seed germination. — J. exp. Bot. 63: 6555–6563, 2012.
Potocka, I., Baldwin, T.C., Kurczynska, E.U.: Distribution of lipid transfer protein 1 (LTP1) epitopes associated with morphogenic events during somatic embryogenesis of Arabidopsis thaliana. — Plant Cell Rep. 31: 2031–2045, 2012.
Safi, H., Saibi, W., Alaoui, M.M., Hmyene, A., Masmoudi, K., Hanin, M., Brini, F.: A wheat lipid transfer protein (TdLTP4) promotes tolerance to abiotic and biotic stress in Arabidopsis thaliana. — Plant Physiol. Biochem. 89: 64–75, 2015.
Salama, K.H.A., Mansour, M.M.F., Ali, F.Z.M., Abou-Hadid, A.F.: NaCl-induced changes in plasma membrane lipids and proteins of Zea mays L. cultivars in their response to salinity. — Acta. Physiol. Plant. 29: 351–359, 2007.
Seo, P.J., Lee, S.B., Suh, M.C., Park, M.J., Go, Y.S., Park, C.M.: The MYB96 transcription factor regulates cuticular wax biosynthesis under drought conditions in Arabidopsis. — Plant Cell 23:1138-1152, 2011.
Shin, D.H., Lee, J.Y., Hwang, K.Y., Kim, K.K., Suh, S.W.: High resolution crystal structure of the nonspecific lipidtransfer protein from maize seedlings. — Structure 3: 189–199, 1995.
Silverstein, K.A., Moskal, W.A.J., Wu, H.A., Underwood, B.A., Graham, M.A., Town, C.D., Van den Bosch, K.A.: Small cysteine-rich peptides resembling antimicrobial peptides have been under-predicted in plants. — Plant J. 51: 262–280, 2007.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., Kumar, S.: MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. — Mol. Biol. Evol. 28: 2731–2739, 2011.
Tapia, G., Morales-Quintana, L., Parra, C., Berbel, A., Alcorta, M.: Study of nsLTPs in Lotus japonicus genome reveal a specific epidermal cell member (LjLTP10) regulated by drought stress in aerial organs with a putative role in cutin formation. — Plant mol. Biol. 82: 485–501, 2013.
Torres-Schumann, S., Godoy, J.A., Pintor-Toro, J.A.: A probable lipid transfer protein gene is induced by NaCl in stems of tomato plants. — Plant mol. Biol. 18: 749–757, 1992.
Tsuboi, S., Osafune, T., Tsugeki, R., Nishimura, M., Yamada, M.: Nonspecific lipid transfer protein in castor bean cotyledon cells: subcellular localization and a possible role in lipid metabolism. — J. Biochem. 111: 500–508, 1992.
Vignols, F., Lund, G., Pammi, S., Tremousaygue, D., Grellet, F., Kader, J.C., Puigdomenech, P., Delseny, M.: Characterization of a rice gene coding for a lipid transfer protein. — Gene 142: 265–270, 1994.
Vignols, F., Wigger, M., García Garrido, J.M., Grellet, F., Kader, J.C., Delseny, M.: Rice lipid transfer protein (LTP) genes belong to a complex multigene family and are differentially regulated. — Gene 195: 177–186, 1997.
Wahid, A., Gelani, S., Ashraf, M., Foolad, M.: Heat tolerance in plants: an overview. — Environ. exp. Bot. 61: 199–223, 2007.
Wang, C., **e, C., Chi, F., Hu, W., Mao, G., Sun, D., Li, C., Sun, Y.: BcLTP, a novel lipid transfer protein in Brassica chinensis, may secrete and combine extracellular CaM. — Plant Cell Rep. 27: 159–169, 2008.
Wang, F., Zang, X., Kabir, M.R., Liu, K., Liu, Z., Ni, Z., Yao, Y., Hu, Z., Sun, Q., Peng, H.A.: Wheat lipid transfer protein 3 could enhance the basal thermotolerance and oxidative stress resistance of Arabidopsis. — Gene 550: 18–26, 2014.
Wang, H.W., Hwang, S.G., Karuppanapandian, T., Liu, A., Kim, W., Jang, C.S.: Insight into the molecular evolution of nonspecific lipid transfer proteins via comparative analysis between rice and sorghum. — DNA Res. 19: 179–194, 2012.
Wu, T.M., Lin, K.C., Liau, W.S., Chao, Y.Y., Yang, L.H., Chen, S.Y., Lu, C.A., Hong, C.Y.: A set of GFP-based organelle marker lines combined with DsRed-based gateway vectors for subcellular localization study in rice (Oryza sativa, L.). — Plant mol. Biol. 90: 107–115, 2015.
Yeats, T.H., Rose, J.K.C.: The biochemistry and biology of extracellular plant lipid-transfer proteins (LTPs). — Protein Sci. 17: 191–198, 2008.
Yoo, S.D., Cho, Y.H., Sheen, J.: Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. — Nat. Protocols 2: 1565–1572, 2007.
Yoshida, S., Forno, D.A., Cock, J.H., Gomez, K.A.: Laboratory Manual for Physiological Studies of Rice. 2nd Ed. — The International Rice Research Institute, Los Baños 1972.
Yu, G., Hou, W., Du, X., Wang, L., Wu, H., Zhao, L., Kong, L., Wang, H.: Identification of wheat non-specific lipid transfer proteins involved in chilling tolerance. — Plant Cell Rep. 33: 1757–1766, 2014.
Yu, K., Soares, J.M., Mandal, M.K., Wang, C., Chanda, B., Gifford, A.N., Fowler, J.S., Navarre, D., Kachroo, A., Kachroo, P.: A feedback regulatory loop between G3P and lipid transfer proteins DIR1 and AZI1 mediates azelaicacid-induced systemic immunity. — Cell Rep. 3: 1266–1278, 2013.
Yubero-Serrano, E.M., Moyano, E., Medina-Escobar, N., Muñoz-Blanco, J., Caballero, J.L.: Identification of a strawberry gene encoding a non-specific lipid transfer protein that responds to ABA, wounding and cold stress. — J. exp. Bot. 54: 1865–1877, 2003.
Zamani, S., Bybordi, A., Khorshidi, A., Nezami, T.: Effects of NaCl salinity levels on lipids and proteins of canola (Brassica napus L.) cultivars. — Adv. environ. Biol. 4: 397–403, 2010.
Zhang, D., Liang, W., Yin, C., Zong, J., Gu, F.: OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. — Plant Physiol. 154: 49–162, 2010.
Zhang, Y., Su, J., Duan, S., Ao, Y., Dai, J., Liu, J., Wang, P., Li, Y., Liu, B., Feng, D., Wang, J., Wang, H.: A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. — Plant Methods 7: 30–43, 2011.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgments: We are grateful to the Joint Center for Instruments and Researches of the College of Bioresources and Agriculture at the National Taiwan University for confocal microscopy and technical support. This work was supported by a research grant (MOST 104-2313-B-002 -013 -MY3) from the Ministry of Science and Technology of Taiwan to C.-Y. Hong.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lin, KC., Wu, TM., Chandrika, N.N.P. et al. Molecular characterization and subcellular localization of salt-inducible lipid transfer proteins in rice. Biol Plant 61, 501–510 (2017). https://doi.org/10.1007/s10535-016-0671-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-016-0671-x