Abstract
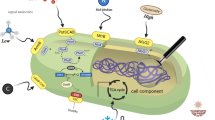
Actinomyces are gram-positive bacteria known for their valuable secondary metabolites. Redirecting metabolic flux towards desired products in actinomycetes requires precise and dynamic regulation of gene expression. In this study, we integrated the CRISPR interference (CRISPRi) system with a cumate-inducible promoter to develop an inducible gene downregulation method in Saccharopolyspora erythraea, a prominent erythromycin-producing actinobacterium. The functionality of the cumate-inducible promoter was validated using the gusA gene as a reporter, and the successful inducible expression of the dCas9 gene was confirmed. The developed inducible CRISPRi strategy was then employed to downregulate the expression of target genes rppA in the wild-type strain NRRL2338 and sucC in the high erythromycin-producing strain E3. Through dynamic control of sucC expression, a significant enhancement in erythromycin production was achieved in strain E3. This study demonstrated the effectiveness of an inducible gene downregulation approach using CRISPRi and a cumate-inducible promoter, providing valuable insights for optimizing natural product production in actinomyces.






Similar content being viewed by others
References
Alam MT, Merlo ME, Hodgson DA, Wellington EM, Takano E, Breitling R (2010) Metabolic modeling and analysis of the metabolic switch in Streptomyces coelicolor. BMC Genomics 11:202. https://doi.org/10.1186/1471-2164-11-202
Baltz RH (2016) Genetic manipulation of secondary metabolite biosynthesis for improved production in Streptomyces and other actinomycetes. J Ind Microbiol Biotechnol 43(2–3):343–370. https://doi.org/10.1007/s10295-015-1682-x
Bu Q, Li Y, **e H, Li J, Lv Z, Su Y, Li Y (2021) Rational engineering strategies for achieving high-yield, high-quality and high-stability of natural product production in actinomycetes. Metab Eng 67:198–215. https://doi.org/10.1016/j.ymben.2021.06.003
Choi YJ, Morel L, Le Francois T, Bourque D, Bourget L, Groleau D, Massie B, Miguez CB (2010) Novel, versatile, and tightly regulated expression system for Escherichia coli strains. Appl Environ Microbiol 76(15):5058–5066. https://doi.org/10.1128/AEM.00413-10
Herai S, Hashimoto Y, Higashibata H, Maseda H, Ikeda H, Omura S, Kobayashi M (2004) Hyper-inducible expression system for streptomycetes. Proc Natl Acad Sci USA 101(39):14031–14035. https://doi.org/10.1073/pnas.0406058101
Horbal L, Fedorenko V, Luzhetskyy A (2014) Novel and tightly regulated resorcinol and cumate-inducible expression systems for Streptomyces and other actinobacteria. Appl Microbiol Biotechnol 98(20):8641–8655. https://doi.org/10.1007/s00253-014-5918-x
Horbal L, Siegl T, Luzhetskyy A (2018) A set of synthetic versatile genetic control elements for the efficient expression of genes in Actinobacteria. Sci Rep 8(1):491. https://doi.org/10.1038/s41598-017-18846-1
Hwang KS, Kim HU, Charusanti P, Palsson B, Lee SY (2014) Systems biology and biotechnology of Streptomyces species for the production of secondary metabolites. Biotechnol Adv 32(2):255–268. https://doi.org/10.1016/j.biotechadv.2013.10.008
Ke X, Jiang X, Huang M, Tian X, Chu J (2022) Engineering of succinyl-CoA metabolism in view of succinylation regulation to improve the erythromycin production. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-022-12060-4
Kim DK, Gu B, Kim DG, Oh MK (2023) Quorum sensing-based metabolic engineering of the precursor supply in Streptomyces coelicolor to improve heterologous production of neoaureothin. Biotechnol Bioeng 120(7):2039–2044. https://doi.org/10.1002/bit.28400
Magdevska V, Gaber R, Goranovič D, Kuščer E, Boakes S, Durán Alonso MB, Santamaría RI, Raspor P, Leadlay PF, Fujs S, Petković H (2010) Robust reporter system based on chalcone synthase rppA gene from Saccharopolyspora erythraea. J Microbiol Methods 83(2):111–119. https://doi.org/10.1016/j.mimet.2010.08.001
Myronovskyi M, Luzhetskyy A (2016) Native and engineered promoters in natural product discovery. Nat Prod Rep 33(8):1006–1019. https://doi.org/10.1039/c6np00002a
Myronovskyi M, Welle E, Fedorenko V, Luzhetskyy A (2011) Beta-glucuronidase as a sensitive and versatile reporter in actinomycetes. Appl Environ Microbiol 77(15):5370–5383. https://doi.org/10.1128/AEM.00434-11
Noguchi Y, Kashiwagi N, Uzura A, Ogino C, Kondo A, Ikeda H, Sota M (2018) Development of a strictly regulated xylose-induced expression system in Streptomyces. Microb Cell Factories 17(1):151. https://doi.org/10.1186/s12934-018-0991-y
Rateb ME, Ebel R, Jaspars M (2018) Natural product diversity of actinobacteria in the Atacama Desert. Antonie Van Leeuwenhoek 111(8):1467–1477. https://doi.org/10.1007/s10482-018-1030-z
Rodríguez-García A, Combes P, Pérez-Redondo R, Smith MC, Smith MC (2005) Natural and synthetic tetracycline-inducible promoters for use in the antibiotic-producing bacteria Streptomyces. Nucleic Acids Res 33(9):e87. https://doi.org/10.1093/nar/gni086
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3(6):1101–1108. https://doi.org/10.1038/nprot.2008.73
Seo SO, Schmidt-Dannert C (2019) Development of a synthetic cumate-inducible gene expression system for Bacillus. Appl Microbiol Biotechnol 103(1):303–313. https://doi.org/10.1007/s00253-018-9485-4
Takano E, White J, Thompson CJ, Bibb MJ (1995) Construction of thiostrepton-inducible, high-copy-number expression vectors for use in Streptomyces spp. Gene 166(1):133–137. https://doi.org/10.1016/0378-1119(95)00545-2
Tian J, Yang G, Gu Y, Sun X, Lu Y, Jiang W (2020) Develo** an endogenous quorum-sensing based CRISPRi circuit for autonomous and tunable dynamic regulation of multiple targets in Streptomyces. Nucleic Acids Res 48(14):8188–8202. https://doi.org/10.1093/nar/gkaa602
Tong Y, Whitford CM, Blin K, Jorgensen TS, Weber T, Lee SY (2020) CRISPR-Cas9, CRISPRi and CRISPR-BEST-mediated genetic manipulation in streptomycetes. Nat Protoc 15(8):2470–2502. https://doi.org/10.1038/s41596-020-0339-z
Wang W, Li S, Li Z, Zhang J, Fan K, Tan G, Ai G, Lam SM, Shui G, Yang Z, Lu H, ** P, Li Y, Chen X, **a X, Liu X, Dannelly HK, Yang C, Yang Y, Zhang S, Alterovitz G, **ang W, Zhang L (2020) Harnessing the intracellular triacylglycerols for titer improvement of polyketides in Streptomyces. Nat Biotechnol 38(1):76–83. https://doi.org/10.1038/s41587-019-0335-4
Xu F, Ke X, Hong M, Huang M, Chen C, Tian X, Hang H, Chu J (2021) Exploring the metabolic fate of propanol in industrial erythromycin-producing strain via (13)C labeling experiments and enhancement of erythromycin production by rational metabolic engineering of Saccharopolyspora erythraea. Biochem Biophys Res Commun 542:73–79. https://doi.org/10.1016/j.bbrc.2021.01.024
Zhao Y, Li L, Zheng G, Jiang W, Deng Z, Wang Z, Lu Y (2018) CRISPR/dCas9-mediated multiplex gene repression in Streptomyces. Biotechnol J 13(9):e1800121. https://doi.org/10.1002/biot.201800121
Acknowledgements
We thank Professor Weihong Jiang (Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, China) and Professor Yinhua Lu (Shanghai Normal University, Shanghai, China) for providing pSET-dcas9 and pSET-dCas9-actII4-NT-S1.
Funding
This research was financially supported by the National Key Research Development Program of China (No. 2022YFC2105403), the National Natural Science Foundation of China (No. 21276081), the Scientific Research Think Tank of Biological Manufacturing Industry in Qingdao (QDSWZK202004), the Fundamental Research Funds for the Central Universities (JKF01211116, JKVF1211036, JKF01211708).
Author information
Authors and Affiliations
Contributions
All authors read and approved the manuscript, contributed significantly to the work, and conceived the project. XK, XJ, XWT and JC designed the experiments. XJ and XK performed the experiments and supported by JC and XWT. XJ and XK analyzed the results. XJ, XK wrote the manuscript with the help of XWT and JC.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethics approval
This article does not contain any studies on human or animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article will be submitted to the Bioprocessing and Bioengineering section.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, X., Ke, X., Tian, X. et al. An inducible CRISPRi circuit for tunable dynamic regulation of gene expression in Saccharopolyspora erythraea. Biotechnol Lett 46, 161–172 (2024). https://doi.org/10.1007/s10529-023-03462-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03462-z