Abstract
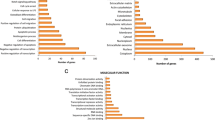
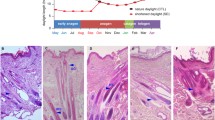
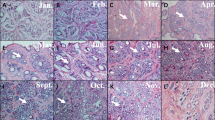
The Cashmere goat (Capra hircus) is renowned for its high-quality fiber production trait. The hair cycle in Cashmere goat has an annual rhythm. To deepen the understanding of the molecular foundation of annual rhythm in the skin of Cashmere goat, we did a comparative analysis of the Cashmere goat skin transcriptome all year round. 4002 Differentially expressed genes (DEGs) were identified with seasonal variations. 12 months transcriptome were divided into four developmental stages: Jan–Mar, Apr–Jul, Aug–Oct, and Nov–Dec based on gene expression patterns. 13 modules of highly correlated genes in skin were identified using WGCNA. Ten of these modules were consistent with the development stages. The gene function of those genes in each module was analyzed by functional enrichment. The results indicated that Wnt and Hedgehog signaling pathways were inhibited from January to March and activated from April to July. The cutaneous immune system of Cashmere goats has high activity from August to October. Fatty acid metabolism dominates goat skin from November to December. This study provides new information related to the annual skin development cycle, which could provide molecular biological significance for understanding the seasonal development and response to the annual rhythm of skin.




Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article. The sequencing data have been submitted to NCBI SRA under BioProject ID: PRJNA000000.
References
Abdallah F, Mijouin L, Pichon C (2017) Skin immune landscape: inside and outside the organism. Mediators Inflamm 2017:5095293
Aubin-Houzelstein G (2012) Notch signaling and the develo** hair follicle. Adv Exp Med Biol 727:142–160
Bai CB, Auerbach W, Lee JS, Stephen D, Joyner AL (2002) Gli2, but not Gli1, is required for initial Shh signaling and ectopic activation of the Shh pathway. Development 129(20):4753–4761
Baroli B (2010) Penetration of nanoparticles and nanomaterials in the skin: fiction or reality? J Pharm Sci 99(1):21–50
Baroni A, Buommino E, De Gregorio V, Ruocco E, Ruocco V, Wolf R (2012) Structure and function of the epidermis related to barrier properties. Clin Dermatol 30(3):257–262
Berthon AS, Szarek E, Stratakis CA (2015) PRKACA: the catalytic subunit of protein kinase A and adrenocortical tumors. Front Cell Dev Biol 3:26
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pages F, Trajanoski Z, Galon J (2009) ClueGO: a cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25(8):1091–1093
Buscher D, Bosse B, Heymer J, Ruther U (1997) Evidence for genetic control of Sonic hedgehog by Gli3 in mouse limb development. Mech Dev 62(2):175–182
Chuong C-M, Randall VA, Widelitz RB, Wu P, Jiang T-X (2012) Physiological regeneration of skin appendages and implications for regenerative medicine. Physiology 27(2):61–72
Dbrowska AK, Rotaru GM, Derler S, Spano F, Rossi RM (2016) Materials used to simulate physical properties of human skin. Skin Res Technol 22(1):3–14
Dbrowska AK, Spano F, Derler S, Adlhart C, Spencer ND, Rossi RM (2017) The relationship between skin function, barrier properties, and body-dependent factors. Skin Res Technol. https://doi.org/10.1111/srt.12424
de Hoon MJ, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20(9):1453–1454
Fan YX, Wu RB, Qiao X, Zhang YJ, Wang RJ, Su R, Wu JH, Dong Y, Li JQ (2015) Hair follicle transcriptome profiles during the transition from anagen to catagen in Cashmere goat (Capra hircus). Genet Mol Res 14(4):17904–17915
Flanagan DJ, Vincan E, Phesse TJ (2019) Wnt signaling in cancer: not a binary ON:OFF switch. Cancer Res 79(23):1362
Fuchs E (2016) Epithelial skin biology: three decades of developmental biology, a hundred questions answered and a thousand new ones to address. Curr Top Dev Biol 116:357–374
Fukuoka Y, Hite MR, Dellinger AL, Schwartz LB (2013) Human skin mast cells express complement factors C3 and C5. J Immunol 191(4):1827–1834
Gabriela Bindea JG, Mlecnik B (2013) CluePedia cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 29(5):661–663
Gong H, Wang H, Wang Y, Bai X, Liu B, He J, Wu J, Qi W, Zhang W (2018) Skin transcriptome reveals the dynamic changes in the Wnt pathway during integument morphogenesis of chick embryos. PLoS ONE 13(1):e0190933
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat Biotechnol 29(7):644–652
Grachtchouk M, Pero J, Yang SH, Ermilov AN, Michael LE, Wang A, Wilbert D, Patel RM, Ferris J, Diener J, Allen M, Lim S, Syu LJ, Verhaegen M, Dlugosz AA (2011) Basal cell carcinomas in mice arise from hair follicle stem cells and multiple epithelial progenitor populations. J Clin Invest 121(5):1768–1781
Kanitakis J (2002) Anatomy, histology and immunohistochemistry of normal human skin. Eur J Dermatol 12(4):390–399
Lalli PN, Strainic MG, Lin F, Medof ME, Heeger PS (2007) Decay accelerating factor can control T cell differentiation into IFN-γ-producing effector cells via regulating local C5a-induced IL-12 production. J Immunol 179(9):5793–5802
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559
Li J, Witten DM, Johnstone IM, Tibshirani R (2012) Normalization, testing, and false discovery rate estimation for RNA-sequencing data. Biostatistics 13(3):523–538
Liu Z, Rebowe RE, Wang Z, Li Y, Wang Z, DePaolo JS, Guo J, Qian C, Liu W (2014) KIF3a promotes proliferation and invasion via Wnt signaling in advanced prostate cancer. Mol Cancer Res 12(4):491–503
Losquadro WD (2017) Anatomy of the skin and the pathogenesis of nonmelanoma skin cancer. Facial Plast Surg Clin North Am 25(3):283–289
Masuya H, Sagai T, Moriwaki K, Shiroishi T (1997) Multigenic control of the localization of the zone of polarizing activity in limb morphogenesis in the mouse. Dev Biol 182(1):42–51
Mill P (2003) Sonic hedgehog-dependent activation of Gli2 is essential for embryonic hair follicle development. Genes Dev 17(2):282
Negera E, Walker SL, Lema T, Aseffa A, Lockwood DN, Dockrell HM (2018) Complement C1q expression in erythema nodosum leprosum. PLoS Negl Trop Dis 12(3):e0006321
Nosenko MA, Ambaryan SG, Drutskaya MS (2019) Proinflammatory cytokines and skin wound healing in mice. Mol Biol (mosk) 53(5):741–754
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14(4):417–419
Peng Q, Li K, Patel H, Sacks SH, Zhou W (2006) Dendritic cell synthesis of C3 is required for full T cell activation and development of a Th1 phenotype. J Immunol 176(6):3330
Plikus MV, Van Spyk EN, Pham K, Geyfman M, Kumar V, Takahashi JS, Andersen B (2015) The circadian clock in skin: implications for adult stem cells, tissue regeneration, cancer, aging, and immunity. J Biol Rhythms 30(3):163–182
Rajesh A, Wise L, Hibma M (2019) The role of langerhans cells in pathologies of the skin. Immunol Cell Biol 97(8):700–713
Rile N, Liu Z, Gao L, Qi J, Zhao M, **e Y, Su R, Zhang Y, Wang R, Li J, **ao H, Li J (2018) Expression of Vimentin in hair follicle growth cycle of inner Mongolian cashmere goats. BMC Genomics 19(1):38
Rimkus TK, Carpenter RL, Qasem S, Chan M, Lo HW (2016) Targeting the sonic hedgehog signaling pathway: review of smoothened and GLI inhibitors. Cancers (basel) 8(2):22
Rishikaysh P, Dev K, Diaz D, Qureshi WM, Filip S, Mokry J (2014) Signaling involved in hair follicle morphogenesis and development. Int J Mol Sci 15(1):1647–1670
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11(3):R25
Schepp-Berglind J, Atkinson C, Elvington M, Qiao F, Mannon P, Tomlinson S (2012) Complement-dependent injury and protection in a murine model of acute dextran sulfate sodium-induced colitis. J Immunol 188(12):6309–6318
Schneider MR, Schmidt-Ullrich R, Paus R (2009) The hair follicle as a dynamic miniorgan. Curr Biol 19(3):R132–R142
Song S, Yang M, Li Y, Rouzi M, Zhao Q, Pu Y, He X, Mwacharo JM, Yang N, Ma Y, Jiang L (2018) Genome-wide discovery of lincRNAs with spatiotemporal expression patterns in the skin of goat during the cashmere growth cycle. BMC Genomics 19(1):495
Spiegelman VS, Tang W, Katoh M, Slaga TJ, Fuchs SY (2002) Inhibition of HOS expression and activities by Wnt pathway. Oncogene 21(5):856–860
Strainic MG, Liu J, Huang D, An F, Lalli PN, Muqim N, Shapiro VS, Dubyak GR, Heeger PS, Medof ME (2008) Locally produced complement fragments C5a and C3a provide both costimulatory and survival signals to naive CD4+ T cells. Immunity 28(3):425–435
Su R, Gong G, Zhang L, Yan X, Wang F, Zhang L, Qiao X, Li X, Li J (2020) Screening the key genes of hair follicle growth cycle in inner Mongolian cashmere goat based on RNA sequencing. Arch Anim Breed 63(1):155–164
Suzuki R, Shimodaira H (2006) Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22(12):1540–1542
Turnham RE, Scott JD (2016) Protein kinase A catalytic subunit isoform PRKACA; history, function and physiology. Gene 577(2):101–108
Veltri A, Lang C, Lien WH (2018) Concise review: Wnt signaling pathways in skin development and epidermal stem cells. Stem Cells 36(1):22–35
Wang E, Harel S, Christiano AM (2016) JAK-STAT signaling jump starts the hair cycle. J Invest Dermatol 136(11):2131–2132
Wang ECE, Higgins CA (2020) Immune cell regulation of the hair cycle. Exp Dermatol 29(3):322–333
Wang X, Tredget EE, Wu Y (2012) Dynamic signals for hair follicle development and regeneration. Stem Cells Dev 21(1):7–18
Yang F, Liu Z, Zhao M, Mu Q, Che T, **e Y, Ma L, Mi L, Li J, Zhao Y (2020) Skin transcriptome reveals the periodic changes in genes underlying cashmere (ground hair) follicle transition in cashmere goats. BMC Genomics 21(1):392
Yoshinori A, Nobuyuki T (2017) Roles of the hedgehog signaling pathway in epidermal and hair follicle development, homeostasis, and cancer. J Develop Biol 5(4):12
Zhang QL, Li JP, Chen Y, Chang Q, Li YM, Yao JY, Jiang HZ, Zhao ZH, Guo D (2014) Growth and viability of Liaoning cashmere goat hair follicles during the annual hair follicle cycle. Genet Mol Res 13(2):4433–4443
Zhu H, Lu W, Laurent C, Shaw GM, Lammer EJ, Finnell RH (2005) Genes encoding catalytic subunits of protein kinase A and risk of spina bifida. Birth Defects Res A Clin Mol Teratol 73(9):591–596
Acknowledgements
I would like to thank my dearest daughter, Han Wu. It would not be possible to write this article without the support from her.
Funding
This work was supported by the Doctoral Scientific Research Foundation of Inner Mongolia University for Nationalities (BS526, BS527) to Dr. Sile Hu and Jianghong Wu, the Natural Science Foundation of Inner Mongolian (2019LH03012, 2018MS03034, 2019MS03078) to Dr. Sile Hu, Jianghong Wu, and Chun Li, the Innovation Foundation of Inner Mongolia Academy of Agricultural and Animal Husbandry Sciences (2017CXJJM03-2) to Dr. Jianghong Wu, and the Opening Foundation of Inner Mongolia Engineering Research Center of Personalized Medicine, the Science Research Project of Inner Mongolia University for the Nationalities (MDK2019088, MDK2019086) to Dr. Sile Hu, Jianghong Wu.
Author information
Authors and Affiliations
Contributions
JW conceived and designed the experiments. SH, CL, WB, and HH performed the experiments. SH and CL analyzed the data. WB, HB, and HH contributed with reagents, materials, and analysis tools. SH and JW wrote the paper. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the grant, scholarship, and/or funding for this study do not lead to any conflict of interest. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Ethical Approval
This manuscript report data were collected from goat. The goat experiments were performed according to the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China; revised in June 2004) and approved by the ethics committee of Inner Mongolia University for Nationalities.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, S., Li, C., Wu, D. et al. The Dynamic Change of Gene-Regulated Networks in Cashmere Goat Skin with Seasonal Variation. Biochem Genet 60, 527–542 (2022). https://doi.org/10.1007/s10528-021-10114-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10114-2