Abstract
Cystathionine-β-synthase (CBS) domain-containing proteins (CDCPs) constitute a large family in plants, and members of this family have been implicated in a variety of biological processes. However, the precise functions and the underlying mechanisms of most members of this family in plants remain to be elucidated. CBSDUF proteins belong to the CDCP superfamily, which contains one domain of unknown function (DUF21) and an N terminus that is adjacent to two intracellular CBS domains. In this study, a comprehensive genome database analysis of soybean was performed to investigate the role(s) of these CBSDUFs and to explore their nomenclature, classification, chromosomal distribution, exon–intron organization, protein structure, and phylogenetic relationships; the analysis identified a total of 18 putative CBSDUF genes. Using specific protein domains and phylogenetic analysis, the CBSDUF gene family was subdivided into eight groups. The soybean CBSDUF genes showed an uneven distribution on 12 chromosomes of Glycine max. RNA-seq transcriptome data from different tissues in public databases revealed tissue-specific and differential expression profiles of the GmCBSDUFs, and qPCR analysis revealed that certain groups of soybean CBSDUFs are likely involved in specific stress responses. In addition, GmCBSDUF3 transgenic Arabidopsis was subjected to phenotypic analysis under NaCl, PEG, and ABA stress treatments. The overexpression of GmCBSDUF3 could enhance tolerance to drought and salt stress in Arabidopsis. This study presents a first comprehensive look at soybean CBSDUF proteins and provides valuable resources for functionally elucidating this protein subgroup within the CBS domain-containing protein family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Expression profiling studies in different organisms have suggested that proteins with unknown functions play important roles in many biological processes (Gollery et al. 2006). These proteins have been divided into two types: one includes proteins with obscure features that lack defined motifs or domains (POFs) and the other includes proteins with defined features that contain at least one previously defined domain or motif (PDFs). Among the latter, a group of proteins containing the cystathionine-β-synthase (CBS) domain might play important roles in stress response/tolerance in Arabidopsis under various stress conditions (Kushwaha et al. 2009). Since the CBS domain was first identified in the Archaebacterium Methanococcus jannaschii (Bateman 1997), CDCPs have been found to represent a large superfamily of evolutionarily conserved proteins. Kushwaha et al. identified CDCPs in whole-genome analyses of Oryza sativa and Arabidopsis thaliana and found that the CBS domain coexists with other functional domain(s) in most of these proteins, which may indicate their probable functions. Based on whether they have additional domain(s), these proteins were further classified into different subclasses: CBSX, CBSCLC, CBSSIS, CBSPPR, CBSIMPDH, CBSCBS, CBSCBSPB and CBSDUF. These subclasses possess various functions, including cytoplasmic targeting, subcellular localization of chloride channels (CLC), protein–protein interaction, protein regulation, sensing of cellular energy status, and maintenance of intracellular ion gradients (Bateman 1997). For example, the highly conserved structure of CBS domains from CLC plays a role in regulating the common gate (Estevez et al. 2004). AKINbc, a CDCP containing four CBS domains, contributes to SnRK1 heterotrimeric complexes and interacts with two proteins implicated in plant pathogen resistance (Gissot et al. 2006). OsCBSX4, a CDCP, could improve abiotic stress tolerance in plants (Singh et al. 2012). OsBi1, a CDCP, could be induced by BPH and is related to resistance to brown plant hopper in rice plants (Wang et al. 2004). OsCBSX3, a CDCP, is involved in rice resistance to M. oryzae (Singh et al. 2012).
However, very few studies have been reported on the CBSDUF subgroup. The CBSDUF subgroup protein contains one domain of unknown function (DUF21) (PF01595) and an N terminus that is adjacent to two intracellular CBS domains. This transmembrane region has no known function. Many of the sequences in this family are annotated as hemolysins because of their similarity to Q54318 (HLYC_BRAHO), which does not contain this domain. Therefore, the functions of DUF21 are still unknown. DUF21 often exists together with CBS domains and plays important roles in plant growth and development. The characteristics of the CBSDUFs in this subgroup are not yet clear. In our previous study, we identified CDCPs in soybean, but there was no detailed analysis of the CBSDUF subgroup. We found that overexpression of soybean GmCBS21, which belongs to the CBSDUF subgroup, possesses a novel function to improve low nitrogen tolerance in A. thaliana in our previous study (Hao et al. 2016). In addition, Sinharoy et al. found that a protein containing the CBS-DUF21 domain from Medicago truncatula is required for rhizobial infection and symbiotic nitrogen fixation (Sinharoy and Liu 2016). Therefore, considering the above studies, we speculate that proteins in the CBSDUF subgroup may play an important role in regulating biotic and abiotic stress, especially in legumes, and are worthy of further exploration. Soybean is one of the most important oil crops in the world and provides a large proportion of the protein used by humans and animals (Kereszt et al. 2007). However, to date, few data (Hao et al. 2016) are available about proteins in the CBSDUF subgroup in soybean. In this study, we took advantage of bioinformatics and publicly available data to identify and analyze soybean CBSDUF genes on a genome-wide scale. A total of 18 CBSDUFs were identified, and their phylogenetic relationships, gene structures, protein structures, conserved motifs, and expression patterns were analyzed in detail. Furthermore, the expression of CBSDUFs in response to various abiotic stresses as well as low nitrogen treatments in a low N-tolerant soybean variety (Pohuang) was determined. Our results provide a basis for further investigation of the evolution and functions of CBSDUFs.
Results
Identification and Phylogenetic Analysis of the Soybean DUF21- and CBS-Domain-Containing Proteins
Eighteen putative GmCBSDUF members were found in the NCBI database and used as queries to conduct BLAST searches against the public genome database (https://phytozome.jgi.doe.gov/pz/portal.html#). If more than one transcript existed, the primary transcript was selected as a representative. Using the same approach, 8, 10, 10, 4, 9, 4, and 4 putative CBSDUF members were identified from common bean (Phaseolus vulgaris), M. truncatula, Lotus japonicus, sorghum, Arabidopsis, rice, and maize, respectively. Table 1 shows the information of CBSDUF genes. Based on available information in the Phytozome 12 database, functional annotations for soybean CBSDUFs were obtained. Less information about the functions of the CBSDUF genes was found. The main functional annotations showed that most of the CBSDUF genes were predicted to be ancient conserved domain protein-related, metal transporter CNNM, or hemolysin-related. The specific functions of these genes remain to be discovered.
A phylogenetic tree was built with 67 protein sequences from eight plant species to investigate the phylogenetic relationships among CBSDUFs from soybean, three other legumes, Arabidopsis, and three gramineous plants (Fig. 1). The soybean CBSDUFs were named GmCBSDUF1 to GmCBSDUF18 according to their chromosomal positions. The genes from the other plant species were named by the same method. Based on the results of phylogenetic tree analysis, we divided these CBSDUFs into eight groups: Group A to Group H (Fig. 1). Group A included 21 members, and it covered eight species. All members of Group B and Group E were dicotyledonous plants. Group C was monocot-specific. Group D did not include legume members. Group F and Group G were legume-specific. The legume CBSDUFs show a very close evolutionary relationship, and the CBSDUFs from gramineous plants show a close evolutionary relationship. Compared to other species, the soybean CBSDUF gene family is extensively expanded. The number of soybean CBSDUFs was almost as many as those from rice, maize, sorghum, and Arabidopsis combined (Table 1). The number of GmCBSDUF genes is approximately two times more than those of Arabidopsis, common bean, M. truncatula, or L. japonicus and four times more abundant than those of rice, maize, or sorghum. The reason for this increase may be the multiple whole-genome duplication events of the soybean genome (Schmutz et al. 2010). The number of CBSDUF genes in dicotyledonous plants is much greater than that in monocotyledonous plants. Therefore, we speculate that CBSDUF plays an important role in dicots than monocots. The phylogenetic relationships may reflect some distinction between legume plant CBSDUFs and the four nonlegume plant CBSDUFs and indicate that the potential biological functions of some CBSDUFs are specific to legume plants.
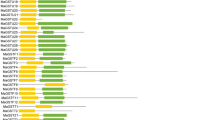
Phylogenetic relationships of the CBSDUFs. Phylogenetic relationships of the CBSDUFs from soybean (Gm), common bean (Pv), Medicago truncatula (Mt), Lotus japonicus (Lj), Arabidopsis (At), rice (Os), maize (Zm), and sorghum (Sb). The phylogenetic tree was constructed using Mega 6.0. The 67 CBSDUF proteins from eight plant species can be divided into eight groups (a–h); the branches are shown in different colors (Color figure online)
Gene Structure and Protein Structure of GmCBSDUFs
Exon–intron structural diversity often plays a key role in the evolution of gene families. To investigate the exon–intron organization of GmCBSDUFs, gene structures were mapped on the basis of the genomic and coding region sequences. The results showed that GmCBSDUFs have 8–15 exons and highly similar gene structures in the conserved region (Fig. 2). The size of GmCBSDUF genes is mainly affected by their intron size. GmCBSDUF12 is the largest gene and has the longest total intron length.
The soybean genome has undergone significant changes in the long-term evolutionary process. Some CBSDUF proteins are highly homologous in the terminal nodes, suggesting that they are putative paralogous pairs. In the study, a total of seven putative paralogous pairs (4/6, 10/14, 11/13, 5/17, 2/3, 8/16, 1/12) were identified, with sequence identities ranging from 60.47 to 99.26%.
To some extent, functional information can be derived from structural similarity. Knowledge of the structure is often essential for interpreting functional data. GmCBSDUF protein structures are shown in Fig. S1. It is clear that GmCBSDUF proteins have a highly conserved hydrophobicity profile, with one hydrophobic segment located at the N terminus. SMART allows the identification and annotation of genetically mobile domains and the analysis of domain architectures. The results are shown in Fig. 3. The major domains are the DUF21 and CBS domains. The DUF21 domain is found in the N terminus of each protein, adjacent to two intracellular CBS domains, and has no known function. In addition, most GmCBSDUF proteins possess 3–4 transmembrane helices except for GmCBSDUF10, GmCBSDUF11, and GmCBSDUF13, which have 2, 1, and 5, respectively. Interestingly, all GmCBSDUFs transmembrane domains pass through the DUF21 domain. Therefore, we speculate that the domain of unknown function DUF21 may play a role in ion channel or signal transduction. In this study, the secondary and tertiary structures of GmCBSDUF proteins were predicted (Fig. 4). The structures were analyzed and compared to the results of Fig. 2. Proteins with high identities also have similar secondary structures, such as GmCBSDUF4/6, GmCBSDUF11/13, GmCBSDUF10/14, GmCBSDUF5/17, GmCBSDUF2/3, GmCBSDUF8/16, and GmCBSDUF1/12. Interaction with a ligand molecule is essential for many proteins to carry out their biological function. This interaction is generally specific, not only in terms of the molecules involved in the interaction but also in the location (i.e., the site of ligand binding) in which the interaction takes place. The results showed that although most GmCBSDUF proteins have similar structures, they have different binding sites, suggesting that they may display different functions.
Main domains detected in soybean CBSDUF proteins by SMART. The blue rectangle represents the transmembrane region; the gray rectangle represents the DUF21 domain; the pink pentagon represents the CBS domain; the green hexagon represents the CorC_HlyC domain; and the orange rectangle represents the SCOP domain (Color figure online)
Protein structure analysis of soybean CBSDUF proteins. a The secondary structure analysis of soybean CBSDUF proteins.  protein binding region,
protein binding region,  polynucelotide-binding region,
polynucelotide-binding region,  helix,
helix,  strand,
strand,  disordered region,
disordered region,  buried,
buried,  exposed,
exposed,  helical transmembrane region. b The tertiary protein structures were predicted by using Phyre2 (Color figure online)
helical transmembrane region. b The tertiary protein structures were predicted by using Phyre2 (Color figure online)
Tissue-Specific Expression Profiling of GmCBSDUFs
Based on the publicly available soybean RNA-Seq data (Libault et al. 2010), the expression patterns of 18 GmCBSDUFs were investigated in various tissues, including (1) root hair cells isolated at 84 h after sowing (HAS), (2) root hair cells isolated at 120 HAS, (3) root tips, (4) roots, (5) mature nodules, (6) leaves, (7) shoot apical meristems, (8) flowers, and (9) green pods. An expression heat map was constructed (Fig. 5a). The results showed that (1) all GmCBSDUFs were expressed in at least one tissue; (2) GmCBSDUF2/3/5 were expressed in all tissues, and their expression levels were relatively high; (3) GmCBSDUF9 had the lowest expression under all conditions; (4) GmCBSDUF8 was expressed only in the underground tissues; and (5) GmCBSDUF9 was expressed only in one shoot apical meristem. In addition, GmCBSDUF1/12 as well as GmCBSDUF16/13 showed similar expression patterns. Moreover, based on the publicly available soybean RNA-Seq data (Libault et al. 2010), expression heat maps of 14 GmCBSDUFs (except GmCBSDUF7/11/13/16, which were not or barely expressed in roots) in root hairs harvested at 12, 24, and 48 h after Bradyrhizobium japonicum inoculation (HAI), in mock-inoculated root hairs at 24 HAI, and in stripped roots at 48 HAI were also constructed (Fig. 5b). Based on the rhizobial inoculation method according to Libault et al. (2010), a B. japonicum cell suspension or water (mock inoculation) was sprayed on soybean seedlings growing on B&D agar medium. The results showed that inoculation with B. japonicum significantly increased the expression of GmCBSDUF8/9, but not other GmCBSDUFs, in root hairs. Therefore, we suspect that GmCBSDUF8/9 may be required for bacterial recognition, nodulation, and nitrogen fixation.
Tissue-specific expression profiles of GmCBSDUF genes. a Gene expression patterns of GmCBSDUF genes in nine different tissues, according to RNA-Seq data (Libault et al. 2010). SAM shoot apical meristem, HAS hours after sowing. b Comparison of the expression of soybean GmCBSDUF genes in root hairs (RH) and stripped roots inoculated (IN) and mock-inoculated (UN) with B. japonicum at 12, 24, and 48 h after B. japonicum inoculation (HAI). HAI IN RH: Root hair inoculated with B. japonicum; HAI UN RH: Root hair not inoculated by B. japonicum. Stripped roots: A soybean root after the strip** of root hairs. The color scale above the heat map indicates gene expression levels. The green color indicates a low expression level, and the red color indicates a high expression level (Color figure online)
Furthermore, the soybean (Glycine max) genome database (Phytozome 12) provides high-resolution gene expression data for a diverse set of 17 soybean GeneAtlas tissue samples, such as flower (open and unopened), lateral root (standard), leaf (ammonia, nitrate, urea, standard and symbiotic condition), nodule (symbiotic condition), root tip (standard), root (ammonia, nitrate, urea, standard and symbiotic condition), shoot tip (standard), stem (standard), and 9 soybean normal tissue samples (flower, leaf, nodule, pod, root, root hair, seed, SAM, and stem). These data were also analyzed and represented as heat maps (Fig. S3). Expression analyses of all GmCBSDUF genes revealed that the different members have different tissue-specific expression. Among all 18 analyzed genes, GmCBSDUF5 showed the highest level of constitutive expression in all tissues, followed by GmCBSDUF3, GmCBSDUF2, and GmCBSDUF12. This high level of constitutive expression indicates a significant role in all these soybean tissues (Fig. S3). A cluster of genes showed low levels of expression in all tissues. They are GmCBSDUF8/9/11/13. GmCBSDUF16 is highly expressed only in root nodules, but its expression is very low in symbiotic conditions. These results are basically consistent with the results in Fig. 5, which makes the analysis of tissue expression patterns of GmCBSDUF genes more sufficient and meaningful. Analysis of the expression patterns of these genes will be helpful to the study of their function. All these expression profiles suggest functional redundancy and divergence among the soybean GmCBSDUFs during plant growth and development.
Promoter Analysis
Based on the soybean genome database (https://www.phytozome.net/soybean), the promoter regions located 2 kb upstream of the translation start codons of the GmCBSDUF genes were analyzed using the PlantCARE promoter analysis program (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/). Multiple elements were identified, and the stress and hormone signaling-related sites are shown in Table 2. The table describes information pertaining to functions, such as elements in response to hormones, including abscisic acid (ABRE, CE1, and MRE) (Narusaka et al. 2003), salicylic acid (TCA element) (Liu et al. 2020), ethylene (ERE) (Song et al. 2009) reported that some AtCBS genes, such as AtCBSX2, AtCBSX3, and AtCBSCBS1, were stably expressed under any stress conditions, while some, such as AtCBSX1 and 15, were more sensitive to all stress conditions in both roots and shoots, and some, such as AtCBSDUFCH2, AtCBSDUF1, AtCBSDUF2, and AtCBSCBS2, were sensitive to stress conditions only in roots. In this study, the expression patterns of soybean CBSDUF genes under abiotic stresses were analyzed (Fig. 6). In contrast to other subgroup members, the results showed that GmCBSDUF7/8/11/16 was upregulated after exposure to cold, drought, salt, and H2O2, while GmCBSDUF17/18 was downregulated by cold, H2O2, salt and ABA, suggesting that these GmCBSDUF genes may play a role in crosstalk between signaling pathways responding to drought, H2O2, salinity, cold, and ABA. The results presented here will be helpful for future studies of the biological functions of GmCBSDUF proteins. Remarkably, we found that GmCBSDUF7/8/11/13/16 showed significant differences in expression under stress treatments. Therefore, we speculate that these genes are inducible and may play an important role in stress response. We will further examine this prospect in subsequent studies.
In conclusion, we performed a comprehensive bioinformatics analysis and provided detailed information on the soybean CBSDUF gene subgroup. Specifically, our results show that the soybean genome contains 18 CBSDUF genes, the largest subgroup among the identified CBSDUF gene subgroups in the study. Our analysis revealed the possible function of each GmCBSDUF gene in response to cold, salt, H2O2, ABA, dehydration, and low nitrogen, identified their potential clients and functional interactions, and revealed the specific responses of some GmCBSDUF genes to specific stresses. By interaction network prediction, some candidate interacting genes were found. At the same time, we preliminarily explored the function of GmCBSDUF3, which might improve the ability to resist abiotic stress in plants. This result provides an impetus for additional investigation of the biological roles and interacting proteins of the CBSDUF protein family in soybean, and a functional analysis of the genes in this family will be carried out systematically. In the future, we will use functional genomics in combination with a transgenic approach to verify the utility of those proteins with defined features as tools to improve stress tolerance in crop plants. Based on the present research and the characteristics of each family member, the research on functional analysis was classified and summarized. We will use gene knockout and transgenic technology to study the functions of the GmCBSDUFs. At the same time, the functions of the two domains, CBS and DUF21, will be studied by site-directed mutagenesis. In addition, due to the lack of information about this family of proteins, the biological pathways involving these genes are still unknown. We will screen for interacting proteins with yeast two-hybrid technology and provide evidence for their mechanisms of action. We will also determine the expression of transgenic plants under specific conditions by high-throughput sequencing technology and infer the gene regulatory network. The ideas provided here would also have a way for expounding the definite role of CBSDUF proteins in plants.
Materials and Methods
Identification of DUF21 and CBS Domain-Containing Proteins in Soybean
The known DUF21 and CBS domain-containing protein sequences from soybean, Arabidopsis, common bean, M. truncatula, L. japonicus, rice, maize, and sorghum were obtained from the NCBI database and used as queries to conduct BLAST searches against the public genome database (https://phytozome.jgi.doe.gov/pz/portal.html#) and L. japonicus genome database (https://www.kazusa.or.jp/lotus/). Sequences with an E value < 1.0 were selected for further analysis. A search with the keywords PF00571 for the CBS domain and PF01595 for the DUF21 domain was conducted for putative soybean CBSDUFs by searching ontologies against the Phytozome (v12.0) database (https://www.phytozome.net). If more than one transcript existed, the primary transcript was selected as a representative.
Phylogenetic, Gene, and Protein Structure Analyses
Multiple alignment analysis was performed with ClustalX 1.83 software (Thompson et al. 1997). Phylogenetic trees were generated by the neighbor-joining (NJ) method and bootstrap analysis (1000 replicates), and phylogenetic analysis was performed using MEGA6 software (Hall 2013). The exon/intron structures of the CBS genes were determined by comparing the coding sequences and corresponding genomic sequences in the gene structure display server (GSDS, https://gsds.cbi.pku.edu.cn/) (Guo et al. 2007). The protein transmembrane topology was predicted using TMHMM Server v2.0, and tertiary protein structures were predicted using Phyre. Domain architecture was analyzed by SMART (a Simple Modular Architecture Research Tool).
Plant Materials and Treatments
For low nitrogen treatment, seeds of a low N-tolerant soybean variety (Pohuang) were germinated. After 7 days, the seedlings were grown hydroponically in half-strength modified Hoagland solution until the first trifoliate leaf was fully developed and then grown in normal nitrogen solution (2 mM Ca(NO3)2·4H2O, 2.5 mM KNO3, 0.5 mM NH4NO3, 0.5 mM KH2PO4, 1 mM MgSO4·7H2O, 0.05 mM Fe-EDTA, 0.005 mM KI, 0.1 mM H3BO3, 0.1 mM MnSO4·H2O, 0.03 mM ZnSO4·7H2O, 0.0001 mM CuSO4·5H2O, 0.001 mM Na2MO4·2H2O, 0.0001 mM CoCl2·6H2O) or low nitrogen solution (0.2 mM Ca(NO3)2·4H2O, 1.8 mM CaCl2·2H2O, 0.25 mM KNO3, 1.125 mM K2SO4, 0.05 mM NH4NO3, 0.5 mM KH2PO4, 1 mM MgSO4·7H2O, 0.05 mM Fe-EDTA, 0.005 mM KI, 0.1 mM H3BO3, 0.1 mM MnSO4·H2O, 0.03 mM ZnSO4·7H2O, 0.0001 mM CuSO4·5H2O, 0.001 mM Na2MO4·2H2O, 0.0001 mM CoCl2·6H2O) at 25 °C in a chamber with a 12-h light and 12-h dark photoperiod. All treatments were performed over a continuous time course (0 h, 0.5 h, 2 h, 6 h, 12 h, and 3, 6, and 9 days). Roots, stems, and leaves from control and stress-treated plants (five plants were collected as mixed samples at each time point) were collected as samples in three biological replicates for RNA preparation, and the samples were quickly frozen in liquid nitrogen and stored at − 80 °C until use.
Soybean seeds were geminated in water at 25 °C in the dark under conditions of a 12--h light and 12-h dark photoperiod and 70% humidity. Salt, dehydration, cold, H2O2, and abscisic acid (ABA) stresses were applied to 2-week-old soybean seedlings. For salt stress, the roots of seedlings were dipped into solutions of 200 mM NaCl. For dehydration, the root systems of whole plants were placed onto filter paper with 70% humidity at room temperature for induction of a rapid drought treatment (Feng et al. 2015). For H2O2 stress, the roots of seedlings were dipped into solutions of 25 mM H2O2. For ABA treatment, soybean seedlings were sprayed with 100 μM ABA. For cold treatment, soybean seedlings were subjected to 4 °C. All stress treatments lasted from 0 to 12 h. Each treatment contained three independent replicates. At 0, 0.5, 5, and 12 h after each treatment, soybean seedlings were harvested, and five plants were collected as mixed samples at each time point, frozen in liquid nitrogen, and stored at − 80 °C until extraction of total RNA for qRT-PCR assays.
Expression Analysis of GmCBSDUFs
Total RNA was isolated from soybean tissues using TRIzol reagent (Invitrogen) and treated with DNase I (Invitrogen) to avoid genomic DNA contamination. First-strand cDNA was synthesized using Superscript II reverse transcriptase (Invitrogen). Gene-specific primers were designed according to gene sequences using Primer 5.0 software (Table S1). The quantitative RT-PCR was performed with a CFX96TM real-time system (Bio-Rad) in a 20 μl system containing 2 μl of a tenfold diluted cDNA, 10 μl of 2 × SYBR green real-time PCR master mix (Takara), and 1 μl each of 10 μM forward and reverse primers. β-actin was used as the internal control. Statistical analyses were performed using the t-test, and p < 0.05 and < 0.01 were considered significant and extremely significant differences, respectively.
Vector Construction, Arabidopsis Transformation, and Stress Treatment
The full-length coding sequence (the primers 5′ ATGGCGGCAGAGATACCG 3′ and 5′ CTATTGATTCCTTAGTGACTCACT 3′.) of GmCBSDUF3 was TA cloned into the plant expression vector pCXSN. The recombinant construct containing the 35S::GmCBSDUF3 (Fig. S2A) cassette was introduced into Agrobacterium tumefaciens strain GV3101 and then transformed into Arabidopsis (Columbia) via the floral dip method. The transgenic plants were screened on MS medium with 100 mg/L hygromycin and confirmed by PCR analyses. The expression levels of GmCBSDUF3 in transgenic plants were determined by qPCR (Fig. S2B).
Seeds of transgenic overexpressing Arabidopsis and WT plants were grown on 10 × 10 cm MS agar plates. They were routinely kept for 2 days in darkness at 4 °C to break dormancy and transferred in a light growth chamber under a day/night 16/8 h cycle at 23 °C. For stress treatment, the seeds of transgenic lines or WT were kept on MS media supplemented with 50 mM NaCl, 2% PEG, or 1.5 μM ABA. Each treatment contained three independent replicates.
References
Bateman A (1997) The structure of a domain common to archaebacteria and the homocystinuria disease protein. Trends Biochem Sci 22:12–13
Brenna A, Talora C (2019) WC-1 and the proximal GATA sequence mediate a Cis-/trans- acting repressive regulation of light-dependent gene transcription in the dark. Int J Mol Sci 20(12):e2854
Collart FR, Osipiuk J, Trent J, Olsen GJ, Huberman E (1996) Cloning and characterization of the gene encoding IMP dehydrogenase from Arabidopsis thaliana. Gene 174:217–220
Diédhiou CJ, Golldack D (2006) Salt-dependent regulation of chloride channel transcripts in rice. Plant Sci 170:793–800
Espinoza C, Medina C, Somerville S, Arce-Johnson P (2007) Senescence-associated genes induced during compatible viral interactions with grapevine and Arabidopsis. J Exp Bot 58:3197–3212
Estevez R, Pusch M, Ferrer-Costa C, Orozco M, Jentsch TJ (2004) Functional and structural conservation of CBS domains from CLC chloride channels. J Physiol 557:363–378
Fabro G, Di Rienzo JA, Voigt CA, Savchenko T, Dehesh K, Somerville S, Alvarez ME (2008) Genome-wide expression profiling Arabidopsis at the stage of Golovinomyces cichoracearum haustorium formation. Plant Physiol 146:1421–1439
Feng ZJ, Cui XY, Cui XY, Chen M, Yang GX, Ma YZ, He GY, Xu ZS (2015) The soybean GmDi19–5 interacts with GmLEA3.1 and increases sensitivity of transgenic plants to abiotic stresses. Front Plant Sci 6:179
Gissot L, Polge C, Jossier M, Girin T, Bouly JP, Kreis M, Thomas M (2006) AKINbetagamma contributes to SnRK1 heterotrimeric complexes and interacts with two proteins implicated in plant pathogen resistance through its KIS/GBD sequence. Plant Physiol 142:931–944
Gollery M, Harper J, Cushman J, Mittler T, Girke T, Zhu JK, Bailey-Serres J, Mittler R (2006) What makes species unique? The contribution of proteins with obscure features. Genome Biol 7:R57
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29:1023–1026
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235
Hao Q, Shang W, Zhang C, Chen H, Chen L, Yuan S, Chen S, Zhang X, Zhou X (2016) Identification and comparative analysis of CBS domain-containing proteins in soybean (Glycine max) and the primary function of GmCBS21 in enhanced tolerance to low nitrogen stress. Int J Mol Sci 17:620
Hechenberger M, Schwappach B, Fischer WN, Frommer WB, Jentsch TJ, Steinmeyer K (1996) A family of putative chloride channels from Arabidopsis and functional complementation of a yeast strain with a CLC gene disruption. J Biol Chem 271:33632–33638
Kereszt A, Li D, Indrasumunar A, Nguyen CD, Nontachaiyapoom S, Kinkema M, Gresshoff PM (2007) Agrobacterium rhizogenes-mediated transformation of soybean to study root biology. Nat Protoc 2:948–952
Kumari S, Sabharwal VP, Kushwaha HR, Sopory SK, Singla-Pareek SL, Pareek A (2009) Transcriptome map for seedling stage specific salinity stress response indicates a specific set of genes as candidate for saline tolerance in Oryza sativa L. Funct Integr Genom 9:109–123
Kushwaha HR, Singh AK, Sopory SK, Singla-Pareek SL, Pareek A (2009) Genome wide expression analysis of CBS domain containing proteins in Arabidopsis thaliana (L.) Heynh and Oryza sativa L. reveals their developmental and stress regulation. BMC Genom 10:200
Li H, Han X, Qiu W, Xu D, Wang Y, Yu M, Hu X, Zhuo R (2019) Identification and expression analysis of the GDSL esterase/lipase family genes, and the characterization of SaGLIP8 in Sedum alfredii hance under cadmium stress. Peer J 7:e6741
Libault M, Farmer A, Joshi T, Takahashi K, Langley RJ, Franklin LD, He J, Xu D, May G, Stacey G (2010) An integrated transcriptome atlas of the crop model Glycine max, and its use in comparative analyses in plants. Plant J 63:86–99
Liu YM, Qu JT, Zhang L, Xu XY, Wei G, Zhao ZF, Ren MZ, Cao MJ (2020) Identification and characterization of the TCA cycle genes in maize. BMC Plant Biol 19(1):592
Lv Q, Tang R, Hua L, Gao X, Li Y, Zheng H, Zhang H (2009) Cloning and molecular analyses of the Arabidopsis thaliana chloride channel gene family. Plant Sci 176:650–661
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J 34:137–148
Sahu BB, Shaw BP (2009) Isolation, identification and expression analysis of salt-induced genes in Suaeda maritima, a natural halophyte, using PCR-based suppression subtractive hybridization. BMC Plant Biol 9:69
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Singh AK, Kumar R, Pareek A, Sopory SK, Singla-Pareek SL (2012) Overexpression of rice CBS domain containing protein improves salinity, oxidative, and heavy metal tolerance in transgenic tobacco. Mol Biotechnol 52:205–216
Sinharoy S, Liu C (2016) A Medicago truncatula cystathionine-beta-synthase-like domain-containing protein is required for rhizobial infection and symbiotic nitrogen fixation. Plant Physiol 170:2204–2217
Song W, Wang F, Chen L, Ma R, Zuo X, Cao A, **e S, Chen X, ** X, Li H (2019) GhVTC1, the key gene for ascorbate biosynthesis in cell elongation under control of ethylene. Cells 8(9):1039
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang X, Ren X, Zhu L, He G (2004) OsBi1, a rice gene, encodes a novel protein with a CBS-like domain and its expression is induced in responses to herbivore feeding. Plant Sci 166:1581–1588
Wang Q, Lai T, Qin G, Tian S (2009) Response of jujube fruits to exogenous oxalic acid treatment based on proteomic analysis. Plant Cell Physiol 50:230–242
Wu C, Zheng C, Ji G, Jiang P (2019) Synergistic effects of HSE and LTR elements from hsp70 gene promoter of Ulva prolifera (Ulvophyceae, Chlorophyta) upon temperature induction1. J Phycol 55(3):738–743
**n S, Tao C, Li H (2016) Cloning and functional analysis of the promoter of an Ascorbate Oxidase gene from Gossypium hirsutum. PLoS ONE 11(9):e0161695
Xu Z, Wang M, Guo Z, Zhu X, **a Z (2019) Identification of a 119-bp promoter of the maize sulfite oxidase gene(ZmSO) that confers high-level gene expression and ABA or drought inducibility in transgenic plants. Int J Mol Sci 20(13):e3326
Yu TF, Zhao WY, Fu JD, Liu YW, Chen M, Zhou YB, Ma YZ, Xu ZS, ** YJ (2018) Genome-wide analysis of CDPK family in foxtail millet and determination of SiCDPK24 Functions in drought stress. Front Plant Sci 9:651
Zhang RX, Qin LJ, Zhao DG (2017) Overexpression of the OsIMP gene increases the accumulation of inositol and confers enhanced cold tolerance in tobacco through modulation of the antioxidant enzymes activities. Genes (Basel) 8(7):179
Acknowledgements
This work was funded by the National Key Research and Development Program of China (2018YFD1000900).
Author information
Authors and Affiliations
Contributions
QH and XZ designed the experiments. QH and YY performed the experiments. QH, WS, CZ, and LC analyzed the data. HC, SY, ZS, XZ, SC, ZY, and DQ contributed to the reagents/materials/analysis tools. QH and XZ wrote the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Research Involving Human and Animal Participants
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10528_2020_9991_MOESM1_ESM.tif
Supplementary file1 (TIF 531 kb)—Figure S1. Transmembrane topology analysis of soybean CBSDUF proteins. The protein transmembrane topology was predicted by using TMHMM Server v2.0. The predicted transmembrane helixes are shown as red peaks.
10528_2020_9991_MOESM2_ESM.tif
Supplementary file2 (TIF 132 kb)—Figure S2. Generation and RT-PCR testing of GmCBSDUF3 lines (A) GmCBSDUF3 overexpression plasmid profile; (B) relative expression levels of GmCBSDUF3 lines.
10528_2020_9991_MOESM3_ESM.tif
Supplementary file3 (TIF 98 kb)—Figure S3. Tissue-specific expression patterns of GmCBSDUF genes. The figure shows the gene expression patterns of GmCBSDUF genes in 17 soybean GeneAtlas tissue samples, including flower (open and unopened), lateral root (standard), leaf (ammonia, nitrate, urea, standard and symbiotic condition), nodule (symbiotic condition), root tip (standard), root (ammonia, nitrate, urea, standard and symbiotic condition), shoot tip (standard), stem (standard) and 9 soybeannormal tissue samples (flower, leaf, nodule, pod, root, root hair, seed, SAM and stem). The data were obtained from the soybean genome database Phytozome 12.
10528_2020_9991_MOESM4_ESM.tif
Supplementary file4 (TIF 572 kb)—Figure S4. Expression of soybean GmCBSDUF genes in response to low nitrogen stresses.A: Leaves, B: stems, C: roots.Data were obtained by real-time PCR normalized against the reference gene ACT11 and are shown as apercentage of expression in the control at 0 h. Blue columns represent the expression under normalnitrogen conditions, and red columns represent the expression under low nitrogen conditions. GmCBSDUF13, which was not expressed in soybean roots, stems and leaves under normal conditions,was not induced under low nitrogen stress and is not shown in this figure.
10528_2020_9991_MOESM6_ESM.xlsx
Supplementary file6 (XLSX 15 kb)—Table S2. Responses of 18 soybean CBSDUF genes to abiotic stresses by qRT-PCR with the 2-ΔΔCT method
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hao, Q., Yang, Y., Shan, Z. et al. Genome-Wide Investigation and Expression Profiling Under Abiotic Stresses of a Soybean Unknown Function (DUF21) and Cystathionine-β-Synthase (CBS) Domain-Containing Protein Family. Biochem Genet 59, 83–113 (2021). https://doi.org/10.1007/s10528-020-09991-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-020-09991-w