Abstract
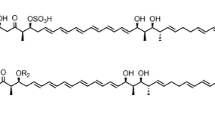
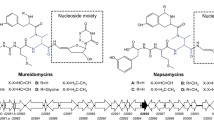
Maduramicin is the most efficient and possesses the largest market share of all anti-coccidiosis polyether antibiotics (ionophore); however, its biosynthetic gene cluster (BGC) has yet to been identified, and the associated strains have not been genetically engineered. Herein, we performed whole-genome sequencing of a maduramicin-producing industrial strain of Actinomadura sp. J1-007 and identified its BGC. Additionally, we analyzed the identified BGCs in silico to predict the biosynthetic pathway of maduramicin. We then developed a conjugation method for the non-spore-forming Actinomadura sp. J1-007, consisting of a site-specific integration method for gene overexpression. The maduramicin titer increased by 30% to 7.16 g/L in shake-flask fermentation following overexpression of type II thioesterase MadTE that is the highest titer at present. Our findings provide insights into the biosynthetic mechanism of polyethers and provide a platform for the metabolic engineering of maduramicin-producing microorganisms for overproduction and development of maduramicin analogs in the future.





Similar content being viewed by others
Abbreviations
- BGC:
-
Biosynthetic gene cluster
- PKS:
-
Polyketide synthase
- ORF:
-
Open reading frame
- T1PKS:
-
Type I polyketide synthase
References
Bhatt A, Stark CB, Harvey BM, Gallimore AR, Demydchuk YA, Spencer JB, Staunton J, Leadlay PF (2005) Accumulation of an E,E,E-triene by the monensin-producing polyketide synthase when oxidative cyclization is blocked. Angew Chem Int Ed Engl 44:7075–7078. https://doi.org/10.1002/anie.200501757
Caldeira C, Neves WS, Cury PM, Serrano P, Baptista MA, Burdmann EA (2001) Rhabdomyolysis, acute renal failure, and death after monensin ingestion. Am J Kidney Dis 38:1108–1112. https://doi.org/10.1053/ajkd.2001.28618
Cane DE, Celmer WD, Westley JW (1983) Unified stereochemical model of polyether antibiotic structure and biogenesis. J Am Chem Soc 105:3594–3600. https://doi.org/10.1021/ja00349a040
Chen X, Li Y, Feng M, Hu X, Zhang H, Zhang R, Dong X, Liu C, Zhang Z, Jiang S, Huang S, Chen L (2019) Maduramicin induces cardiac muscle cell death by the ROS-dependent PTEN/Akt-Erk1/2 signaling pathway. J Cell Physiol 234:10964–10976. https://doi.org/10.1002/jcp.27830
Dairi T, Hamano Y, Furumai T, Oki T (1999) Development of a self-cloning system for Actinomadura verrucosospora and identification of polyketide synthase genes essential for production of the angucyclic antibiotic pradimicin. Appl Environ Microbiol 65:2703–2709. https://doi.org/10.1089/oli.1.1999.9.313
Dalloul RA, Lillehoj HS (2006) Poultry coccidiosis: recent advancements in control measures and vaccine development. Expert Rev Vaccines 5:143–163. https://doi.org/10.1586/14760584.5.1.143
de Jong J, Stoisser B, Wagner K, Tomassen M, Driessen J, Hofmann P, Putzka HA (2004) Determination of maduramicin in feedingstuffs and premixtures by liquid chromatography: development, validation, and interlaboratory study. J AOAC Int 87:1033–1041. https://doi.org/10.1023/B:JANC.0000047015.09774.7b
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Folz SD, Lee BL, Nowakowski LH, Conder GA (1988) Anticoccidial evaluation of halofuginone, lasalocid, maduramicin, monensin and salinomycin. Vet Parasitol 28:1–9. https://doi.org/10.1016/0304-4017(88)90013-1
Gallimore AR, Stark CBW, Bhatt A, Harvey BM, Demydchuk Y, Bolanos-Garcia V, Fowler DJ, Staunton J, Leadlay PF, Spencer JB (2006) Evidence for the role of the monB genes in polyether ring formation during monensin biosynthesis. Chem Biol 13:453–460. https://doi.org/10.1016/j.chembiol.2006.01.013
Harned RL, Hidy PH, Corum CJ, Jones KL (1951) Nigericin a new crystalline antibiotic from an unidentified Streptomyces. Antibiot Chemother (Northfield) 1(9):594–596. https://doi.org/10.1016/S1876-0813(08)60032-9
Harvey BM, Hong H, Jones MA, Hughes-Thomas ZA, Goss RM, Heathcote ML, Bolanos-Garcia VM, Kroutil W, Staunton J, Leadlay PF, Spencer JB (2006) Evidence that a novel thioesterase is responsible for polyketide chain release during biosynthesis of the polyether ionophore monensin. ChemBioChem 7:1435–1442. https://doi.org/10.1002/cbic.200500474
Harvey BM, Mironenko T, Sun Y, Hong H, Deng Z, Leadlay PF, Weissman KJ, Haydock SF (2007) Insights into polyether biosynthesis from analysis of the nigericin biosynthetic gene cluster in Streptomyces sp. DSM4137. Chem Biol 14:703–714. https://doi.org/10.1016/j.chembiol.2007.05.011
Heathcote ML, Staunton J, Leadlay PF (2001) Role of type II thioesterases: evidence for removal of short acyl chains produced by aberrant decarboxylation of chain extender units. Chem Biol 8:207–220. https://doi.org/10.1016/s1074-5521(01)00002-3
Hotta K, Chen X, Paton RS, Minami A, Li H, Swaminathan K, Mathews II, Watanabe K, Oikawa H, Houk KN, Kim CY (2012) Enzymatic catalysis of anti-Baldwin ring closure in polyether biosynthesis. Nature 483:355–358. https://doi.org/10.1038/nature10865
Huang M, Liu B, Liu R, Li J, Liu T (2018) The aglycone polyether nanchangmycin and its homologues exhibit apoptotic and antiproliferative activities against cancer stem cells. ACS Pharmacol Transl Sci 1:84–95. https://doi.org/10.1021/acsptsci.8b00007
Huttel W, Spencer JB, Leadlay PF (2014) Intermediates in monensin biosynthesis: a late step in biosynthesis of the polyether ionophore monensin is crucial for the integrity of cation binding. Beilstein J Org Chem 10:361–368. https://doi.org/10.3762/bjoc.10.34
Jha AK, Paudel S, Dhakal D, Van PT, Ghimire GP, Sohng JK (2015) Genetic evidence for the involvement of glycosyltransferase PdmQ and PdmS in biosynthesis of pradimicin from Actinomadura hibisca. Microbiol Res 174:9–16. https://doi.org/10.1016/j.micres.2015.02.006
Jiang W, Zhu TF (2016) Targeted isolation and cloning of 100-kb microbial genomic sequences by Cas9-assisted targeting of chromosome segments. Nat Protoc 11:960–975. https://doi.org/10.1038/nprot.2016.055
Jiang C, Wang H, Kang Q, Liu J, Bai L (2012) Cloning and characterization of the polyether salinomycin biosynthesis gene cluster of Streptomyces albus XM211. Appl Environ Microbiol 78:994–1003. https://doi.org/10.1128/AEM.06701-11
Jiang W, Zhao X, Gabrieli T, Lou C, Ebenstein Y, Zhu TF (2015) Cas9-assisted targeting of chromosome segments CATCH enables one-step targeted cloning of large gene clusters. Nat Commun 6:8101. https://doi.org/10.1038/ncomms9101
Kotowska M, Pawlik K (2014) Roles of type II thioesterases and their application for secondary metabolite yield improvement. Appl Microbiol Biotechnol 98:7735–7746. https://doi.org/10.1007/s00253-014-5952-8
Kouyoumdjian JA, Morita MP, Sato AK, Pissolatti AF (2001) Fatal rhabdomyolysis after acute sodium monensin (Rumensin) toxicity: case report. Arq Neuro-psiquiatr 59:596–598. https://doi.org/10.1590/s0004-282x2001000400022
Leadlay PF, Staunton J, Oliynyk M, Bisang C, Cortes J, Frost E, Hughes-Thomas ZA, Jones MA, Kendrew SG, Lester JB, Long PF, McArthur HA, McCormick EL, Oliynyk Z, Stark CB, Wilkinson CJ (2001) Engineering of complex polyketide biosynthesis-insights from sequencing of the monensin biosynthetic gene cluster. J Ind Microbiol Biotechnol 27:360–367. https://doi.org/10.1038/sj/jim/7000204
Li C, Hazzard C, Florova G, Reynolds KA (2009) High titer production of tetracenomycins by heterologous expression of the pathway in a Streptomyces cinnamonensis industrial monensin producer strain. Metab Eng 11:319–327. https://doi.org/10.1016/j.ymben.2009.06.004
Liu CM, Hermann TE, Downey A, Prosser BL, Schildknecht E, Palleroni NJ, Westley JW, Miller PA (1983) Novel polyether antibiotics X-14868A, B, C, and D produced by a Nocardia. Discovery, fermentation, biological as well as ionophore properties and taxonomy of the producing culture. J Antibiot (Tokyo) 36:343–350. https://doi.org/10.7164/antibiotics.36.343
Liu T, You D, Valenzano C, Sun Y, Li J, Yu Q, Zhou X, Cane DE, Deng Z (2006) Identification of NanE as the thioesterase for polyether chain release in nanchangmycin biosynthesis. Chem Biol 13:945–955. https://doi.org/10.1016/j.chembiol.2006.07.006
Liu T, Lin X, Zhou X, Deng Z, Cane DE (2008) Mechanism of thioesterase-catalyzed chain release in the biosynthesis of the polyether antibiotic nanchangmycin. Chem Biol 15:449–458. https://doi.org/10.1016/j.chembiol.2008.04.006
Liu T, Cane DE, Deng Z (2009) The enzymology of polyether biosynthesis. Methods Enzymol 459:187–214. https://doi.org/10.1016/S0076-6879(09)04609-6
Liu R, Deng Z, Liu T (2018) Streptomyces species: ideal chassis for natural product discovery and overproduction. Metab Eng 50:74–84. https://doi.org/10.1016/j.ymben.2018.05.015
Logan NB, McKenzie ME, Conway DP, Chappel LR, Hammet NC (1993) Anticoccidial efficacy of semduramicin. 2. Evaluation against field isolates including comparisons with salinomycin, maduramicin, and monensin in battery tests. Poult Sci 72:2058–2063. https://doi.org/10.3382/ps.0722058
Lu C, Zhang X, Jiang M, Bai L (2016) Enhanced salinomycin production by adjusting the supply of polyketide extender units in Streptomyces albus. Metab Eng 35:129–137. https://doi.org/10.1016/j.ymben.2016.02.012
Ma T, Zhou Y, Li X, Zhu F, Cheng Y, Liu Y, Deng Z, Liu T (2016) Genome mining of astaxanthin biosynthetic genes from Sphingomonas sp. ATCC 55669 for heterologous overproduction in Escherichia coli. Biotechnol J 11:228–237. https://doi.org/10.1002/biot.201400827
Maron MI, Magle CT, Czesny B, Turturice BA, Huang R, Zheng W, Vaidya AB, Williamson KC (2015) Maduramicin rapidly eliminates malaria parasites and potentiates the gametocytocidal activity of the pyrazoleamide PA21A050. Antimicrob Agents Chemother 60:1492–1499. https://doi.org/10.1128/AAC.01928-15
Mead JR, You X, Pharr JE, Belenkaya Y, Arrowood MJ, Fallon MT, Schinazi RF (1995) Evaluation of maduramicin and alborixin in a SCID mouse model of chronic cryptosporidiosis. Antimicrob Agents Chemother 39:854–858. https://doi.org/10.1128/aac.39.4.854
Migita A, Watanabe M, Hirose Y, Watanabe K, Tokiwano T, Kinashi H, Oikawa H (2009) Identification of a gene cluster of polyether antibiotic lasalocid from Streptomyces lasaliensis. Biosci Biotechnol Biochem 73:169–176. https://doi.org/10.1271/bbb.80631
Minami A, Ose T, Sato K, Oikawa A, Kuroki K, Maenaka K, Oguri H, Oikawa H (2014) Allosteric regulation of epoxide opening cascades by a pair of epoxide hydrolases in monensin biosynthesis. ACS Chem Biol 9:562–569. https://doi.org/10.1021/cb4006485
Myadoh S (1993) Research on antibiotic screening in Japan over the last decade: a producing microorganism approach. Actinomycetologica 7:100–106. https://doi.org/10.3209/saj.7_100
Noack S, Chapman HD, Selzer PM (2019) Anticoccidial drugs of the livestock industry. Parasitol Res 118:2009–2026. https://doi.org/10.1007/s00436-019-06343-5
Oliynyk M, Stark CB, Bhatt A, Jones MA, Hughes-Thomas ZA, Wilkinson C, Oliynyk Z, Demydchuk Y, Staunton J, Leadlay PF (2003) Analysis of the biosynthetic gene cluster for the polyether antibiotic monensin in Streptomyces cinnamonensis and evidence for the role of monB and monC genes in oxidative cyclization. Mol Microbiol 49:1179–1190. https://doi.org/10.1046/j.1365-2958.2003.03571.x
Paget MS, Chamberlin L, Atrih A, Foster SJ, Buttner MJ (1999) Evidence that the extracytoplasmic function sigma factor sigmaE is required for normal cell wall structure in Streptomyces coelicolor A3(2). J Bacteriol 181:204–211. https://doi.org/10.1128/JB.00622-08
Paudel S, Lee HC, Kim BS, Sohng JK (2011) Enhancement of pradimicin production in Actinomadura hibisca P157-2 by metabolic engineering. Microbiol Res 167:32–39. https://doi.org/10.1016/j.micres.2011.02.007
Raza M, Bharti H, Singal A, Nag A, Ghosh PC (2018) Long circulatory liposomal maduramicin inhibits the growth of Plasmodium falciparum blood stages in culture and cures murine models of experimental malaria. Nanoscale 10:13773–13791. https://doi.org/10.1039/c8nr02442a
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Shichijo Y, Migita A, Oguri H, Watanabe M, Tokiwano T, Watanabe K, Oikawa H (2008) Epoxide hydrolase Lsd19 for polyether formation in the biosynthesis of lasalocid A: direct experimental evidence on polyene-polyepoxide hypothesis in polyether biosynthesis. J Am Chem Soc 130:12230–12231. https://doi.org/10.1021/ja8040543
Shimshoni JA, Britzi M, Pozzi PS, Edery N, Berkowitz A, Bouznach A, Cuneah O, Soback S, Bellaiche M, Younis A, Blech E, Oren P, Galon N, Shlosberg A, Perl S (2014) Acute maduramicin toxicosis in pregnant gilts. Food Chem Toxicol 68:283–289. https://doi.org/10.1016/j.fct.2014.03.034
Shlosberg A, Perl S, Harmelin A, Hanji V, Bellaiche M, Bogin E, Cohen R, Markusfeld-Nir O, Shpigel N, Eisenberg Z, Furman M, Brosh A, Holzer Z, Aharoni Y (1997) Acute maduramicin toxicity in calves. Vet Rec 140:643–646. https://doi.org/10.1136/vr.140.25.643
Singh T, Gupta RP (2003) Clinico-haematological and mineral studies on experimental maduramicin toxicity in chickens. Vet Parasitol 116:345–353. https://doi.org/10.1016/S0304-4017(03)00299-1
Smith L, Hong H, Spencer JB, Leadlay PF (2008) Analysis of specific mutants in the lasalocid gene cluster: evidence for enzymatic catalysis of a disfavoured polyether ring closure. ChemBioChem 9:2967–2975. https://doi.org/10.1002/cbic.200800585
Story P, Doube A (2004) A case of human poisoning by salinomycin, an agricultural antibiotic. N Z Med J 117:U799
Sun Y, Zhou X, Dong H, Tu G, Wang M, Wang B, Deng Z (2003) A complete gene cluster from Streptomyces nanchangensis NS3226 encoding biosynthesis of the polyether ionophore nanchangmycin. Chem Biol 10:431–441. https://doi.org/10.1016/S1074-5521(03)00092-9
Tan GY, Deng K, Liu X, Tao H, Chang Y, Chen J, Chen K, Sheng Z, Deng Z, Liu T (2017) Heterologous biosynthesis of spinosad: an omics-guided large polyketide synthase gene cluster reconstitution in Streptomyces. ACS Synth Biol 6:995–1005. https://doi.org/10.1021/acssynbio.6b00330
Thorpe HM, Wilson SE, Smith MC (2000) Control of directionality in the site-specific recombination system of the Streptomyces phage phiC31. Mol Microbiol 38:232–241. https://doi.org/10.1046/j.1365-2958.2000.02142.x
Tsou H, Rajan S, Fiala R, Mowery PC, Bullock MW, Borders DB, James JC, Martin JH, Morton GO (1984) Biosynthesis of the antibiotic maduramicin. Origin of the carbon and oxygen atoms as well as the 13C NMR assignments. J Antibiot (Tokyo) 37:1651–1663. https://doi.org/10.7164/antibiotics.37.1651
Tsou HR, Rajan S, Chang TT, Fiala RR, Stockton GW, Bullock MW (1987) The utilization of molecular oxygen during the biosynthesis of maduramicin. J Antibiot (Tokyo) 40:94–99. https://doi.org/10.7164/antibiotics.40.94
Voeykova T, Emelyanova L, Tabakov V, Mkrtumyan N (1998) Transfer of plasmid pTO1 from Escherichia coli to various representatives of the order Actinomycetales by intergeneric conjugation. FEMS Microbiol Lett 162:47–52. https://doi.org/10.1111/j.1574-6968.1998.tb12977.x
Wang H, Li Z, Jia R, Yin J, Li A, **a L, Yin Y, Muller R, Fu J, Stewart AF, Zhang Y (2018) ExoCET: exonuclease in vitro assembly combined with RecET recombination for highly efficient direct DNA cloning from complex genomes. Nucleic Acids Res 46:e28. https://doi.org/10.1093/nar/gkx1249
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Muller R, Wohlleben W, Breitling R, Takano E, Medema MH (2015) antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43(1):237–243. https://doi.org/10.1093/nar/gkv437
Wohlert SE, Lomovskaya N, Kulowski K, Fonstein L, Occi JL, Gewain KM, MacNeil DJ, Hutchinson CR (2001) Insights about the biosynthesis of the avermectin deoxysugar l-oleandrose through heterologous expression of Streptomyces avermitilis deoxysugar genes in Streptomyces lividans. Chem Biol 8:681–700. https://doi.org/10.1016/S1074-5521(01)00043-6
Wong FT, Hotta K, Chen X, Fang MY, Watanabe K, Kim CY (2015) Epoxide hydrolase–lasalocid A structure provides mechanistic insight into polyether natural product biosynthesis. J Am Chem Soc 137:86–89. https://doi.org/10.1021/ja511374k
Yakisich JS, Azad N, Kaushik V, O’Doherty GA, Iyer AK (2017) Nigericin decreases the viability of multidrug-resistant cancer cells and lung tumorspheres and potentiates the effects of cardiac glycosides. Tumour B 39:1010428317694310. https://doi.org/10.1177/1010428317694310
Zhang Y, Lin CY, Li XM, Tang ZK, Qiao J, Zhao GR (2016) DasR positively controls monensin production at two-level regulation in Streptomyces cinnamonensis. J Ind Microbiol Biotechnol 43:1681–1692. https://doi.org/10.1007/s10295-016-1845-4
Zhang X, Lu C, Bai L (2017) Mechanism of salinomycin overproduction in Streptomyces albus as revealed by comparative functional genomics. Appl Microbiol Biotechnol 101:4635–4644. https://doi.org/10.1007/s00253-017-8278-5
Zhou Y, Meng Q, You D, Li J, Chen S, Ding D, Zhou X, Zhou H, Bai L, Deng Z (2008) Selective removal of aberrant extender units by a type II thioesterase for efficient FR-008/candicidin biosynthesis in Streptomyces sp. strain FR-008. Appl Environ Microbiol 74:7235–7242. https://doi.org/10.1128/AEM.01012-08
Zhu Z, Li H, Yu P, Guo Y, Luo S, Chen Z, Mao X, Guan W, Li Y (2017) SlnR is a positive pathway-specific regulator for salinomycin biosynthesis in Streptomyces albus. Appl Microbiol Biotechnol 101:1547–1557. https://doi.org/10.1007/s00253-016-7918-5
Acknowledgements
This work was supported by funding from National Key R&D Program of China 2018YFA0900400, Hubei Natural Science Fund Project 2017CFA054, and J1 Biotech Co., Ltd.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, R., Fang, F., An, Z. et al. Genomics-driven discovery of the biosynthetic gene cluster of maduramicin and its overproduction in Actinomadura sp. J1-007. J Ind Microbiol Biotechnol 47, 275–285 (2020). https://doi.org/10.1007/s10295-019-02256-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02256-5