Abstract
Background
Inhibition of vascular endothelial growth factor receptor 2 (VEGFR-2) tyrosine kinase by small molecules has become a promising target in the treatment of cancer.
Objective
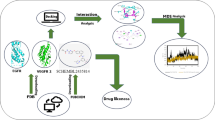
In this study, we approached pharmacophore modeling coupled with a structure-based virtual screening workflow to identify the potent inhibitors.
Methods
The top selected hit compounds have been rescored using the MM/GBSA approach. To understand the molecular reactivity, electronic properties, and stability of those inhibitors, we have employed density functional theory and molecular dynamics. Following that, the best 21 hit compounds have been further post-processed with a Quantum ligand partial charge-based rescoring process and further validated by implementing molecular dynamics simulation.
Results
The ten hit compounds have been hypothesized and considered as potent inhibitors of VEGFR-2 tyrosine kinase. This study also signifies the contribution of QM-based ligand partial charge, which is more accurate in predicting reliable free binding energy and filtering large ligand libraries to hit optimization, rather than assigning those of the force field-based method. From the binding pattern analysis of all the complexes, amino acids, such as Glu885, Cys919, Cys1045, Thr916, Thr919, and Asp1046, were found to have comprehensive interaction with the hit compounds.
Conclusion
Hence, this could prove to be useful as a potential inhibition site of the VEGFR-2 tyrosine kinase domain for future researchers. Moreover, this study also emphasizes the conformational changes upon ATP binding, based on either the receptor’s rigidity or flexibility.












Similar content being viewed by others
Data Availability
The authors confirm that the data supporting the findings of this study are available within the article and/or its supplementary materials.
References
Holmes K, Roberts OL, Thomas AM, Cross MJ (2007) Vascular endothelial growth factor receptor-2: structure, function, intracellular signalling and therapeutic inhibition. Cell Signal 19:2003–2012. https://doi.org/10.1016/j.cellsig.2007.05.013
Kowanetz M, Ferrara N (2006) Vascular endothelial growth factor signaling pathways: therapeutic perspective. Clin Cancer Res 12:5018–5022. https://doi.org/10.1158/1078-0432.CCR-06-1520
Roy H, Bhardwaj S, Ylä-Herttuala S (2006) Biology of vascular endothelial growth factors. FEBS Lett 580:2879–2887. https://doi.org/10.1016/j.febslet.2006.03.087
Shibuya M (2011) Vascular endothelial growth factor (VEGF) and its receptor (VEGFR) signaling in angiogenesis: a crucial target for anti- and pro-angiogenic therapies. Genes Cancer 2:1097–1105. https://doi.org/10.1177/1947601911423031
Cébe-Suarez S, Zehnder-Fjällman A, Ballmer-Hofer K (2006) The role of VEGF receptors in angiogenesis; complex partnerships. Cell Mol Life Sci 63:601. https://doi.org/10.1007/s00018-005-5426-3
Shibuya M (2008) Vascular endothelial growth factor-dependent and -independent regulation of angiogenesis. BMB Rep 41:278–286
Chae S-S, Kamoun WS, Farrar CT et al (2010) Angiopoietin-2 interferes with anti-VEGFR2–induced vessel normalization and survival benefit in mice bearing gliomas. Clin Cancer Res 16:3618–3627. https://doi.org/10.1158/1078-0432.CCR-09-3073
Kanno S, Oda N, Abe M et al (2000) Roles of two VEGF receptors, Flt-1 and KDR, in the signal transduction of VEGF effects in human vascular endothelial cells. Oncogene 19:2138–2146. https://doi.org/10.1038/sj.onc.1203533
Sharma S, Johnson D, Abouammoh M et al (2012) Rate of serious adverse effects in a series of bevacizumab and ranibizumab injections. Can J Ophthalmol 47:275–279. https://doi.org/10.1016/j.jcjo.2012.03.026
Ferrara N (2004) Vascular endothelial growth factor: basic science and clinical progress. Endocr Rev 25:581–611. https://doi.org/10.1210/er.2003-0027
Cook KM, Figg WD (2010) Angiogenesis inhibitors: current strategies and future prospects. CA Cancer J Clin 60:222–243. https://doi.org/10.3322/caac.20075
Weis SM, Cheresh DA (2011) Tumor angiogenesis: molecular pathways and therapeutic targets. Nat Med 17:1359–1370. https://doi.org/10.1038/nm.2537
Li X, Wu X, Zhao P et al (2011) Efficacy and safety of sunitinib in the treatment of metastatic renal cell carcinoma. Chin Med J (Engl) 124:2920
Kiselyov AS, Semenova M, Semenov VV, Piatnitski E (2006) 2-((1H-Azol-1-yl)methyl)-N-arylbenzamides: novel dual inhibitors of VEGFR-1/2 kinases. Bioorg Med Chem Lett 16:1726–1730. https://doi.org/10.1016/j.bmcl.2005.11.105
Borzilleri RM, Bhide RS, Barrish JC et al (2006) Discovery and evaluation of N-cyclopropyl-2,4-difluoro-5-((2-(pyridin-2-ylamino)thiazol-5-ylmethyl)amino)benzamide (BMS-605541), a selective and orally efficacious inhibitor of vascular endothelial growth factor receptor-2. J Med Chem 49:3766–3769. https://doi.org/10.1021/jm060347y
Nakamura H, Sasaki Y, Uno M et al (2006) Synthesis and biological evaluation of benzamides and benzamidines as selective inhibitors of VEGFR tyrosine kinases. Bioorg Med Chem Lett 16:5127–5131. https://doi.org/10.1016/j.bmcl.2006.07.075
Schrödinger Release 2022-2: LigPrep, Schrödinger, LLC, New York, NY, 2021
Shelley JC, Cholleti A, Frye LL et al (2007) Epik: a software program for pKaprediction and protonation state generation for drug-like molecules. J Comput Aided Mol Des 21:681–691. https://doi.org/10.1007/s10822-007-9133-z
Greenwood JR, Calkins D, Sullivan AP, Shelley JC (2010) Towards the comprehensive, rapid, and accurate prediction of the favorable tautomeric states of drug-like molecules in aqueous solution. J Comput Aided Mol Des 24:591–604. https://doi.org/10.1007/s10822-010-9349-1
Harder E, Damm W, Maple J et al (2016) OPLS3: a force field providing road coverage of drug-like small molecules and proteins. J Chem Theory Comput 12:281–296. https://doi.org/10.1021/acs.jctc.5b00864
Berman H, Henrick K, Nakamura H (2003) Announcing the worldwide Protein Data Bank. Nat Struct Mol Biol 10:980. https://doi.org/10.1038/nsb1203-980
Madhavi Sastry G, Adzhigirey M, Day T et al (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27:221–234. https://doi.org/10.1007/s10822-013-9644-8
Halgren TA, Murphy RB, Friesner RA et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem 47:1750–1759. https://doi.org/10.1021/jm030644s
Friesner RA, Murphy RB, Repasky MP et al (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein−ligand complexes. J Med Chem 49:6177–6196. https://doi.org/10.1021/jm051256o
Schrödinger Release 2021-4: QikProp, Schrödinger, LLC, New York, NY, 2021
Schrödinger Release 2022-2: Prime, Schrödinger, LLC, New York, NY, 2021
Li J, Abel R, Zhu K et al (2011) The VSGB 2.0 model: a next generation energy model for high resolution protein structure modeling. Proteins Struct, Funct, Bioinform 79:2794–2812. https://doi.org/10.1002/prot.23106
Matysiak J (2007) Evaluation of electronic, lipophilic and membrane affinity effects on antiproliferative activity of 5-substituted-2-(2,4-dihydroxyphenyl)-1,3,4-thiadiazoles against various human cancer cells. Eur J Med Chem 42:940–947. https://doi.org/10.1016/j.ejmech.2006.12.033
Baseden KA, Tye JW (2014) Introduction to density functional theory: calculations by hand on the helium atom. J Chem Educ 91:2116–2123. https://doi.org/10.1021/ed5004788
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652. https://doi.org/10.1063/1.464913
Gill PMW, Johnson BG, Pople JA, Frisch MJ (1992) The performance of the Becke—Lee—Yang—Parr (B—LYP) density functional theory with various basis sets. Chem Phys Lett 197:499–505. https://doi.org/10.1016/0009-2614(92)85807-M
Stephens PJ, Devlin FJ, Chabalowski CF, Frisch MJ (1994) Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J Phys Chem 98:11623–11627. https://doi.org/10.1021/j100096a001
Easton RE, Giesen DJ, Welch A et al (1996) The MIDI! basis set for quantum mechanical calculations of molecular geometries and partial charges. Theor Chim Acta 93:281–301
Bochevarov AD, Harder E, Hughes TF et al (2013) Jaguar: a high-performance quantum chemistry software program with strengths in life and materials sciences. Int J Quantum Chem 113:2110–2142. https://doi.org/10.1002/qua.24481
Krieger E, Vriend G (2015) New ways to boost molecular dynamics simulations. J Comput Chem 36:996–1007. https://doi.org/10.1002/jcc.23899
Wang J, Wolf RM, Caldwell JW et al (2004) Development and testing of a general amber force field. J Comput Chem 25:1157–1174. https://doi.org/10.1002/jcc.20035
Sprenger KG, Jaeger VW, Pfaendtner J (2015) The general AMBER force field (GAFF) can accurately predict thermodynamic and transport properties of many ionic liquids. J Phys Chem B 119:5882–5895. https://doi.org/10.1021/acs.jpcb.5b00689
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. https://doi.org/10.1063/1.464397
Plimpton SJ, Pollock R, Stevens MJ (1997) Particle-mesh Ewald and rRESPA for parallel molecular dynamics simulations. Proc 8th SIAM Conf on Parallel Processing for Scientific Computing
Zhan C-G, Nichols JA, Dixon DA (2003) Ionization potential, electron affinity, electronegativity, hardness, and electron excitation energy: molecular properties from density functional theory orbital energies. J Phys Chem A 107:4184–4195. https://doi.org/10.1021/jp0225774
Parr RG, Zhou Z (1993) Absolute hardness: unifying concept for identifying shells and subshells in nuclei, atoms, molecules, and metallic clusters. Acc Chem Res 26:256–258. https://doi.org/10.1021/ar00029a005
Hoque MM, Halim MA, Sarwar MG, Khan MdW (2015) Palladium-catalyzed cyclization of 2-alkynyl-N-ethanoyl anilines to indoles: synthesis, structural, spectroscopic, and mechanistic study. J Phys Org Chem 28:732–742. https://doi.org/10.1002/poc.3477
Ehrlich P (1909) Über den jetzigen Stand der Chemotherapie. Ber Dtsch Chem Ges 42:17–47. https://doi.org/10.1002/cber.19090420105
Schueler FW (1961) Chemobiodynamics and drug design. Acad Med 36:285–286
Hu E, Tasker A, White RD et al (2008) Discovery of aryl aminoquinazoline pyridones as potent, selective, and orally efficacious inhibitors of receptor tyrosine kinase c-Kit. J Med Chem 51:3065–3068. https://doi.org/10.1021/jm800188g
McTigue M, Murray BW, Chen JH et al (2012) Molecular conformations, interactions, and properties associated with drug efficiency and clinical performance among VEGFR TK inhibitors. Proc Natl Acad Sci 109:18281–18289
Li J, Zhou N, Luo K et al (2014) In silico discovery of potential VEGFR-2 inhibitors from natural derivatives for anti-angiogenesis therapy. Int J Mol Sci 15:15994–16011
Weiss MM, Harmange J-C, Polverino AJ et al (2008) Evaluation of a series of naphthamides as potent, orally active vascular endothelial growth factor receptor-2 tyrosine kinase inhibitors. J Med Chem 51:1668–1680. https://doi.org/10.1021/jm701098w
Potashman MH, Bready J, Coxon A et al (2007) Design, synthesis, and evaluation of orally active benzimidazoles and benzoxazoles as vascular endothelial growth factor-2 receptor tyrosine kinase inhibitors. J Med Chem 50:4351–4373. https://doi.org/10.1021/jm070034i
Papakyriakou A, Kefalos P, Sarantis P et al (2014) A zebrafish in vivo phenotypic assay to identify 3-aminothiophene-2-carboxylic acid-based angiogenesis inhibitors. Assay Drug Dev Technol 12:527–535. https://doi.org/10.1089/adt.2014.606
Morozov AV, Kortemme T, Tsemekhman K, Baker D (2004) Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations. Proc Natl Acad Sci 101:6946–6951. https://doi.org/10.1073/pnas.0307578101
Kangas E, Tidor B (1998) Optimizing electrostatic affinity in ligand–receptor binding: theory, computation, and ligand properties. J Chem Phys 109:7522–7545
Ryde U, Söderhjelm P (2016) Ligand-binding affinity estimates supported by quantum-mechanical methods. Chem Rev 116:5520–5566. https://doi.org/10.1021/acs.chemrev.5b00630
Ekins S, Waller CL, Swaan PW et al (2000) Progress in predicting human ADME parameters in silico. J Pharmacol Toxicol Methods 44:251–272. https://doi.org/10.1016/S1056-8719(00)00109-X
Selick HE, Beresford AP, Tarbit MH (2002) The emerging importance of predictive ADME simulation in drug discovery. Drug Discov Today 7:109–116. https://doi.org/10.1016/S1359-6446(01)02100-6
Yu H, Adedoyin A (2003) ADME-Tox in drug discovery: integration of experimental and computational technologies. Drug Discov Today 8:852–861. https://doi.org/10.1016/s1359-6446(03)02828-9
Ioakimidis L, Thoukydidis L, Mirza A et al (2008) Benchmarking the reliability of QikProp. Correlation between experimental and predicted values. QSAR Comb Sci 27:445–456. https://doi.org/10.1002/qsar.200730051
Mi Lobanov, Bogatyreva NS, Galzitskaia OV (2008) Radius of gyration is indicator of compactness of protein structure. Mol Biol 42:701–706
Acknowledgements
The authors would like to acknowledge Shafi Mahmud for facilitating YASARA software (V.20.08.10 License: 1965847**) for simulating protein-ligand complexes.
Author information
Authors and Affiliations
Contributions
1) MRP and YMR conceived and designed the experiments.
2) Shafi Mahmud performed the MD simulation.
3) YMR, SA, MRP, and SJ analyzed and interpreted the data.
4) YMR, MRP, and SA wrote the paper.
Corresponding author
Ethics declarations
Ethics approval
None.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parves, M.R., Riza, Y.M., Alam, S. et al. Molecular dynamics-based insight of VEGFR-2 kinase domain: a combined study of pharmacophore modeling and molecular docking and dynamics. J Mol Model 29, 17 (2023). https://doi.org/10.1007/s00894-022-05427-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-022-05427-x