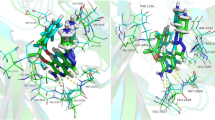
Abstract
Crizotinib is an anticancer tyrosine kinase inhibitor that is approved for use as a first-line treatment for some non-small-cell lung cancers. L1196M is the most frequently observed mutation in NSCLC patients. This mutation, known as the gatekeeper mutation in the ALK kinase domain, confers resistance to crizotinib by sterically blocking the binding of the drug. However, the molecular mechanism of crizotinib resistance caused by the L1196M mutation is still unclear. Molecular dynamics simulation was therefore utilized in this study to investigate the mechanism by which the L1196M mutation may affect crizotinib resistance. Our results suggest that larger fluctuations in some important regions of the mutant complex compared to the wild-type complex may contribute to the resistance of the mutant complex to crizotinib. Also, mutation-induced alterations to the secondary structure of the complex as well as unstable hydrogen-bonding patterns in the A-loop and P-loop regions decrease the total binding energy of the complex. This study therefore provides a molecular explanation for the resistance to crizotinib caused by the L1196M mutation, which could aid the design of more efficient and selective drugs.




Similar content being viewed by others
Abbreviations
- RTKs:
-
Receptor tyrosine kinase
- NSCLC:
-
Non-small-cell lung cancer
- EML4:
-
Echinoderm microtubule-associated protein-like 4
- FDA:
-
Food and Drug Administration
- GAFF:
-
Generalized AMBER force field
- SD:
-
Steepest descent
- CG:
-
Conjugated gradient
- PME:
-
Particle mesh Ewald
- RMSD:
-
Root-mean-square deviation
- RMSF:
-
Root-mean-square fluctuation
- ALK:
-
Anaplastic lymphoma kinase
References
Casaluce F, Sgambato A, Claudia Sacco P, Palazzolo G, Maione P, Rossi A, Ciardiello F, Gridelli C (2016) Resistance to crizotinib in advanced non-small cell lung cancer (NSCLC) with ALK rearrangement: mechanisms, treatment strategies and new targeted therapies. Curr Clin Pharmacol 11(2):77–87
Katayama R, Lovly CM, Shaw AT (2015) Therapeutic targeting of anaplastic lymphoma kinase in lung cancer: a paradigm for precision cancer medicine. Clin Cancer Res 21(10):2227–2235
Huang Q, Johnson TW, Bailey S, Brooun A, Bunker KD, Burke BJ, Collins MR, Cook AS, Cui JJ, Dack KN (2014) Design of potent and selective inhibitors to overcome clinical anaplastic lymphoma kinase mutations resistant to crizotinib. J Med Chem 57(4):1170–1187
Shaw AT, Yeap BY, Solomon BJ, Riely GJ, Gainor J, Engelman JA, Shapiro GI, Costa DB, Ou S-HI, Butaney M (2011) Effect of crizotinib on overall survival in patients with advanced non-small-cell lung cancer harbouring ALK gene rearrangement: a retrospective analysis. Lancet Oncol 12(11):1004–1012
Choi YL, Soda M, Yamashita Y, Ueno T, Takashima J, Nakajima T, Yatabe Y, Takeuchi K, Hamada T, Haruta H (2010) EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N Engl J Med 363(18):1734–1739
Doebele RC, Pilling AB, Aisner DL, Kutateladze TG, Le AT, Weickhardt AJ, Kondo KL, Linderman DJ, Heasley LE, Franklin WA (2012) Mechanisms of resistance to crizotinib in patients with ALK gene rearranged non-small cell lung cancer. Clin Cancer Res 18(5):1472–1482
Priya Doss CG, Chakraborty C, Chen L, Zhu H (2014) Integrating in silico prediction methods, molecular docking, and molecular dynamics simulation to predict the impact of ALK missense mutations in structural perspective. BioMed Res Int 895831
Sun H-Y, Ji F-Q (2012) A molecular dynamics investigation on the crizotinib resistance mechanism of C1156Y mutation in ALK. Biochem Biophys Res Commun 423(2):319–324
Gainor JF, Dardaei L, Yoda S, Friboulet L, Leshchiner I, Katayama R, Dagogo-Jack I, Gadgeel S, Schultz K, Singh M (2016) Molecular mechanisms of resistance to first- and second-generation ALK inhibitors in ALK-rearranged lung cancer. Cancer Discovery CD-16-0596
Dehghanian F, Kay M, Vallian S (2017) F1174V mutation alters the ALK active conformation in response to crizotinib in NSCLC: insight from molecular simulations. J Mol Graph Model 75:287–293
Webb B, Sali A (2016) Comparative protein structure modeling using Modeller. Curr Protoc Bioinformatics 5.6.1–5.6.37
Thomsen R, Christensen MH (2006) MolDock: a new technique for high-accuracy molecular docking. J Med Chem 49(11):3315–3321
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65(3):712–725
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33(12):889–897
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J Comput Chem 23(16):1623–1641
Wang J, Wang W, Kollman PA, Case DA (2006) Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model 25(2):247–260
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935
Case DA, Cheatham TE, Darden T, Gohlke H, Luo R, Merz KM, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26(16):1668–1688
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092
Ryckaert J-P, Ciccotti G, Berendsen HJ (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22(12):2577–2637
Li L, Uversky VN, Dunker AK, Meroueh SO (2007) A computational investigation of allostery in the catabolite activator protein. J Am Chem Soc 129(50):15668–15676
Stoica I, Sadiq SK, Coveney PV (2008) Rapid and accurate prediction of binding free energies for saquinavir-bound HIV-1 proteases. J Am Chem Soc 130(8):2639–2648
Kongsted J, Ryde U (2009) An improved method to predict the entropy term with the MM/PBSA approach. J Comput Aided Mol Des 23(2):63–71
Ai X, Shen S, Shen L, Lu S (2015) An interaction map of small-molecule kinase inhibitors with anaplastic lymphoma kinase (ALK) mutants in ALK-positive non-small cell lung cancer. Biochimie 112:111–120
Azarova AM, Gautam G, George RE (2011) Emerging importance of ALK in neuroblastoma. Semin Cancer Biol 21:267–75
Kumar A, Ramanathan K (2015) Analyzing resistance pattern of non-small cell lung cancer to crizotinib using molecular dynamic approaches. Indian J Biochem Biophys 52:23–28
Lu H, Tonge PJ (2010) Drug–target residence time: critical information for lead optimization. Curr Opin Chem Biol 14(4):467–474
Tummino PJ, Copeland RA (2008) Residence time of receptor–ligand complexes and its effect on biological function. Biochemistry 47(20):5481–5492
Kumar A, Ramanathan K (2014) Exploring the structural and functional impact of the ALK F1174L mutation using bioinformatics approach. J Mol Model 20(7):1–7
Kumar A, Shanthi V, Ramanathan K (2015) Computational investigation and experimental validation of crizotinib resistance conferred by C1156Y mutant anaplastic lymphoma kinase. Mol Inf 34(2–3):105–114
Chen J, Zhang S, Liu X, Zhang Q (2010) Insights into drug resistance of mutations D30N and I50V to HIV-1 protease inhibitor TMC-114: free energy calculation and molecular dynamic simulation. J Mol Model 16(3):459–468
Leonis G, Steinbrecher T, Papadopoulos MG (2013) A contribution to the drug resistance mechanism of darunavir, amprenavir, indinavir, and saquinavir complexes with HIV-1 protease due to flap mutation I50V: a systematic MM-PBSA and thermodynamic integration study. J Chem Inf Model 53(8):2141–2153
Sun H, Li Y, Li D, Hou T (2013) Insight into crizotinib resistance mechanisms caused by three mutations in ALK tyrosine kinase using free energy calculation approaches. J Chem Inf Model 53(9):2376–2389
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
There is no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 330 kb)
Rights and permissions
About this article
Cite this article
Kay, M., Dehghanian, F. Exploring the crizotinib resistance mechanism of NSCLC with the L1196M mutation using molecular dynamics simulation. J Mol Model 23, 323 (2017). https://doi.org/10.1007/s00894-017-3495-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-017-3495-5