Abstract
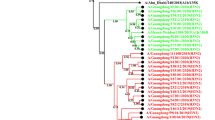
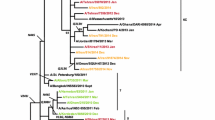
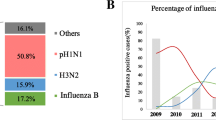
We investigated and analysed the molecular evolution of hemagglutinin (HA) and neuraminidase (NA) of influenza A(H1N1)pdm09 virus during the 2017–2018 and 2018–2019 influenza seasons in Bei**g, China. We collected and extracted RNA from influenza A(H1N1)pdm09 strains from Peking University People’s Hospital and analyzed their HA and NA genes by RT-PCR and sequencing. Phylogenetic analysis of HA and NA sequences was used to compare the amino acid sequences of 51 strains with those of reference strains. All strains belonged to subclade 6B.1, with S162N and I216T substitutions (H1 numbering). Our strains differed from strain A/Michigan/45/2015, with the substitutions S91R, S181T and I312V in the HA antigenic epitope. An E189G mutation was detected in the 190 helix of the receptor binding region of HA. A new potential glycosylation site, 179 (NQT), which was not detected before the 2015 influenza season, was identified. Two strains were mutated at I223, the NA inhibitor resistance site. During 2012-2019, amino acids of HA and NA mutated over time. Co-occurrence mutations N146D, S200P, S202I and A273T in HA appeared along with Q51K, F74S and D416N in NA in six strains during two influenza seasons. Our work reveals the molecular changes and phylogenetic characteristics of influenza A(H1N1)pdm09 virus and suggests that a vaccine probably provides suboptimal protection. The biological characteristics of the new glycosylation and drug-resistance sites detected in this work need to be studied further. The co-occurrence of mutations in HA and NA might affect the characteristics of the virus and need to be given more attention.



Similar content being viewed by others
References
World Health Organization (2016) Recommended composition of influenza virus vaccines for use in the 2016–2017 northern hemisphere influenza season. Wkly Epidemiol Rec 91:121–132
World Health Organization (2018) Recommended composition of influenza virus vaccines for use in the 2018–2019 northern hemisphere influenza season. Wkly Epidemiol Rec 93:133–141
Abed Y, Pizzorno A, Bouhy X, Rheaume C, Boivin G (2014) Impact of potential permissive neuraminidase mutations on viral fitness of the H275Y oseltamivir-resistant influenza A(H1N1)pdm09 virus in vitro, in mice and in ferrets. J Virol 88:1652–1658
Air GM (2012) Influenza neuraminidase. Influ Other Respir Viruses 6:245–256
Al Khatib HA, Al Thani AA, Gallouzi I, Yassine HM (2019) Epidemiological and genetic characterization of pH1N1 and H3N2 influenza viruses circulated in MENA region during 2009–2017. BMC Infect Dis 19:314
Barbezange C, Jones L, Blanc H, Isakov O, Celniker G, Enouf V, Shomron N, Vignuzzi M, van der Werf S (2018) Seasonal genetic drift of human influenza A virus quasispecies revealed by deep sequencing. Front Microbiol 9:2596
Chambers C, Skowronski DM, Sabaiduc S, Winter AL, Dickinson JA, De Serres G, Gubbay JB, Drews SJ, Martineau C, Eshaghi A, Krajden M, Bastien N, Li Y (2016) Interim estimates of 2015/16 vaccine effectiveness against influenza A(H1N1)pdm09, Canada, February 2016. Euro Surveill 21:30168
Chan PK, Lee N, Joynt GM, Choi KW, Cheung JL, Yeung AC, Lam P, Wong R, Leung BW, So HY, Lam WY, Hui DC (2011) Clinical and virological course of infection with haemagglutinin D222G mutant strain of 2009 pandemic influenza A (H1N1) virus. J Clin Virol 50:320–324
Fang Q, Gao Y, Chen M, Guo X, Yang X, Yang X, Wei L (2014) Molecular epidemiology and evolution of A(H1N1)pdm09 and H3N2 virus during winter 2012–2013 in Bei**g, China. Infect Genet Evol 26:228–240
Fang Q, Gao Y, Chen M, Guo X, Yang X, Wei L (2015) Molecular epidemiology and evolution of influenza A and B viruses during winter 2013–2014 in Bei**g, China. J Arch Virol 160:1083–1095
Fitzner J, Qasmieh S, Mounts AW, Alexander B, Besselaar T, Briand S, Brown C, Clark S, Dueger E, Gross D, Hauge S, Hirve S, Jorgensen P, Katz MA, Mafi A, Malik M, McCarron M, Meerhoff T, Mori Y, Mott J, Olivera M, Ortiz JR, Palekar R, Rebelo-de-Andrade H, Soetens L, Yahaya AA, Zhang W, Vandemaele K (2018) Revision of clinical case definitions: influenza-like illness and severe acute respiratory infection. Bull World Health Organ 96:122–128
Garcia V, Aris-Brosou S (2014) Comparative dynamics and distribution of influenza drug resistance acquisition to protein m2 and neuraminidase inhibitors. Mol Biol Evol 31:355–363
Goka EA, Vallely PJ, Mutton KJ, Klapper PE (2014) Mutations associated with severity of the pandemic influenza A(H1N1)pdm09 in humans: a systematic review and meta-analysis of epidemiological evidence. Arch Virol 159:3167–3183
Guarnaccia T, Carolan LA, Maurer-Stroh S, Lee RT, Job E, Reading PC, Petrie S, McCaw JM, McVernon J, Hurt AC, Kelso A, Mosse J, Barr IG, Laurie KL (2013) Antigenic drift of the pandemic 2009 A(H1N1) influenza virus in A ferret model. PLoS Pathog 9:e1003354
Hussain M, Galvin HD, Haw TY, Nutsford AN, Husain M (2017) Drug resistance in influenza A virus: the epidemiology and management. Infect Drug Resist 10:121–134
Jones S, Nelson-Sathi S, Wang Y, Prasad R, Rayen S, Nandel V, Hu Y, Zhang W, Nair R, Dharmaseelan S, Chirundodh DV, Kumar R, Pillai RM (2019) Evolutionary, genetic, structural characterization and its functional implications for the influenza A (H1N1) infection outbreak in India from 2009 to 2017. Sci Rep 9:14690
Khandaker I, Suzuki A, Kamigaki T, Tohma K, Odagiri T, Okada T, Ohno A, Otani K, Sawayama R, Kawamura K, Okamoto M, Oshitani H (2013) Molecular evolution of the hemagglutinin and neuraminidase genes of pandemic (H1N1) 2009 influenza viruses in Sendai, Japan, during 2009–2011. Virus Genes 21(11):30168
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
L’Huillier AG, Abed Y, Petty TJ, Cordey S, Thomas Y, Bouhy X, Schibler M, Simon A, Chalandon Y, van Delden C, Zdobnov E, Boquete-Suter P, Boivin G, Kaiser L (2015) E119D neuraminidase mutation conferring pan-resistance to neuraminidase inhibitors in an A(H1N1)pdm09 isolate from a stem-cell transplant recipient. J Infect Dis 212:1726–1734
Medina RA, Garcia-Sastre A (2011) Influenza A viruses: new research developments. Nat Rev Microbiol 9:590–603
Mesman AW, Westerhuis BM, Ten Hulscher HI, Jacobi RH, de Bruin E, van Beek J, Buisman AM, Koopmans MP, van Binnendijk RS (2016) Influenza virus A(H1N1)2009 antibody-dependent cellular cytotoxicity in young children prior to the H1N1 pandemic. J Gen Virol 97:2157–2165
Morlighem JE, Aoki S, Kishima M, Hanami M, Ogawa C, Jalloh A, Takahashi Y, Kawai Y, Saga S, Hayashi E, Ban T, Izumi S, Wada A, Mano M, Fukunaga M, Kijima Y, Shiomi M, Inoue K, Hata T, Koretsune Y, Kudo K, Himeno Y, Hirai A, Takahashi K, Sakai-Tagawa Y, Iwatsuki-Horimoto K, Kawaoka Y, Hayashizaki Y, Ishikawa T (2011) Mutation analysis of 2009 pandemic influenza A(H1N1) viruses collected in Japan during the peak phase of the pandemic. PLoS One 6:e18956
Neumann G, Noda T, Kawaoka Y (2009) Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature 459:931–939
Nguyen HT, Fry AM, Gubareva LV (2012) Neuraminidase inhibitor resistance in influenza viruses and laboratory testing methods. Antivir Ther 17:159–173
Pizzorno A, Abed Y, Bouhy X, Beaulieu E, Mallett C, Russell R, Boivin G (2012) Impact of mutations at residue I223 of the neuraminidase protein on the resistance profile, replication level, and virulence of the 2009 pandemic influenza virus. Antimicrob Agents Chemother 56:1208–1214
Russell RJ, Haire LF, Stevens DJ, Collins PJ, Lin YP, Blackburn GM, Hay AJ, Gamblin SJ, Skehel JJ (2006) The structure of H5N1 avian influenza neuraminidase suggests new opportunities for drug design. Nature 443:45–49
Shao W, Li X, Goraya MU, Wang S, Chen JL (2017) Evolution of influenza A virus by mutation and re-assortment. Int J Mol Sci 18(8):1650
Tamura D, DeBiasi RL, Okomo-Adhiambo M, Mishin VP, Campbell AP, Loechelt B, Wiedermann BL, Fry AM, Gubareva LV (2015) Emergence of multidrug-resistant influenza A(H1N1)pdm09 virus variants in an immunocompromised child treated with oseltamivir and zanamivir. J Infect Dis 212:1209–1213
Tamura K (1992) Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol Biol Evol 9:678–687
Trebbien R, Pedersen SS, Vorborg K, Franck KT, Fischer TK (2017) Development of oseltamivir and zanamivir resistance in influenza A(H1N1)pdm09 virus, Denmark, 2014. Euro Surveill 22(3):30445
Wang Q, Yue N, Zheng M, Wang D, Duan C, Yu X, Zhang X, Bao C, ** H (2018) Influenza vaccination coverage of population and the factors influencing influenza vaccination in mainland China: a meta-analysis. Vaccine 36:7262–7269
Webster RG, Rott R (1987) Influenza virus A pathogenicity: the pivotal role of hemagglutinin. Cell 50:665–666
Williams JA, Gui L, Hom N, Mileant A, Lee KK (2018) Dissection of epitope-specific mechanisms of neutralization of influenza virus by intact IgG and Fab fragments. J Virol 92(6):e02006-17
Woods CJ, Malaisree M, Pattarapongdilok N, Sompornpisut P, Hannongbua S, Mulholland AJ (2012) Long time scale GPU dynamics reveal the mechanism of drug resistance of the dual mutant I223R/H275Y neuraminidase from H1N1-2009 influenza virus. Biochemistry 51:4364–4375
Zhou B, Deng YM, Barnes JR, Sessions OM, Chou TW, Wilson M, Stark TJ, Volk M, Spirason N, Halpin RA, Kamaraj US, Ding T, Stockwell TB, Salvatore M, Ghedin E, Barr IG, Wentworth DE (2017) Multiplex reverse transcription-PCR for simultaneous surveillance of influenza A and B viruses. J Clin Microbiol 55:3492–3501
Acknowledgements
We thank all the doctors in the Department of Infectious Disease of PKUPH for recruiting patients and collecting nasal specimens. We also thank the laboratory staff in Peking University Hepatology Institute of PKUPH for their guidance.
Funding
This study was supported by the National Natural Science Foundation of China (no. 81541139). The funding sources were not involved.
Author information
Authors and Affiliations
Contributions
Conceptualization, methodology, and supervision were performed by Yan Gao. Material preparation, data collection, and analysis were performed by Baiyi Liu. Baiyi Liu, Yue Wang, Yafen Liu, Yisi Liu, and Yuanyuan Chen collected nasal specimens for this research. Xu Cong and Ying Ji provided help with experimental techniques. The original draft of the manuscript was written by Baiyi Liu, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethics approval
Informed consent was obtained from all patients before collecting nasal samples. This study followed the Declaration of Helsinki Principles, and ethical approval for the study was obtained from the Research Ethics Committee at PKUPH (IRB No. 2016PHB100-01). All analysed data were anonymized.
Additional information
Handling Editor: Ayato Takada.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, B., Wang, Y., Liu, Y. et al. Molecular evolution and characterization of hemagglutinin and neuraminidase of influenza A(H1N1)pdm09 viruses isolated in Bei**g, China, during the 2017–2018 and 2018–2019 influenza seasons. Arch Virol 166, 179–189 (2021). https://doi.org/10.1007/s00705-020-04869-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04869-z