Abstract
Background
Hirschsprung’s disease (HSCR) is a congenital disorder resulting from abnormal development of the enteric nervous system (ENS). Given the complexity of its pathogenesis, it is important to investigate the role of epigenetic inheritance in its development. As Circ-MTCL1 is abundant in brain tissue and colon tissue, whether it has a significant part in the development of ENS is worth exploring. This study clarifies its role in HSCR and identifies the specific molecular mechanisms involved.
Methods
Diseased and dilated segment colon tissues diagnosed as HSCR were collected for the assessment of gene expression levels using RT-PCR. EdU and CCK-8 assays were adopted to evaluate cell proliferation, and Transwell assay was adopted to assess cell migration. The interaction between Circ-MTCL1, miR-145-5p and SMAD3 was confirmed by dual luciferase reporter gene analysis, RT-PCR and Western blotting.
Results
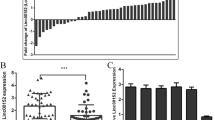
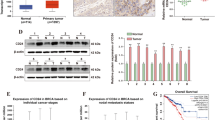
Circ-MTCL1 was down-regulated in the aganglionic colon tissues. The decreased expression of Circ-MTCL1 associated with a reduction in cell migration and proliferation. Bioinformatics analysis and cellular experiments confirmed its role might have been associated with the inhibition of miR-145-5p. MiR-145-5p was up-regulated in HSCR diseased segment colon tissues, exhibiting a negative correlation with Circ-MTCL1. Overexpression of miR-145-5p reversed the inhibition of cell migration and proliferation associated with Circ-MTCL1 down-regulation. The expression of SMAD3 was inhibited by miR-145-5p. The overexpression of SMAD3 eliminated the miR-145-5p-associated inhibition of cell migration and proliferation. Overexpression of miR-145-5p reversed the inhibitory effects of Circ-MTCL1 down-regulation-associated inhibition of cell migration and proliferation, while suppressing SMAD3 expression. Conversely, overexpression of SMAD3 counteracted the miR-145-5p-associated inhibition of cell migration and proliferation.
Conclusions
Circ-MTCL1 may function as a miR-145-5p sponge, regulating the expression of SMAD3 and influencing cell migration and proliferation, thus participating in the development of HSCR.






Similar content being viewed by others
References
Butler TN, Trainor PA (2013) The developmental etiology and pathogenesis of Hirschsprung Disease. Transl Res 162(1):1–15
Lake JI, Heuckeroth RO (2013) Enteric nervous system development: migration, differentiation, and Disease. Am J Physiol Gastrointest Liver Physiol 305(1):G1–24
Soret R et al (2020) Glial cell-derived neurotrophic factor induces enteric neurogenesis and improves Colon structure and function in mouse models of Hirschsprung Disease. Gastroenterology 159(5):1824–1838e17
Belknap WM (2003) Hirschsprung’s Disease. Curr Treat Options Gastroenterol 6(3):247–256
Schriemer D et al (2016) Regulators of gene expression in enteric neural crest cells are putative Hirschsprung Disease genes. Dev Biol 416(1):255–265
Widowati T et al (2016) RET and EDNRB mutation screening in patients with Hirschsprung Disease: functional studies and its implications for genetic counseling. Eur J Hum Genet 24(6):823–829
Musser MA, Correa H, Southard-Smith EM (2015) Enteric neuron imbalance and proximal dysmotility in ganglionated intestine of the Sox10(Dom/+) Hirschsprung mouse model. Cell Mol Gastroenterol Hepatol 1(1):87–101
Huang J et al (2016) Genetic variation in the GDNF promoter affects its expression and modifies the severity of Hirschsprung’s Disease (HSCR) in rats carrying Ednrb(Sl) mutations. Gene 575(1):144–148
Huang Z et al (2023) The m(6)a methyltransferase METTL3 affects cell proliferation and migration by regulating YAP expression in Hirschsprung Disease. Pediatr Surg Int 39(1):126
Zheng H et al (2023) Downregulation of miR-144 blocked the proliferation and invasion of nerve cells in Hirschsprung Disease by regulating Transcription Factor AP 4 (TFAP4). Pediatr Surg Int 39(1):251
Kristensen LS et al (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20(11):675–691
Peng L et al (2017) Circular RNA ZNF609 functions as a competitive endogenous RNA to regulate AKT3 expression by sponging mir-150-5p in Hirschsprung’s Disease. Oncotarget 8(1):808–818
Wen Z et al (2019) Circular RNA CCDC66 targets DCX to regulate cell proliferation and migration by sponging mir-488-3p in Hirschsprung’s Disease. J Cell Physiol 234(7):10576–10587
Zhou L et al (2018) Down-regulation of circ-PRKCI inhibits cell migration and proliferation in Hirschsprung Disease by suppressing the expression of miR-1324 target PLCB1. Cell Cycle 17(9):1092–1101
Wang Z et al (2022) Circular RNA MTCL1 promotes advanced laryngeal squamous cell carcinoma progression by inhibiting C1QBP ubiquitin degradation and mediating beta-catenin activation. Mol Cancer 21(1):92
Li JH et al (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42(Database issue):D92–D97
Beermann J et al (2016) Non-coding RNAs in Development and Disease: background, mechanisms, and therapeutic approaches. Physiol Rev 96(4):1297–1325
Wu W, Zhao F, Zhang J (2023) circAtlas 3.0: a gateway to 3 million curated vertebrate circular RNAs based on a standardized nomenclature scheme. Nucleic Acids Res,
Panda AC et al (2018) Analysis of circular RNAs using the web Tool CircInteractome. Methods Mol Biol 1724:43–56
Liu M et al (2019) Circbank: a comprehensive database for circRNA with standard nomenclature. RNA Biol 16(7):899–905
Agarwal V et al (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife, 4
Chen Y, Wang X (2020) miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res 48(D1):D127–D131
Dweep H, Gretz N, Sticht C (2014) miRWalk database for miRNA-target interactions. Methods Mol Biol 1182:289–305
Vejnar CE, Blum M, Zdobnov EM (2013) miRmap web: Comprehensive microRNA target prediction online. Nucleic Acids Res, 41(Web Server issue): p. W165–W168
Szklarczyk D et al (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 51(D1):D638–D646
Tang X et al (2020) HDAC8 cooperates with SMAD3/4 complex to suppress SIRT7 and promote cell survival and migration. Nucleic Acids Res 48(6):2912–2923
Li Z et al (2020) The emerging landscape of circular RNAs in immunity: breakthroughs and challenges. Biomark Res 8:25
Guria A et al (2019) Circular RNAs-The Road Less traveled. Front Mol Biosci 6:146
**a RP et al (2022) Circ-ITCH overexpression promoted cell proliferation and migration in Hirschsprung Disease through miR-146b-5p/RET axis. Pediatr Res 92(4):1008–1016
You X et al (2015) Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci 18(4):603–610
Zhang Q et al (2022) Mir-145-5p inhibits the proliferation of glioma stem cells by targeting translationally controlled Tumor protein. J Cancer 13(5):1490–1500
Wang Y et al (2021) Umbilical mesenchymal stem cell-derived exosomes facilitate spinal cord functional recovery through the miR-199a-3p/145-5p-mediated NGF/TrkA signaling pathway in rats. Stem Cell Res Ther 12(1):117
Zhao F et al (2019) [Effect of enhancer of zeste homolog 2 on the expression of glial cell line-derived neurotrophic factor family receptor α-1 in the colon tissue of children with Hirschsprung’s Disease]. Zhongguo Dang Dai Er Ke Za Zhi 21(10):1033–1037
Attisano L, Lee-Hoeflich ST (2001) The Smads Genome Biol 2(8):REVIEWS3010
Xu W et al (2017) MiR-1 suppresses Tumor cell proliferation in Colorectal cancer by inhibition of Smad3-mediated Tumor glycolysis. Cell Death Dis 8(5):e2761
Tang PC et al (2022) Single-cell RNA sequencing uncovers a neuron-like macrophage subset associated with cancer pain. Sci Adv 8(40):eabn5535
Huang C et al (2022) EZH2-triggered methylation of SMAD3 promotes its activation and Tumor Metastasis. J Clin Invest, 132(5)
Acknowledgements
This study was supported by the Guangdong Basic and Applied Basic Research Foundation of China (2019A1515011086).
Author information
Authors and Affiliations
Contributions
W.C., L.C.: preparation manuscript, statistics, and writing the manuscript. Y.Y.: performing the experiments. H.X., L.N., Y.J.: literature search, literature analyses. Y.H.: reviewed the manuscript. W.K. and Y.L.: guide the project, and revise the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethical statement
The studies involving human participants were reviewed and approved by Ethics Committee of Zhujiang Hospital of Southern Medical University (2023-KY-117-02). Written informed consent was obtained from patients’ parents for the publication of any potentially identifiable images or data included in this article.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, W., Caiyun, L., Yang, Y. et al. Circular RNA MTCL1 targets SMAD3 by sponging miR-145‐5p for regulation of cell proliferation and migration in Hirschsprung’s disease. Pediatr Surg Int 40, 25 (2024). https://doi.org/10.1007/s00383-023-05621-9
Accepted:
Published:
DOI: https://doi.org/10.1007/s00383-023-05621-9