Abstract
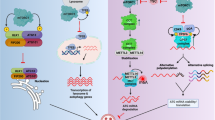
In this report, we discuss recent discoveries concerning the effects and specificity of different RNA-binding proteins (RBPs) as they pertain to macroautophagy/autophagy. Autophagy is a fundamental cellular degradation and recycling pathway, which has attracted substantial attention because defects in this process are associated with a wide range of human disorders including cancer, neurodegeneration, and metabolic diseases. Autophagy must be tightly controlled—either too much or too little can be deleterious. Therefore, understanding the complex regulation of autophagy is critical to achieve the goal of modulating the process for therapeutic purposes. Autophagy occurs constitutively, but is upregulated in response to stress. Here, we highlight a role for various RBPs in regulating particular autophagy-related (ATG) mRNAs. We briefly summarize recent publications, which focus on the RBPs Dhh1, Pat1, Lsm1–Lsm7 and Dcp2 in the post-transcriptional regulation of certain mRNAs that encode key components of the autophagy machinery. Finally, we consider how the established role of these and other RBPs in enhancing decap** and downregulating mRNAs is not their only function when it comes to regulating stress-related transcripts. Most ATG genes are downregulated during growth, in contrast to the vast majority of the genome; we discuss how certain regulatory factors play a key role in maintaining autophagy at a basal level during growth, while allowing for a rapid increase when cells encounter various stress conditions.
Similar content being viewed by others
References
Barve G, Sanyal P, Manjithaya R (2018) Septin localization and function during autophagy. Curr Genet 64:1037–1041
Bernard A et al (2015a) Rph1/KDM4 mediates nutrient-limitation signaling that leads to the transcriptional induction of autophagy. Curr Biol 25:546–555
Bernard A, ** M, Xu Z, Klionsky DJ (2015b) A large-scale analysis of autophagy-related gene expression identifies new regulators of autophagy. Autophagy 11:2114–2122
Chowdhury A, Mukhopadhyay J, Tharun S (2007) The decap** activator Lsm1p–7p–Pat1p complex has the intrinsic ability to distinguish between oligoadenylated and polyadenylated RNAs. RNA 13:998–1016
Chowdhury A, Kalurupalle S, Tharun S (2014) Pat1 contributes to the RNA binding activity of the Lsm1–7–Pat1 complex. RNA 20:1465–1475
Delorme-Axford E, Klionsky DJ (2018) On the edge of degradation: autophagy regulation by RNA decay. Wiley Interdiscip Rev RNA 17:e1522
Delorme-Axford E et al (2018) The exoribonuclease Xrn1 is a post-transcriptional negative regulator of autophagy. Autophagy 14:898–912
Ejzykowicz DE et al (2017) Hygromycin B hypersensitive (hhy) mutants implicate an intact trans-Golgi and late endosome interface in efficient Tor1 vacuolar localization and TORC1 function. Curr Genet 63:531–551
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24:24–41
Feng Y, Yao Z, Klionsky DJ (2015) How to control self-digestion: transcriptional, post-transcriptional, and post-translational regulation of autophagy. Trends Cell Biol 25:354–363
Feng Y et al (2016) Phosphorylation of Atg9 regulates movement to the phagophore assembly site and the rate of autophagosome formation. Autophagy 12:648–658
Garre E, Pelechano V, Sanchez Del Pino M, Alepuz P, Sunnerhagen P (2018) The Lsm1–7/Pat1 complex binds to stress-activated mRNAs and modulates the response to hyperosmotic shock. PLoS Genet 14:e1007563
Gatica D, Lahiri V, Klionsky DJ (2018) Cargo recognition and degradation by selective autophagy. Nat Cell Biol 20:233–242
Gatica D et al (2019) The Pat1–Lsm complex stabilizes ATG mRNA during nitrogen starvation-induced autophagy. Mol Cell 73:314–324 e314
He C, Klionsky DJ (2009) Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43:67–93
He F, Celik A, Wu C, Jacobson A (2018) General decap** activators target different subsets of inefficiently translated mRNAs. Elife 7:e34409
Hu G et al (2015) A conserved mechanism of TOR-dependent RCK-mediated mRNA degradation regulates autophagy. Nat Cell Biol 17:930–942
Jiang P, Mizushima N (2014) Autophagy and human diseases. Cell Res 24:69–79
** M, Klionsky DJ (2014) Regulation of autophagy: modulation of the size and number of autophagosomes. FEBS Lett 588:2457–2463
** M et al (2014) Transcriptional regulation by Pho23 modulates the frequency of autophagosome formation. Curr Biol 24:1314–1322
Parker R (2012) RNA degradation in Saccharomyces cerevisae. Genetics 191:671–702
Ramachandran V, Shah KH, Herman PK (2011) The cAMP-dependent protein kinase signaling pathway is a key regulator of P body foci formation. Mol Cell 43:973–981
Rybstein MD, Bravo-San Pedro JM, Kroemer G, Galluzzi L (2018) The autophagic network and cancer. Nat Cell Biol 20:243–251
Zhang N, Cao L (2017) Starvation signals in yeast are integrated to coordinate metabolic reprogramming and stress response to ensure longevity. Curr Genet 63:839–843
Acknowledgements
This work was funded by National Institutes of Health Grant GM053396.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gatica, D., Klionsky, D.J. Towards understanding mRNA-binding protein specificity: lessons from post-transcriptional regulation of ATG mRNA during nitrogen starvation-induced autophagy. Curr Genet 65, 847–849 (2019). https://doi.org/10.1007/s00294-019-00943-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00943-5