Abstract
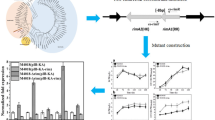
TetR family transcriptional regulators (TFRs) are widespread in actinomycetes, which exhibit diverse regulatory modes in antibiotic biosynthesis. Nitrogen regulators play vital roles in modulation of primary and secondary metabolism. However, crosstalk between TFR and nitrogen regulator has rarely been reported in actinomycetes. Herein, we demonstrated that a novel TFR, SACE_4839, was negatively correlated with erythromycin yield in Saccharopolyspora erythraea A226. SACE_4839 indirectly suppressed erythromycin synthetic gene eryAI and resistance gene ermE and directly inhibited its adjacent gene SACE_4838 encoding a homologue of nitrogen metabolite repression (NMR) regulator NmrA (herein named NmrR). The SACE_4839-binding sites within SACE_4839-nmrR intergenic region were identified. NmrR positively controlled erythromycin biosynthesis by indirectly stimulating eryAI and ermE and directly repressing SACE_4839. NmrR was found to affect growth viability under the nitrogen source supply. Furthermore, NmrR directly repressed glutamine and glutamate utilization-related genes SACE_1623, SACE_5070 and SACE_5979 but activated nitrate utilization-associated genes SACE_1163, SACE_4070 and SACE_4912 as well as nitrite utilization-associated genes SACE_1476 and SACE_4514. This is the first reported NmrA homolog for modulating antibiotic biosynthesis and nitrogen metabolism in actinomycetes. Moreover, combinatorial engineering of SACE_4839 and nmrR in the high-yield S. erythraea WB resulted in a 68.8% increase in erythromycin A production. This investigation deepens the understanding of complicated regulatory network for erythromycin biosynthesis.
Key points
• SACE_4839 and NmrR had opposite contributions to erythromycin biosynthesis.
• NmrR was first identified as a homolog of another nitrogen regulator NmrA.
• Cross regulation between SACE_4839 and NmrR was revealed.









Similar content being viewed by others
Data availability
All the data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Butler MS (2008) Natural products to drugs: natural product-derived compounds in clinical trials. Nat Prod Rep 25:475–516. https://doi.org/10.1039/b514294f
Cane DE (2010) Programming of erythromycin biosynthesis by a modular polyketide synthase. J Biol Chem 285:27517–27523. https://doi.org/10.1074/jbc.R110.144618
Cuthbertson L, Nodwell JR (2013) The TetR family of regulators. Microbiol Mol Biol Rev 77:440–475. https://doi.org/10.1128/MMBR
Han X, Qiu M, Wang B, Yin WB, Nie X, Qin Q, Ren S, Yang K, Zhang F, Zhuang Z, Wang S (2016) Functional analysis of the nitrogen metabolite repression regulator gene nmrA in Aspergillus flavus. Front Microbiol 7:1794. https://doi.org/10.3389/fmicb.2016.01794
Hellman LM, Fried MG (2007) Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat Protoc 2:1849–1861. https://doi.org/10.1038/nprot.2007.249
Hiard S, Marée R, Colson S, Hoskisson PA, Titgemeyer F, van Wezel GP, Joris B, Wehenkel L, Rigali S (2007) PREDetector: a new tool to identify regulatory elements in bacterial genomes. Biochem Biophys Res Commun 357:861–864. https://doi.org/10.1016/j.bbrc.2007.03.180
Jiang N, **ao D, Zhang D, Sun N, Yan B, Zhu X (2009) Negative roles of a novel nitrogen metabolite repression-related gene, TAR1, in laccase production and nitrate utilization by the basidiomycete Cryptococcus neoformans. Appl Environ Microbiol 75:6777–6782. https://doi.org/10.1128/AEM.00708-09
Khosla C, Tang Y, Chen AY, Schnarr NA, Cane DE (2007) Structure and mechanism of the 6-deoxyerythronolide B synthase. Annu Rev Biochem 76:195–221. https://doi.org/10.1146/annurev.biochem.76.053105.093515
Li C, Zhang Q, **a Y, ** K (2021) MaNmrA, a negative transcription regulator in nitrogen catabolite repression pathway, contributes to nutrient utilization, stress resistance, and virulence in entomopathogenic fungus Metarhizium acridum. Biology (basel) 10:1167. https://doi.org/10.3390/biology10111167
Liao CH, Yao LL, Ye BC (2014) Three genes encoding citrate synthases in Saccharopolyspora erythraea are regulated by the global nutrient-sensing regulators GlnR, DasR, and CRP. Mol Microbiol 94:1065–1084. https://doi.org/10.1111/mmi.12818
Liu G, Chater KF, Chandra G, Niu G, Tan H (2013) Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol Rev 77:112–143. https://doi.org/10.1128/MMBR.00054-12
Liu J, Chen Y, Wang W, Ren M, Wu P, Wang Y, Li C, Zhang L, Wu H, Weaver DT, Zhang B (2017) Engineering of an Lrp family regulator SACE_Lrp improves erythromycin production in Saccharopolyspora erythraea. Metab Eng 39:29–37. https://doi.org/10.1016/j.ymben.2016.10.012
Liu J, Chen Y, Li L, Yang E, Wang Y, Wu H, Zhang L, Wang W, Zhang B (2019) Characterization and engineering of the Lrp/AsnC family regulator SACE_5717 for erythromycin overproduction in Saccharopolyspora erythraea. J Ind Microbiol Biotechnol 46:1013–1024. https://doi.org/10.1007/s10295-019-02178-2
Liu J, Li L, Wang Y, Li B, Cai X, Tang L, Dong S, Yang E, Wu H, Zhang B (2021a) Joint engineering of SACE_Lrp and its target MarR enhances the biosynthesis and export of erythromycin in Saccharopolyspora erythraea. Appl Microbiol Biotechnol 105:2911–2924. https://doi.org/10.1007/s00253-021-11228-8
Liu Y, Khan S, Wu P, Li B, Liu L, Ni J, Zhang H, Chen K, Wu H, Zhang B (2021b) Uncovering and engineering a mini-regulatory network of the TetR-family regulator SACE_0303 for yield improvement of erythromycin in Saccharopolyspora erythraea. Front Bioeng Biotechnol 9:692901. https://doi.org/10.3389/fbioe.2021.692901
Macios M, Caddick MX, Weglenski P, Scazzocchio C, Dzikowska A (2012) The GATA factors AREA and AREB together with the co-repressor NMRA, negatively regulate arginine catabolism in Aspergillus nidulans in response to nitrogen and carbon source. Fungal Genet Biol 49:189–198. https://doi.org/10.1016/j.fgb.2012.01.004
Nett M, Ikeda H, Moore BS (2009) Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat Prod Rep 26:1362–1384. https://doi.org/10.1039/b817069j
Oliynyk M, Samborskyy M, Lester JB, Mironenko T, Scott N, Dickens S, Haydock SF, Leadlay PF (2007) Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338. Nat Biotechnol 25:447–453. https://doi.org/10.1038/nbt1297
Pullan ST, Chandra G, Bibb MJ, Merrick M (2011) Genome-wide analysis of the role of GlnR in Streptomyces venezuelae provides new insights into global nitrogen regulation in actinomycetes. BMC Genomics 12:175. https://doi.org/10.1186/1471-2164-12-175
Reeve LM, Baumberg S (1998) Physiological controls of erythromycin production by Saccharopolyspora erythraea are exerted at least in part at the level of transcription. Biotechnol Lett 20:585–589. https://doi.org/10.1023/A:1005357930000
Ren CY, Liu Y, Wei WP, Dai J, Ye BC (2021) Reconstruction of secondary metabolic pathway to synthesize novel metabolite in Saccharopolyspora erythraea. Front Bioeng Biotechnol 9:628569. https://doi.org/10.3389/fbioe.2021.628569
Romero-Rodríguez A, Maldonado-Carmona N, Ruiz-Villafán B, Koirala N, Rocha D, Sánchez S (2018) Interplay between carbon, nitrogen and phosphate utilization in the control of secondary metabolite production in Streptomyces. Antonie Van Leeuwenhoek 111:761–781. https://doi.org/10.1007/s10482-018-1073-1
Stammers DK, Ren J, Leslie K, Nichols CE, Lamb HK, Cocklin S, Dodds A, Hawkins AR (2001) The structure of the negative transcriptional regulator NmrA reveals a structural superfamily which includes the short-chain dehydrogenase/reductases. EMBO J 20:6619–6626. https://doi.org/10.1093/emboj/20.23.6619
Tiffert Y, Supra P, Wurm R, Wohlleben W, Wagner R, Reuther J (2008) The Streptomyces coelicolor GlnR regulon: identification of new GlnR targets and evidence for a central role of GlnR in nitrogen metabolism in actinomycetes. Mol Microbiol 67:861–880. https://doi.org/10.1111/j.1365-2958.2007.06092.x
Tudzynski B (2014) Nitrogen regulation of fungal secondary metabolism in fungi. Front Microbiol 5:656. https://doi.org/10.3389/fmicb.2014.00656
van der Heul HU, Bilyk BL, McDowall KJ, Seipke RF, van Wezel GP (2018) Regulation of antibiotic production in Actinobacteria: new perspectives from the post-genomic era. Nat Prod Rep 35:575–604. https://doi.org/10.1039/c8np00012c
Wu H, Chen M, Mao Y, Li W, Liu J, Huang X, Zhou Y, Ye BC, Zhang L, Weaver DT, Zhang B (2014a) Dissecting and engineering of the TetR family regulator SACE_7301 for enhanced erythromycin production in Saccharopolyspora erythraea. Microb Cell Fact 13:158. https://doi.org/10.1186/s12934-014-0158-4
Wu P, Pan H, Zhang C, Wu H, Yuan L, Huang X, Zhou Y, Ye BC, Weaver DT, Zhang L, Zhang B (2014b) SACE_3986, a TetR family transcriptional regulator, negatively controls erythromycin biosynthesis in Saccharopolyspora erythraea. J Ind Microbiol Biotechnol 41:1159–1167. https://doi.org/10.1007/s10295-014-1449-9
Wu H, Wang Y, Yuan L, Mao Y, Wang W, Zhu L, Wu P, Fu C, Müller R, Weaver DT, Zhang L, Zhang B (2016) Inactivation of SACE_3446, a TetR family transcriptional regulator, stimulates erythromycin production in Saccharopolyspora erythraea. Synth Syst Biotechnol 1:39–46. https://doi.org/10.1016/j.synbio.2016.01.004
Wu H, Chu Z, Zhang W, Zhang C, Ni J, Fang H, Chen Y, Wang Y, Zhang L, Zhang B (2019) Transcriptome-guided target identification of the TetR-like regulator SACE_5754 and engineered overproduction of erythromycin in Saccharopolyspora erythraea. J Biol Eng 13:11. https://doi.org/10.1186/s13036-018-0135-2
Wu P, Chen K, Li B, Zhang Y, Wu H, Chen Y, Ren S, Khan S, Zhang L, Zhang B (2021) Polyketide starter and extender units serve as regulatory ligands to coordinate the biosynthesis of antibiotics in actinomycetes. mBio 12: e0229821. https://doi.org/10.1128/mBio.02298-21.
**a H, Zhan X, Mao XM, Li YQ (2020) The regulatory cascades of antibiotic production in Streptomyces. World J Microbiol Biotechnol 36:13. https://doi.org/10.1007/s11274-019-2789-4
Xu Z, Liu Y, Ye BC (2018) PccD regulates branched-chain amino acid degradation and exerts a negative effect on erythromycin production in Saccharopolyspora erythraea. Appl Environ Microbiol 84:e00049-e118. https://doi.org/10.1128/AEM.00049-18
Xu Y, You D, Yao LL, Chu X, Ye BC (2019) Phosphate regulator PhoP directly and indirectly controls transcription of the erythromycin biosynthesis genes in Saccharopolyspora erythraea. Microb Cell Fact 18:206. https://doi.org/10.1186/s12934-019-1258-y
Yao LL, Ye BC (2016) Reciprocal regulation of GlnR and PhoP in response to nitrogen and phosphate limitations in Saccharopolyspora erythraea. Appl Environ Microbiol 82:409–420. https://doi.org/10.1128/AEM.02960-15
Yao LL, Liao CH, Huang G, Zhou Y, Rigali S, Zhang B, Ye BC (2014) GlnR-mediated regulation of nitrogen metabolism in the actinomycete Saccharopolyspora erythraea. Appl Microbiol Biotechnol 98:7935–7948. https://doi.org/10.1007/s00253-014-5878-1
Young JL, Jarai G, Fu YH, Marzluf GA (1990) Nucleotide sequence and analysis of NMR, a negative-acting regulatory gene in the nitrogen circuit of Neurospora crassa. Mol Gen Genet 222:120–128. https://doi.org/10.1007/BF00283032
Yu H, Yao Y, Liu Y, Jiao R, Jiang W, Zhao GP (2007) A complex role of Amycolatopsis mediterranei GlnR in nitrogen metabolism and related antibiotics production. Arch Microbiol 188:89–96. https://doi.org/10.1007/s00203-007-0228-7
Funding
This study was supported by the National Natural Science Foundation of China (grant numbers 31972930, 32170073 and 31800057), the Anhui Provincial Natural Science Foundation (grant number 2208085Y09), the University Synergy Innovation Program of Anhui Province (grant number GXXT-2019–035), and the Anhui Provincial Program on Key Research and Development Project (grant number 201904a07020080).
Author information
Authors and Affiliations
Contributions
BZ, HW and PW conceived and designed the study. SK, XX, JS, XL, KC and ZX performed the experiments. SK, XX, JS, PW, JL and HW analyzed the data. SK and PW wrote the manuscript. HW and BZ checked the final version. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, S., Xu, X., Song, J. et al. Crosstalk of TetR-like regulator SACE_4839 and a nitrogen regulator for erythromycin biosynthesis. Appl Microbiol Biotechnol 106, 6551–6566 (2022). https://doi.org/10.1007/s00253-022-12153-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12153-0