Abstract
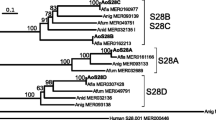
Aspartic proteases exhibit optimum enzyme activity under acidic conditions and have been extensively used in food, fermentation, and leather industries. In this study, a novel aspartic protease precursor (proTlAPA1) from Talaromyces leycettanus was identified and successfully expressed in Pichia pastoris. Subsequently, the auto-activation processing of the zymogen proTlAPA1 was studied by SDS-PAGE and N-terminal sequencing, under different processing conditions. TlAPA1 shared the highest identity of 70.3% with the aspartic endopeptidase from Byssochlamys spectabilis (GAD91729) and was classified into a new subgroup of the aspartic protease A1 family, based on evolutionary analysis. Mature TlAPA1 protein displayed an optimal activity at 60 °C and remained stable at temperatures of 55 °C and below, indicating the thermostable nature of TlAPA1 aspartic protease. During the auto-activation processing of proTlAPA1, a 45-kDa intermediate was identified that divided the processing mechanism into two steps: formation of intermediates and activation of the mature protein (TlAPA1). The former step can be processed without proteolytic activity, while the latter process depended on protease activity completely. The discovery of the novel aspartic protease TlAPA1 and the study of its activation process will contribute to a better understanding of the mechanism of aspartic protease auto-activation.







Similar content being viewed by others
References
Chen Z, Koelsch G, Han HP, Wang XJ, Lin XL, Hartsuck JA, Tang J (1991) Recombinant rhizopuspepsinogen: expression, purification, and activation properties of recombinant rhizopuspepsinogens. J Biol Chem 266:11718–11725
Domingos A, Cardoso PC, Xue ZT, Clemente A, Brodelius PE, Pais MS (2000) Purification, cloning and autoproteolytic processing of an aspartic proteinase from Centaurea calcitrapa. Eur J Biochem 267:6824–6831
Dostal J, Dlouha H, Malon P, Pichova I, Hruskova-Heidingsfeldova O (2005) The precursor of secreted aspartic proteinase sapp1p from Candida parapsilosis can be activated both autocatalytically and by a membrane-bound processing proteinase. J Biol Chem 386:791–799
Dunn BM (2002) Structure and mechanism of the pepsin-like family of aspartic peptidases. Chem Rev 102:4431–4458
Fraser ME, Strynadka NC, Bartlett PA, Hanson JE, James MN (1992) Crystallographic analysis of transition-statemimics bound to penicillopepsin: phosphorus-containing peptide analogues. Biochemistry 31:5201–5214
Gama Salgado JA, Kangwa M, Fernandez-Lahore M (2013) Cloning and expression of an active aspartic proteinase from Mucor circinelloides in Pichia pastoris. BMC Microbiol 13:250–260
Glathe S, Kervinen J, Nimtz M, Li GH, Tobin GJ, Copeland TD, Ashford DA, Wlodawer A, Costa J (1998) Transport and activation of vacuolar aspartic proteinase phytepsin in barley (Hordeum vulgare L.). J Biol Chem 273:31230–31236
Gomi K, Arikawa K, Kamiya N, Kitamoto K, Kumagai C (1993) Cloning and nucleotide sequence of the acid protease-encoding gene (pepA) from Aspergillus oryzae. Biosci Biotechnol Biochem 57:1095–1100
Guo Y, Tu T, Yuan P, Wang Y, Ren Y, Yao B, Luo H (2019) High-level expression and characterization of a novel aspartic protease from Talaromyces leycettanus JCM12802 and its potential application in juice clarification. Food Chem 281:197–203
Hanova I, Brynda J, Houstecka R, Alam N, Sojka D, Kopacek P, Maresova L, Vondrasek J, Horn M, Schueler-Furman O, Mares M (2018) Novel structural mechanism of allosteric regulation of aspartic peptidases via an evolutionarily conserved exosite. Cell Chem Biol 25:318–329
Horimoto Y, Dee DR, Yada RY (2009) Multifunctional aspartic peptidase prosegments. New Biotechnol 25:318–324
Kamitori S, Ohtaki A, Ino H, Takeuchi M (2003) Crystal structures of Aspergillus oryzae aspartic proteinase and its complex with an inhibitor pepstatin at 1.9A resolution. J Mol Biol 326:1503–1511
Kangwa M, Salgado JAG, Fernandez-Lahore HM (2018) Identification and characterization of N-glycosylation site on a Mucor circinelloides aspartic protease expressed in Pichia pastoris: effect on secretion, activity and thermo-stability. AMB Express 8:157–170
Khan AR, Khazanovich-Bernstein N, Bergmann EM, James MNG (1999) Structural aspects of activation pathways of aspartic protease zymogens and viral 3C protease precursors. Proc Natl Acad Sci U S A 96:10968–10975
Koelsch G, Mareš M, Metcalf P, Fusek M (1994) Multiple functions of pro-parts of aspartic proteinase zymogens. FEBS Lett 343:6–10
Kumar S, Sharma NS, Saharan MR, Singh R (2005) Extracellular acid protease from Rhizopus oryzae: purification and characterization. Process Biochem 40:1701–1705
Kumar S, Stecher G, Tamura K (2016) Mega 7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lakshman PLN, Toyokawa Y, Toyama H, Taira T, Yasuda M (2010) Purification and characterisation of two extracellular acid proteinases from Monascus pilosus. Food Chem 121:1216–1224
Lee AY, Gulnik SV, Erickson JW (1998) Conformational switching in an aspartic proteinase. Nat Struct Biol 5:866–871
Mandujano-González V, Villa-Tanaca L, Anducho-Reyes MA, Mercado-Flores Y (2016) Secreted fungal aspartic proteases: a review. Rev Iberoam Micol 33:76–82
Morales R, Watier Y, Bocskei Z (2012) Human prorenin structure sheds light on a novel mechanism of its autoinhibition and on its non-proteolytic activation by the (pro)renin receptor. J Mol Biol 421:100–111
Nascimento AS, Krauchenco S, Golubev AM, Gustchina A, Wlodawer A, Polikarpov I (2008) Statistical coupling analysis of aspartic proteinases based on crystal structures of the Trichoderma reesei enzyme and its complex with pepstatin a. J Mol Biol 382:763–778
Oka T, Ekino K, Fukud K, Nomura Y (2014) Draft genome sequence of the formaldehyde-resistant fungus Byssochlamys spectabilis No. 5. Genome Announc 2:e01162–e01113
Ostermann N, Gerhartz B, Worpenberg S, Trappe J, Eder J (2004) Crystal structure of an activation intermediate of cathepsin E. J Mol Biol 342:889–899
Rao MB, Tanksale AM, Ghatge MS, Deshpande VV (1998) Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev 62:597–635
Rao S, Mizutani O, Hirano T, Masaki K, Iefuji H (2011) Purification and characterization of a novel aspartic protease from basidiomycetous yeast Cryptococcus sp. S-2. J Biosci Bioeng 112:441–446
Raveendran S, Parameswaran B, Ummalyma SB, Abraham A, Mathew AK, MadhavanA RS, Pandey A (2018) Applications of microbial enzymes in food industry. Food Technol Biotechnol 56:16–30
Rawlings ND, Barrett AJ (2013) Introduction: aspartic and glutamic peptidases and their clans. In: Barrett AJ, Rawlings ND, Woessner JF (eds) Handbook of proteolytic enzymes, vol 2, 3rd edn. Elsevier, Amsterdam, pp 3–19
Rawlings ND, Barrett AJ, Thomas PD, Huang X, Bateman A, Finn RD (2018) The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res 46:624–632
Revuelta MV, Kan JALV, Kay J, Have AT (2014) Extensive expansion of A1 family aspartic proteinases in fungi revealed by evolutionary analyses of 107 complete eukaryotic proteomes. Genome Biol Evol 6:1480–1494
Richter C, Tanaka T, Yada RY (1998) Mechanism of activation of the gastric aspartic proteinases: pepsinogen, progastricsin and prochymosin. Biochem J 335:481–490
Silva RR, Suoto TB, Oliveira TB, Oliveira LCG, Karcher D, Juliano MA, Juliano L, Oliveira AHC, Rodrigues A, Rosa JC (2016) Evaluation of the catalytic specificity, biochemical properties, and milk clotting abilities of an aspartic peptidase from Rhizomucor miehei. J Ind Microbiol Biotechnol 43:1059–1069
Souza PM, Bittencourt MLD, Caprara CC, Freitas M, Almeida RPC, Silveira D, Fonseca YM, Ferreira EX, Pessoa A, Magalhaes PO (2015) A biotechnology perspective of fungal proteases. Braz J Microbiol 46:337–346
Souza PMD, Werneck G, Aliakbarian B, Siqueira F, Ferreira FEX, Perego P, Converti A, Magalhaes PO, Junior AP (2017) Production, purification and characterization of an aspartic protease from Aspergillus foetidus. Food Chem Toxicol 109:1103–1110
Sun Q, Chen F, Geng F, Luo Y, Gong S, Jiang Z (2018) A novel aspartic protease from Rhizomucor miehei expressed in Pichia pastoris and its application on meat tenderization and preparation of turtle peptides. Food Chem 245:570–577
Takenaka S, Umeda M, Senba H, Koyama D, Tanaka K, Yoshida KI, Doi M (2017) Heterologous expression and characterisation of the Aspergillus aspartic protease involved in the hydrolysis and decolorisation of red-pigmented proteins. J Sci Food Agric 97:95–101
Vishwanatha KS, Rao AGA, Singh SA (2009) Characterisation of acid protease expressed from Aspergillus oryzae MTCC 5341. Food Chem 114:402–407
Wang C, Luo H, Niu C, Shi P, Huang H, Meng K, Bai Y, Wang K, Hua H, Yao B (2015) Biochemical characterization of a thermophilic β-mannanase from Talaromyces leycettanus JCM12802 with high specific activity. Appl Microbiol Biotechnol 99:1217–1228
Wang X, Ma R, **e X, Liu W, Tu T, Zheng F, You S, Ge J, **e H, Yao B, Luo H (2017) Thermostability improvement of a Talaromyces leycettanus xylanase by rational protein engineering. Sci Rep 7:152–187
Yang X, Cong H, Song J, Zhang J (2013) Heterologous expression of an aspartic protease gene from biocontrol fungus Trichoderma asperellum in Pichia pastoris. World J Microbiol Biotechnol 29:2087–2094
Yegin S, Dekker P (2013) Progress in the field of aspartic proteinases in cheese manufacturing: structures, functions, catalytic mechanism, inhibition, and engineering. Dairy Sci Technol 93:565–594
Yegin S, Fernandez-Lahore M (2013) A thermostable aspartic proteinase from Mucor mucedo DSM 809: gene identification, cloning, and functional expression in Pichia pastoris. Mol Biotechnol 54:661–672
Yegin S, Fernandez-Lahore M, Salgado AJG, Guvenc U, Goksungur Y, Tari C (2011) Aspartic proteinases from Mucor spp. in cheese manufacturing. Appl Microbiol Biotechnol 89:949–960
You S, Tu T, Zhang L, Wang Y, Huang H, Ma R, Shi PJ, Bai YG, Su XY, Lin ZM, Luo HY, Yao B (2016) Improvement of the thermostability and catalytic efficiency of a highly active β-glucanase from Talaromyces leycettanus JCM12802 by optimizing residual charge–charge interactions. Biotechnol Biofuels 9:124–135
Zhang D, Tu T, Wang Y, Li Y, Luo X, Zheng F, Wang X, Bai Y, Huang H, Su X, Yao B, Zhang T, Luo H (2017) Improvement of the catalytic performance of a Talaromyces leycettanusα-amylase by changing the linker length. J Agric Food Chem 65:5041–5048
Zhang Y, He S, Simpson BK (2018) Enzymes in food bioprocessing–novel food enzymes, applications, and related techniques. Curr Opin in Food Sci19:30–35
Funding
This work was financially supported by the National Key Research and Development Program of China (2016YFD0501409-02), the Fundamental Research Funds for Central Non-profit Scientific Institution (Y2017JC31), and the China Modern Agriculture Research System (CARS-41).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 254 kb)
Rights and permissions
About this article
Cite this article
Guo, Y., Tu, T., Zheng, J. et al. A novel thermostable aspartic protease from Talaromyces leycettanus and its specific autocatalytic activation through an intermediate transition state. Appl Microbiol Biotechnol 104, 4915–4926 (2020). https://doi.org/10.1007/s00253-020-10569-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10569-0