Abstract
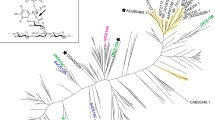
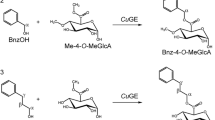
Glucuronoyl esterases (CE15 family) enable targeted cleavage of ester linkages in lignin-carbohydrate complexes (LCCs), particularly those linking lignin and glucuronoyl residues in xylan. A substantial challenge in characterization and kinetic analysis of CE15 enzymes has been the lack of proper substrates. Here, we present an assay using an insoluble LCC-rich lignin fraction from birch; lignin-rich pellet (LRP). The assay employs quantification of enzyme reaction products by LC-MS. The kinetics of four fungal CE15 enzymes, PsGE, CuGE, TtGE, and AfuGE originating from lignocellulose-degrading fungi Punctularia strigosozonata, Cerrena unicolor, Thielavia terrestris, and Armillaria fuscipes respectively were characterized and compared using this new assay. All four enzymes had activity on LRP and showed a clear preference for the insoluble substrate compared with smaller soluble LCC mimicking esters. End-product profiles were near identical for the four enzymes but differences in kinetic parameters were observed. TtGE possesses an alternative active site compared with the three other enzymes as it has the position of the catalytic glutamic acid occupied by a serine. TtGE performed poorly compared with the other enzymes. We speculate that glucuronoyl LCCs are not the preferred substrate of TtGE. Removal of an N-terminal CBM on CuGE affected the catalytic efficiently of the enzyme by reducing Kcat by more than 30%. Reaction products were detected from all four CE15s on a similar substrate from spruce indicating a more generic GE activity not limited to the hardwood. The assay with natural substrate represents a novel tool to study the natural function and kinetics of CE15s.





Similar content being viewed by others
References
Agger JW, Busk PK, Pilgaard B, Meyer AS, Lange L (2017) A new functional classification of glucuronoyl esterases by peptide pattern recognition. Front Microbiol 8:309. https://doi.org/10.3389/fmicb.2017.00309
Bååth JA, Giummarella N, Klaubauf S, Lawoko M, Olsson L (2016) A glucuronoyl esterase from Acremonium alcalophilum cleaves native lignin-carbohydrate ester bonds. FEBS Lett 590:2611–2618. https://doi.org/10.1002/1873-3468.12290
Bååth JA, Mazurkewich S, Knudsen RM, Poulsen J-CN, Olsson L, Lo Leggio L, Larsbrink J (2018) Biochemical and structural features of diverse bacterial glucuronoyl esterases facilitating recalcitrant biomass conversion. Biotechnol Biofuels 11:213. https://doi.org/10.1186/s13068-018-1213-x
Biely P (2016) Microbial glucuronoyl esterases–ten years after discovery. Appl Environ Microbiol 82:7014–7018. https://doi.org/10.1128/AEM.02396-16
Biely P, Malovíková A, Uhliariková I, Li X-L, Wong DWS (2015) Glucuronoyl esterases are active on the polymeric substrate methyl esterified glucuronoxylan. FEBS Lett 589:2334–2339. https://doi.org/10.1016/j.febslet.2015.07.019
Charavgi MD, Dimarogona M, Topakas E, Christakopoulos P, Chrysina ED (2013) The structure of a novel glucuronoyl esterase from Myceliophthora thermophila gives new insights into its role as a potential biocatalyst. Acta Crystallogr Sect D Biol Crystallogr 69:63–73. https://doi.org/10.1107/S0907444912042400
D’Errico C, Börjesson J, Ding H, Krogh KBRM, Spodsberg N, Madsen R, Monrad RN (2016) Improved biomass degradation using fungal glucuronoyl-esterases-hydrolysis of natural corn fiber substrate. J Biotechnol 219:117–123. https://doi.org/10.1016/j.jbiotec.2015.12.024
d’Errico C, Jørgensen JO, Krogh KBRM, Spodsberg N, Madsen R, Monrad RN (2015) Enzymatic degradation of lignin-carbohydrate complexes (LCCs): model studies using a fungal glucuronoyl esterase from Cerrena unicolor. Biotechnol Bioeng 112:914–922. https://doi.org/10.1002/bit.25508
De Santi C, Gani OA, Helland R, Williamson A (2017) Structural insight into a CE15 esterase from the marine bacterial metagenome. Sci Rep 7:17278. https://doi.org/10.1038/s41598-017-17677-4
De Santi C, Willassen NP, Williamson A (2016) Biochemical characterization of a family 15 carbohydrate esterase from a bacterial marine arctic metagenome. PLoS One 11:0159345. https://doi.org/10.1371/journal.pone.0159345
Dilokpimol A, Mäkelä MR, Cerullo G, Zhou M, Varriale S, Gidijala L, Brás JLA, Jütten P, Piechot A, Verhaert R, Faraco V, Hilden KS, de Vries RP (2018) Fungal glucuronoyl esterases: genome mining based enzyme discovery and biochemical characterization. New Biotechnol 40:282–287. https://doi.org/10.1016/j.nbt.2017.10.003
Du X, Gellerstedt G, Li J (2013) Universal fractionation of lignin-carbohydrate complexes (LCCs) from lignocellulosic biomass: an example using spruce wood. Plant J 74:328–338. https://doi.org/10.1111/tpj.12124
Ďuranová M, Špániková S, Wösten HAB, Biely P, de Vries RP (2009) Two glucuronoyl esterases of Phanerochaete chrysosporium. Arch Microbiol 191:133–140. https://doi.org/10.1007/s00203-008-0434-y
Hüttner S, Klaubauf S, de Vries RP, Olsson L (2017) Characterisation of three fungal glucuronoyl esterases on glucuronic acid ester model compounds. Appl Microbiol Biotechnol 101:5301–5311. https://doi.org/10.1007/s00253-017-8266-9
Huynh HH, Ishii N, Matsuo I, Arioka M (2018) A novel glucuronoyl esterase from Aspergillus fumigatus—the role of conserved Lys residue in the preference for 4-O-methyl glucuronoyl esters. Appl Microbiol Biotechnol 102:2191–2201. https://doi.org/10.1007/s00253-018-8739-5
Jeffries TW (1990) Biodegradation of lignin-carbohydrate complexes. Biodegradation 1:163–176. https://doi.org/10.1007/BF00058834
Lin M-I, Hiyama A, Kondo K, Nagata T, Katahira M (2018) Classification of fungal glucuronoyl esterases (FGEs) and characterization of two new FGEs from Ceriporiopsis subvermispora and Pleurotus eryngii. Appl Microbiol Biotechnol 102:9635–9645. https://doi.org/10.1007/s00253-018-9318-5
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:490–495. https://doi.org/10.1093/nar/gkt1178
Monrad RN, Eklöf J, Krogh KBRM, Biely P (2018) Glucuronoyl esterases: diversity, properties and biotechnological potential. A review. Crit Rev Biotechnol 38:1121–1136. https://doi.org/10.1080/07388551.2018.1468316
Mosbech C, Holck J, Meyer AS, Agger JW (2018) The natural catalytic function of CuGE glucuronoyl esterase in hydrolysis of genuine lignin–carbohydrate complexes from birch. Biotechnol Biofuels 11:71. https://doi.org/10.1186/s13068-018-1075-2
Nylander F, Sunner H, Olsson L, Christakopoulos P (2015) Synthesis and enzymatic hydrolysis of a diaryl benzyl ester model of a lignin-carbohydrate complex (LCC). Holzforschung 70:385–488. https://doi.org/10.1515/hf-2014-0347
Pawar PM-A, Koutaniemi S, Tenkanen M, Mellerowicz EJ (2013) Acetylation of woody lignocellulose: significance and regulation. Front Plant Sci 4:118. https://doi.org/10.3389/fpls.2013.00118
Pokkuluri PR, Duke NEC, Wood SJ, M a. C, Li XL, Biely P, Schiffer M (2011) Structure of the catalytic domain of glucuronoyl esterase Cip2 from Hypocrea jecorina. Proteins Struct Funct Bioinforma 79:2588–2592. https://doi.org/10.1002/prot.23088
Špániková S, Biely P (2006) Glucuronoyl esterase – novel carbohydrate esterase produced by Schizophyllum commune. FEBS Lett 580:4597–4601. https://doi.org/10.1016/j.febslet.2006.07.033
Špániková S, Poláková M, Joniak D, Hirsch J, Biely P (2007) Synthetic esters recognized by glucuronoyl esterase from Schizophyllum commune. Arch Microbiol 188:185–189. https://doi.org/10.1007/s00203-007-0241-x
Tang J, Long L, Cao Y, Ding S (2019) Expression and characterization of two glucuronoyl esterases from Thielavia terrestris and their application in enzymatic hydrolysis of corn bran. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-019-09662-w
Tarasov D, Leitch M, Fatehi P (2018) Lignin–carbohydrate complexes: properties, applications, analyses, and methods of extraction: a review. Biotechnol Biofuels 11:269. https://doi.org/10.1186/s13068-018-1262-1
Topakas E, Moukouli M, Dimarogona M, Vafiadi C, Christakopoulos P (2010) Functional expression of a thermophilic glucuronoyl esterase from Sporotrichum thermophile: identification of the nucleophilic serine. Appl Microbiol Biotechnol 87:1765–1772. https://doi.org/10.1007/s00253-010-2655-7
Watanabe T, Koshijima T (1988) Evidence for an ester linkage between lignin and glucuronic acid in lignin-carbohydrate complexes by DDQ-oxidation. Agric Biol Chem 52:2953–2955. https://doi.org/10.1080/00021369.1988.10869116
Acknowledgement
We sincerely thank Novozymes A/S for the donation of GH10 containing preparation.
Funding
This work was partly financed by the grant NNF15OC0015222 funded by the Novo Nordisk Foundation and also supported by the Bio-Value Strategic Platform for Innovation and Research, co-funded by The Danish Council for Strategic Research and The Danish Council for Technology and Innovation, case no: 0603-00522B.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 965 kb)
Rights and permissions
About this article
Cite this article
Mosbech, C., Holck, J., Meyer, A. et al. Enzyme kinetics of fungal glucuronoyl esterases on natural lignin-carbohydrate complexes. Appl Microbiol Biotechnol 103, 4065–4075 (2019). https://doi.org/10.1007/s00253-019-09797-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09797-w