Abstract
HLA class I molecules and killer cell immunoglobulin-like receptors (KIR) form a diverse system of ligands and receptors that individualize human immune systems in ways that improve the survival of individuals and populations. Human settlement of Oceania by island-hop** East and Southeast Asian migrants started ~3,500 years ago. Subsequently, New Zealand was reached ~750 years ago by ancestral Māori. To examine how this history impacted KIR and HLA diversity, and their functional interaction, we defined at high resolution the allelic and haplotype diversity of the 13 expressed KIR genes in 49 Māori and 34 Polynesians. Eighty KIR variants, including four ‘new’ alleles, were defined, as were 35 centromeric and 22 telomeric KIR region haplotypes, which combine to give >50 full-length KIR haplotypes. Two new and divergent variant KIR form part of a telomeric KIR haplotype, which appears derived from Papua New Guinea and was probably obtained by the Asian migrants en route to Polynesia. Māori and Polynesian KIR are very similar, but differ significantly from African, European, Japanese, and Amerindian KIR. Māori and Polynesians have high KIR haplotype diversity with corresponding allotype diversity being maintained throughout the KIR locus. Within the population, each individual has a unique combination of HLA class I and KIR. Characterizing Māori and Polynesians is a paucity of HLA-B allotypes recognized by KIR. Compensating for this deficiency are high frequencies (>50 %) of HLA-A allotypes recognized by KIR. These HLA-A allotypes are ones that modern humans likely acquired from archaic humans at a much earlier time.







Similar content being viewed by others
References
Abi-Rached L et al (2011) The sha** of modern human immune systems by multiregional admixture with archaic humans. Science (New York, NY) 334:89–94. doi:10.1126/science.1209202
Ahlenstiel G, Martin MP, Gao X, Carrington M, Rehermann B (2008) Distinct KIR/HLA compound genotypes affect the kinetics of human antiviral natural killer cell responses. J Clin Invest 118:1017–1026. doi:10.1172/jci32400
Anderson NH, Sadler LC, Stewart AW, Fyfe EM, McCowan LM (2012) Ethnicity, body mass index and risk of pre-eclampsia in a multiethnic New Zealand population. Aust N Z J Obstet Gynaecol 52:552–558. doi:10.1111/j.1479-828X.2012.01475.x
Bari R et al (2009) Significant functional heterogeneity among KIR2DL1 alleles and a pivotal role of arginine 245. Blood 114:5182–5190. doi:10.1182/blood-2009-07-231977
Bashirova AA, Martin MP, McVicar DW, Carrington M (2006) The killer immunoglobulin-like receptor gene cluster: tuning the genome for defense. Annu Rev Genomics Hum Genet 7:277–300. doi:10.1146/annurev.genom.7.080505.115726
Bellwood P, Chambers G, Ross M, Hung H-C (2011) Are ‘cultures’ inherited? Multidisciplinary perspectives on the origins of Austronesian-speaking peoples prior to 1000 BC. In: Roberts B, Vander Linden M (eds) Investigating archaeological cultures: material culture, variability and transmission. Springer, Berlin, pp 321–354
Burley D, Weisler MI, Zhao JX (2012) High precision u/th dating of first Polynesian settlement. PLoS One 7:e48769. doi:10.1371/journal.pone.0048769
Chambers GK (2013) Genetics and the origins of the Polynesians. John Wiley & Sons, Ltd, Chichester. doi:10.1002/9780470015902.a0020808.pub2
Cooper MA, Colonna M, Yokoyama WM (2009) Hidden talents of natural killers: NK cells in innate and adaptive immunity. EMBO Rep 10:1103–1110. doi:10.1038/embor.2009.203
Dohring C, Scheidegger D, Samaridis J, Cella M, Colonna M (1996) A human killer inhibitory receptor specific for HLA-A1,2. J Immunol 156:3098–3101
Duggan AT et al (2014) Maternal history of Oceania from complete mtDNA genomes: contrasting ancient diversity with recent homogenization due to the Austronesian expansion. Am J Hum Genet 94:721–733. doi:10.1016/j.ajhg.2014.03.014
Edinur HA, Dunn PP, Hammond L, Selwyn C, Velickovic ZM, Lea RA, Chambers GK (2012) Using HLA loci to inform ancestry and health in Polynesian and Maori populations. Tissue Antigens 80:509–522. doi:10.1111/tan.12026
Edinur HA et al (2013) HLA and MICA polymorphism in Polynesians and New Zealand Maori: implications for ancestry and health. Hum Immunol 74:1119–1129. doi:10.1016/j.humimm.2013.06.011
Fernandez Vina MA et al (2012) Tracking human migrations by the analysis of the distribution of HLA alleles, lineages and haplotypes in closed and open populations. Philos Trans R Soc Lond Ser B Biol Sci 367:820–829. doi:10.1098/rstb.2011.0320
Frazier WR, Steiner N, Hou L, Dakshanamurthy S, Hurley CK (2013) Allelic variation in KIR2DL3 generates a KIR2DL2-like receptor with increased binding to its HLA-C ligand. J Immunol 190:6198–6208
Friedlaender JS et al (2008) The genetic structure of Pacific Islanders. PLoS Genet 4:e19. doi:10.1371/journal.pgen.0040019
Gendzekhadze K, Norman PJ, Abi-Rached L, Graef T, Moesta AK, Layrisse Z, Parham P (2009) Co-evolution of KIR2DL3 with HLA-C in a human population retaining minimal essential diversity of KIR and HLA class I ligands. Proc Natl Acad Sci U S A 106:18692–18697
Gomez-Lozano N, Estefania E, Williams F, Halfpenny I, Middleton D, Solis R, Vilches C (2005) The silent KIR3DP1 gene (CD158c) is transcribed and might encode a secreted receptor in a minority of humans, in whom the KIR3DP1, KIR2DL4 and KIR3DL1/KIR3DS1 genes are duplicated. Eur J Immunol 35:16–24
Gonzalez-Galarza FF, Christmas S, Middleton D, Jones AR (2011) Allele frequency net: a database and online repository for immune gene frequencies in worldwide populations. Nucleic Acids Res 39:D913–D919. doi:10.1093/nar/gkq1128
Goodridge JP et al (2007) Three common alleles of KIR2DL4 (CD158d) encode constitutively expressed, inducible and secreted receptors in NK cells. Eur J Immunol 37:199–211
Graef T et al (2009) KIR2DS4 is a product of gene conversion with KIR3DL2 that introduced specificity for HLA-A*11 while diminishing avidity for HLA-C. J Exp Med 206:2557–2572
Gumperz JE, Litwin V, Phillips JH, Lanier LL, Parham P (1995) The Bw4 public epitope of HLA-B molecules confers reactivity with natural killer cell clones that express NKB1, a putative HLA receptor. J Exp Med 181:1133–1144
Hiby SE, Walker JJ, O'Shaughnessy KM, Redman CW, Carrington M, Trowsdale J, Moffett A (2004) Combinations of maternal KIR and fetal HLA-C genes influence the risk of preeclampsia and reproductive success. J Exp Med 200:957–965. doi:10.1084/jem.20041214
Hilton HG et al (2012) Mutation at positively selected positions in the binding site for HLA-C shows that KIR2DL1 is a more refined but less adaptable NK cell receptor than KIR2DL3. J Immunol 189:1418–1430. doi:10.4049/jimmunol.1100431
Hollenbach JA, Nocedal I, Ladner MB, Single RM, Trachtenberg EA (2012) Killer cell immunoglobulin-like receptor (KIR) gene content variation in the HGDP-CEPH populations. Immunogenetics 64:719–737. doi:10.1007/s00251-012-0629-x
Hollenbach JA et al (2013) 16(th) IHIW: population global distribution of killer immunoglobulin-like receptor (KIR) and ligands. Int J Immunogenet 40:39–45
Hou L, Chen M, Jiang B, Kariyawasam K, Ng J, Hurley CK (2009) In contrast to other stimulatory natural killer cell immunoglobulin-like receptor loci, several KIR2DS5 alleles predominate in African Americans. Hum Immunol 70:733–737
Hou L, Jiang B, Chen M, Ng J, Hurley CK (2011) The characteristics of allelic polymorphism in killer-immunoglobulin-like receptor framework genes in African Americans. Immunogenetics 63:549–559. doi:10.1007/s00251-011-0536-6
Hurles ME, Maund E, Nicholson J, Bosch E, Renfrew C, Sykes BC, Jobling MA (2003) Native American Y chromosomes in Polynesia: the genetic impact of the Polynesian slave trade. Am J Hum Genet 72:1282–1287. doi:10.1086/374827
Jiang W et al (2012) Copy number variation leads to considerable diversity for B but not A haplotypes of the human KIR genes encoding NK cell receptors. Genome Res 22:1845–1854. doi:10.1101/gr.137976.112
Kayser M et al (2006) Melanesian and Asian origins of Polynesians: mtDNA and Y chromosome gradients across the Pacific. Mol Biol Evol 23:2234–2244. doi:10.1093/molbev/msl093
Khakoo SI, Carrington M (2006) KIR and disease: a model system or system of models? Immunol Rev 214:186–201. doi:10.1111/j.1600-065X.2006.00459.x
Kidd JM et al (2014) Exome capture from saliva produces high quality genomic and metagenomic data. BMC Genomics 15:262. doi:10.1186/1471-2164-15-262
Kikuchi-Maki A, Yusa S, Catina TL, Campbell KS (2003) KIR2DL4 is an IL-2-regulated NK cell receptor that exhibits limited expression in humans but triggers strong IFN-gamma production. J Immunol 171:3415–3425
Kimura R et al (2008) Gene flow and natural selection in oceanic human populations inferred from genome-wide SNP ty**. Mol Biol Evol 25:1750–1761. doi:10.1093/molbev/msn128
Kostyu D et al (1984) HLA in two islands of French Polynesia. Tissue Antigens 23:217–228
Lanier LL (2005) NK cell recognition. Annu Rev Immunol 23:225–274. doi:10.1146/annurev.immunol.23.021704.115526
Leung W et al (2004) Determinants of antileukemia effects of allogeneic NK cells. J Immunol 172:644–650
Liu J, **ao Z, Ko HL, Shen M, Ren EC (2014) Activating killer cell immunoglobulin-like receptor 2DS2 binds to HLA-A*11. Proc Natl Acad Sci U S A 111:2662–2667. doi:10.1073/pnas.1322052111
Long EO, Kim HS, Liu D, Peterson ME, Rajagopalan S (2013) Controlling natural killer cell responses: integration of signals for activation and inhibition. Annu Rev Immunol 31:227–258. doi:10.1146/annurev-immunol-020711-075005
Martin MP, Carrington M (2013) Immunogenetics of HIV disease. Immunol Rev 254:245–264. doi:10.1111/imr.12071
Meyer M et al (2012) A high-coverage genome sequence from an archaic Denisovan individual. Science (New York, NY) 338:222–226. doi:10.1126/science.1224344
Meyer D et al (2007) Single locus polymorphism of classical HLA genes. In: Hansen JA (ed) Immunobiology of the human MHC: Proceedings of the 13th International Histocompatibility Workshop and Conference, vol I. IHWG Press, Seattle, pp 653–704
Middleton D, Meenagh A, Gourraud PA (2007) KIR haplotype content at the allele level in 77 Northern Irish families. Immunogenetics 59:145–158
Moesta AK, Norman PJ, Yawata M, Yawata N, Gleimer M, Parham P (2008) Synergistic polymorphism at two positions distal to the ligand-binding site makes KIR2DL2 a stronger receptor for HLA-C than KIR2DL3. J Immunol 180:3969–3979
Moretta L et al (2014) Human NK Cells: from surface receptors to the therapy of leukemias and solid tumors. Front Immunol 5:87. doi:10.3389/fimmu.2014.00087
Narni-Mancinelli E, Ugolini S, Vivier E (2013) Tuning the threshold of natural killer cell responses. Curr Opin Immunol 25:53–58. doi:10.1016/j.coi.2012.11.005
Norman PJ, Stephens HA, Verity DH, Chandanayingyong D, Vaughan RW (2001) Distribution of natural killer cell immunoglobulin-like receptor sequences in three ethnic groups. Immunogenetics 52:195–205
Norman PJ et al (2007) Unusual selection on the KIR3DL1/S1 natural killer cell receptor in Africans. Nat Genet 39:1092–1099
Norman PJ et al (2013) Co-evolution of human leukocyte antigen (HLA) class I ligands with killer-cell immunoglobulin-like receptors (KIR) in a genetically diverse population of sub-Saharan Africans. PLoS Genet 9:e1003938. doi:10.1371/journal.pgen.1003938
O'Connor GM et al (2014) Mutational and structural analysis of KIR3DL1 reveals a lineage-defining allotypic dimorphism that impacts both HLA and peptide sensitivity. J Immunol 192:2875–2884
Parham P (2005) MHC class I molecules and KIRs in human history, health and survival. Nat Rev Immunol 5:201–214
Parham P, Moffett A (2013) Variable NK cell receptors and their MHC class I ligands in immunity, reproduction and human evolution. Nat Rev Immunol 13:133–144
R Development Core Team (2008) A language and environment for statistical computing. Vienna, Austria
Rajagopalan S, Long EO (2012) Cellular senescence induced by CD158d reprograms natural killer cells to promote vascular remodeling. Proc Natl Acad Sci U S A 109:20596–20601
Roberts RL et al (2013) Prevalence of HLA-B27 in the New Zealand population: effect of age and ethnicity. Arthritis Res Ther 15:R158. doi:10.1186/ar4341
Robinson J, Halliwell JA, McWilliam H, Lopez R, Marsh SG (2013) IPD–the immuno polymorphism database. Nucleic Acids Res 41:D1234–D1240
Shilling HG, Guethlein LA, Cheng NW, Gardiner CM, Rodriguez R, Tyan D, Parham P (2002) Allelic polymorphism synergizes with variable gene content to individualize human KIR genotype. J Immunol 168:2307–2315
Soares P et al (2011) Ancient voyaging and Polynesian origins. Am J Hum Genet 88:239–247. doi:10.1016/j.ajhg.2011.01.009
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tao SD, He YM, Ying YL, He J, Zhu FM, Lv HJ (2014) KIR3DL1 genetic diversity and phenotypic variation in the Chinese Han population. Genes Immun 15:8–15
Thananchai H et al (2007) Cutting edge: allele-specific and peptide-dependent interactions between KIR3DL1 and HLA-A and HLA-B. J Immunol 178:33–37
Thorsby E (2012) The Polynesian gene pool: an early contribution by Amerindians to Easter Island. Philos Trans R Soc Lond Ser B Biol Sci 367:812–819. doi:10.1098/rstb.2011.0319
Tracey MC, Carter JM (2006) Class II HLA allele polymorphism: DRB1, DQB1 and DPB1 alleles and haplotypes in the New Zealand Maori population. Tissue Antigens 68:297–302
Uhrberg M et al (1997) Human diversity in killer cell inhibitory receptor genes. Immunity 7:753–763
Underhill PA, Passarino G, Lin AA, Marzuki S, Oefner PJ, Cavalli-Sforza LL, Chambers GK (2001) Maori origins, Y-chromosome haplotypes and implications for human history in the Pacific. Hum Mutat 17:271–280. doi:10.1002/humu.23
Velickovic M, Velickovic Z, Dunckley H (2006) Diversity of killer cell immunoglobulin-like receptor genes in Pacific Islands populations. Immunogenetics 58:523–532. doi:10.1007/s00251-006-0124-3
Venstrom JM et al (2012) HLA-C-dependent prevention of leukemia relapse by donor activating KIR2DS1. N Engl J Med 367:805–816. doi:10.1056/NEJMoa1200503
Vierra-Green C et al (2012) Allele-level haplotype frequencies and pairwise linkage disequilibrium for 14 KIR loci in 506 European–American individuals. PLoS One 7:e47491
Vivian JP et al (2011) Killer cell immunoglobulin-like receptor 3DL1-mediated recognition of human leukocyte antigen B. Nature 479:401–405
Whyte AL, Marshall SJ, Chambers GK (2005) Human evolution in Polynesia. Hum Biol 77:157–177
Williams F, Maxwell LD, Halfpenny IA, Meenagh A, Sleator C, Curran MD, Middleton D (2003) Multiple copies of KIR 3DL/S1 and KIR 2DL4 genes identified in a number of individuals. Hum Immunol 64:729–732
Wilmshurst JM, Hunt TL, Lipo CP, Anderson AJ (2011) High-precision radiocarbon dating shows recent and rapid initial human colonization of East Polynesia. Proc Natl Acad Sci U S A 108:1815–1820. doi:10.1073/pnas.1015876108
Wilson MJ et al (2000) Plasticity in the organization and sequences of human KIR/ILT gene families. Proc Natl Acad Sci U S A 97:4778–4783. doi:10.1073/pnas.080588597
Winter CC, Long EO (1997) A single amino acid in the p58 killer cell inhibitory receptor controls the ability of natural killer cells to discriminate between the two groups of HLA-C allotypes. J Immunol 158:4026–4028
Wollstein A et al (2010) Demographic history of Oceania inferred from genome-wide data. Curr Biol CB 20:1983–1992. doi:10.1016/j.cub.2010.10.040
Yawata M, Yawata N, Draghi M, Little AM, Partheniou F, Parham P (2006) Roles for HLA and KIR polymorphisms in natural killer cell repertoire selection and modulation of effector function. J Exp Med 203:633–645. doi:10.1084/jem.20051884
Yusa S, Catina TL, Campbell KS (2002) SHP-1- and phosphotyrosine-independent inhibitory signaling by a killer cell Ig-like receptor cytoplasmic domain in human NK cells. J Immunol 168:5047–5057
Zambello R, Teramo A, Barila G, Gattazzo C, Semenzato G (2014) Activating KIRs in chronic lymphoproliferative disorder of NK cells: protection from viruses and disease induction? Front Immunol 5:72. doi:10.3389/fimmu.2014.00072
Acknowledgments
We thank Jyothi Jayaraman for technical assistance, Eric Long for advice, and Derek Middleton for use of genotype data. This study was supported by U.S. National Institutes of Health grants AI17892 (PP, PJN, NNG) and GM109030 (JAH, PJN, PP), the Medical Research Council of the UK (JAT), and by the Victoria University of Wellington, NZ, and Ministry of Higher Education, Malaysia.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
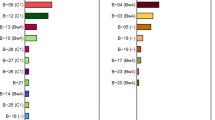
Potential interactions of Māori and Polynesian HLA-A, HLA-B, and HLA-C with KIR. Shown in the two columns on the right of each panel are the frequencies of HLA-C (panel a), HLA-A (panel b), and HLA-B (panel c) allotypes in the Māori and Polynesians. In the central columns are shown the KIR that can recognize each allotype (PDF 15 kb)
Supplementary Figure 2
KIR allele frequencies in Māori and Polynesians. Shown are the KIR alleles and their frequencies in the Māori and Polynesian populations. KIR allotypes that are known or predicted to be unexpressed at the cell surface are given in red. Alleles of the centromeric KIR genes are given in panel a; alleles of the telomeric KIR genes are given in panel b (PDF 19 kb)
Supplementary Figure 3
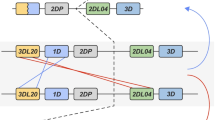
Māori and Polynesian allotype-level KIR haplotypes. In the depiction on the left are shown the Māori and Polynesian centromeric (panel a) and telomeric (panel b) KIR haplotypes with only variation that alters protein sequence or expression being taken into account. Boxes corresponding to KIR A haplotypes are colored red, and KIR B haplotypes are colored blue. In the columns on the right, the frequency of each haplotype is compared with that in Amerindian, European, and sub-Saharan African populations that have also been studied at high resolution. (Gendzekhadze et al. 2009; Norman et al. 2013; Vierra-Green et al. 2012). The Japanese population (Yawata et al. 2006) was not fully typed for centromeric KIR, so only the telomeric segment is shown (PDF 22 kb)
Rights and permissions
About this article
Cite this article
Nemat-Gorgani, N., Edinur, H.A., Hollenbach, J.A. et al. KIR diversity in Māori and Polynesians: populations in which HLA-B is not a significant KIR ligand. Immunogenetics 66, 597–611 (2014). https://doi.org/10.1007/s00251-014-0794-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-014-0794-1