Abstract
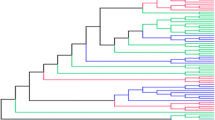
The classical class I HLA loci of humans show an excess of nonsynonymous with respect to synonymous substitutions at codons of the antigen recognition site (ARS), a hallmark of adaptive evolution. Additionally, high polymporphism, linkage disequilibrium, and disease associations suggest that one or more balancing selection regimes have acted upon these genes. However, several questions about these selective regimes remain open. First, it is unclear if stronger evidence for selection on deep timescales is due to changes in the intensity of selection over time or to a lack of power of most methods to detect selection on recent timescales. Another question concerns the functional entities which define the selected phenotype. While most analyses focus on selection acting on individual alleles, it is also plausible that phylogenetically defined groups of alleles (“lineages”) are targets of selection. To address these questions, we analyzed how \({\mathrm{d}}N/{\mathrm{d}}S\) (\(\omega\)) varies with respect to divergence times between alleles and phylogenetic placement (position of branches). We find that \(\omega\) for ARS codons of class I HLA genes increases with divergence time and is higher for inter-lineage branches. Throughout our analyses, we used non-selected codons to control for possible effects of inflation of \(\omega\) associated to intra-specific analysis, and showed that our results are not artifactual. Our findings indicate the importance of considering the timescale effect when analysing \(\omega\) over a wide spectrum of divergences. Finally, our results support the divergent allele advantage model, whereby heterozygotes with more divergent alleles have higher fitness than those carrying similar alleles.



Similar content being viewed by others
References
Albrechtsen A, Moltke I, Nielsen R (2010) Natural selection and the distribution of identity-by-descent in the human genome. Genetics 186(1):295–308
Anisimova M, Nielsen R, Yang Z (2003) Effect of recombination on the accuracy of the likelihood method for detecting positive selection at amino acid sites. Genetics 164(3):1229–36
Apps R, Qi Y, Carlson JM, Chen H, Gao X, Thomas R, Yuki Y, Del Prete GQ, Goulder P, Brumme ZL, Brumme CJ, John M, Mallal S, Nelson G, Bosch R, Heckerman D, Stein JL, Soderberg Ka, Moody MA, Denny TN, Zeng X, Fang J, Moffett A, Lifson JD, Goedert JJ, Buchbinder S, Kirk GD, Fellay J, McLaren P, Deeks SG, Pereyra F, Walker B, Michael NL, Weintrob A, Wolinsky S, Liao W, Carrington M (2013) Influence of HLA-C expression level on HIV control. Science 340(6128):87–91
Bjorkman PJ, Saper MA, Samraoui B, Bennett WS, Strominger JL, Wiley DC (1987) Structure of the human class I histocompatibility antigen, HLA-A2. Nature 329(6139):506–12
Chelvanayagam G (1996) A roadmap for HLA-A, HLA-B, and HLA-C peptide binding specificities. Immunogenetics 45(1):15–26
Dean M, Carrington M, O’Brien SJ (2002) Balanced polymorphism selected by genetic versus infectious human disease. Annu Rev Genom Hum Genet 3:263–292
Doherty PC, Zinkernagel RM (1975) Enhanced immunological surveillance in mice heterozygous at the H-2 gene complex. Nature 256(5512):50–52
Dos Reis M, Yang Z (2013) Why do more divergent sequences produce smaller nonsynonymous/synonymous rate ratios in pairwise sequence comparisons? Genetics 195(1):195–204
Felsenstein J (1989) PHYLIP-phylogeny inference package (version 3.2). Cladistics 5:164–166
Francisco RS, Buhler S, Nunes JM, Bitarello BD, França GS, Meyer D, Sanchez-Mazas A (2015) HLA supertype variation across populations: new insights into the role of natural selection in the evolution of HLA-A and HLA-B polymorphisms. Immunogenetics 67(11–12):651–663. doi:10.1007/s00251-015-0875-9
Garrigan D, Hedrick PW (2003) Detecting adaptive molecular polymorphism : lessons from the MHC. Evolution 57(8):1707–1722
Goldman N, Yang Z (1994) A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol Biol Evol 11(5):725–736
Hedrick PW (2002) Pathogen resistance and genetic variation at MHC loci. Evolution 56(10):1902–1908
Hedrick PW, Thomson G (1983) Evidence for balancing selection at HLA. Genetics 104(3):449–56
Henn B, Botigué LR, Bustamante C, Clark AG, Gravel S (2015) Estimating the mutation load in human genomes. Nat Rev Genetics 16:333–343
Hilton HG, Guethlein LA, Goyos A, Nemat-Gorgani N, Bushnell DA, Norman PJ, Parham P (2015) Polymorphic HLA-C receptors balance the functional characteristics of KIR haplotypes. J Immunol 195:3160–3170
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335(6186):167–170
Hughes AL, Nei M (1989) Nucleotide substitution at major histocompatibility complex class II loci: evidence for overdominant selection. Proc Natl Acad Sci USA 86(3):958–962
Hughes AL, Yeager M (1998) Natural selection at major histocompatibility complex of vertebrates. Annu Rev Genet 32:415–435
Huttley G, Smith MW, Carrington M, O’Brien S (1999) A scan for linkage disequilibrium accross the human genome. Genetics 152(4):1711–1722
Klein J, Sato A (2000) The HLA system first of two parts. Adv Immunol 343(10):702–709
Kryazhimskiy S, Plotkin JB (2008) The population genetics of dN/dS. PLoS Genet 4(12):10
Lenz T, Mueller B, Trillmich F, Wolf JBW (2013) Divergent allele advantage at MHC-DRB through direct and maternal genotypic effects and its consequences for allele pool composition and mating. Proc R Soc B 280:20130714
Martin DP, Lemey P, Lott M, Moulton V, Posada D, Lefeuvre P (2010) RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics 26(19):2462–3
Meyer D, Single RM, Mack SJ, Erlich HA, Thomson G (2006) Signatures of demographic history and natural selection in the human major histocompatibility complex loci. Genetics 173(4):2121–2142
Meyer D, Thomson G (2001) How selection shapes variation of the human major histocompatibility complex: a review. Ann Hum Genet 65(1):1–26
Mugal CF, Wolf JBW, Kaj I (2014) Why time matters: codon evolution and the temporal dynamics of dN/dS. Mol Biol Evol 31(1):212–231
Penn DJ, Damjanovich K, Potts WK (2002) MHC heterozygosity confers a selective advantage against multiple-strain infections. Proc Natl Acad Sci USA 99(17):260–264
Pond SLK, Frost SDW, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21(5):676–679
Prugnolle F, Manica A, Charpentier M, Guégan JF, Guernier V, Balloux F (2005) Pathogen-driven selection and worldwide HLA class I diversity. Curr Biol 15(11):1022–7
Richman A (2000) Evolution of balanced genetic polymorphism. Mol Ecol 9(12):1953–63
Robinson J, Halliwell JA, McWilliam H, Lopez R, Parham P, Marsh SGE (2013) The IMGT/HLA database. Nucleic Acids Res 41:D1222–D1227
Rocha EPC, Smith JM, Hurst LD, Holden MTG, Cooper JE, Smith NH, Feil EJ (2006) Comparisons of dN/dS are time dependent for closely related bacterial genomes. J Theor Biol 239(2):226–235
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sidney J, Grey HM, Kubo RT, Sette A (1996) Practical, biochemical and evolutionary implications of the discovery of HLA class I supermotifs. Immunol Today 17(6):261–6
Single RM, Martin MP, Gao X, Meyer D, Yeager M, Kidd JR, Kidd K, Carrington M (2007) Global diversity and evidence for coevolution of KIR and HLA. Nat Genet 9:1114–1119
Slade R, McCallum H (1992) Overdominant vs. frequency-dependent selection at MHC loci. Genetics 132:861–864
Spurgin LG, Richardson DS (2010) How pathogens drive genetic diversity: MHC, mechanisms and misunderstandings. Proc Biol Sci 277(1684):979–88
Stolestki N, Eyre-Walker A (2011) The positive correlation between dN/dS and dS in mammals is due to runs of adjacent substitutions. Mol Biol Evol 28(4):1371–1380
Takahata N, Nei M (1990) Allelic genealogy under overdominant and frequency-dependent selection and polymorphism of major histocompatibility complex loci. Genetics 124(4):967–978
Takahata N, Satta Y (1998) Footprints of intragenic recombination at HLA loci. Immunogenetics 47(6):430–441
Templeton AR (1996) Contingency tests of neutrality using intra/interspecific gene trees: the rejection of neutrality for the evolution of the mitochondrial Cytochrome Oxidase II gene in the hominoid primates. Genetics 144(3):1263–1270
The 1000 Genomes Project Consortium (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65
Wakeland EK, Boehme S, She JX, Cc Lu, Mclndoe RA, Cheng I, Ye Y, Potts WK (1990) Ancestral polymorphisms of MHC class II genes : divergent allele advantage. Immunol Res 9:115–122
Wolf JBW, Künstner A, Nam K, Jakobsson M, Ellegren H (2009) Nonlinear dynamics of nonsynonymous (dN) and synonymous (dS) substitution rates affects inference of selection. Genome Biol Evol 1:308–319
Yang Z (2006) Computational molecular evolution. Oxford University Press, Oxford
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24(8):1586–1591
Yang Z, Swanson WJ (2002) Codon-substitution models to detect adaptive evolution that account for heterogeneous selective pressures among site classes. Mol Biol Evol 19(1):49–57
Yang Z, Wong WSW, Nielsen R (2005) Bayes empirical bayes inference of amino acid sites under positive selection. Mol Biol Evol 22(4):1107–1118
Yasukochi Y, Satta Y (2014) Nonsynonymous substitution rate heterogeneity in the peptide-binding region among different HLA-DRB1 lineages in humans. G3 (Bethesda) 4:1217–1226
Acknowledgments
The authors thank Kelly Nunes for thoughtful comments on the manuscript, Richard Single for comments on the statistical aspects of this work, Aida M. Andrés for general comments and Débora Y.C.Brandt for help with the 1000 Genomes data sets. BDB was funded by grants #08/56502-6 and #11/12500-2, São Paulo Research Foundation (FAPESP), and #152676/2011-2, Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). RSF was funded by grant #142130/2009-5 (CNPq) and DM was funded by grants #09/09127-8 (FAPESP) and #308960/2009-2 (CNPq).
Author’s Contributions
BDB participated in the design of the study, performed analyses, discussed results and drafted the manuscript. RDF performed analyses to detect recombinants and contributed to discussions. DM conceived of the study, participated in its design and discussion and in the drafting of the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bitarello, B.D., Francisco, R.d. & Meyer, D. Heterogeneity of \({\varvec{\rm d}}{\varvec{N}}/{\varvec{\rm d}}{\varvec{S}}\) Ratios at the Classical HLA Class I Genes over Divergence Time and Across the Allelic Phylogeny. J Mol Evol 82, 38–50 (2016). https://doi.org/10.1007/s00239-015-9713-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-015-9713-9