Abstract
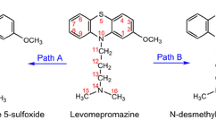
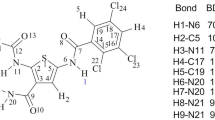
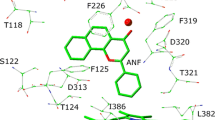
Chlorpromazine, belonging to the first-generation antipsychotics, is known to cause some side effects, such as hepatotoxicity and agranulocytosis. The metabolic mechanisms of chlorpromazine remain elusive up to now, but are thought to result in the formation of some reactive metabolites having side effects on the parent drug. The goal of this work was to explore the metabolic mechanisms of chlorpromazine catalyzed by cytochrome P450 isoenzyme 1A2, a highly important activating enzyme of cytochrome P450 family, using DFT calculation. Three types of metabolic mechanisms were characterized, including S-oxidation, aromatic hydroxylation and N-dealkylation. The calculated results demonstrate that N 14-demethylation is the most thermodynamically and kinetically favorable metabolic pathway of chlorpromazine, followed by S5-oxidation. Then, mono-N-desmethylchlorpromazine is the most feasible chlorpromazine metabolite, which can occur further demethylation to form di-N-desmethylchlorpromazine. Besides, chlorpromazine 5-sulfoxide and 7-hydroxychlorpromazine are both the possible metabolites of chlorpromazine. Interestingly, N-methyl hydroxylation, the rate-limiting step of N-demethylation, proceeds predominantly via a single-electron-transfer mechanism. All the proton transfer processes involved in the aromatic hydroxylation and N-dealkylation prefer to occurrence in a water-assisted enzymatic process. Each metabolic pathway proceeds in the spin-selective manner via the low-spin state of Cpd I. Our results are in good accordance with the experimental observations, which can provide some essential implications for the metabolic mechanisms of chlorpromazine-like drugs.














Similar content being viewed by others
References
Liu X, De Haan S (2009) Cochrane Database Syst Rev 2:CD007778
Anthérieu S, Bachour-El Azzi P, Dumont J, Abdel-Razzak Z, Guguen-Guillouzo C, Fromenty B, Robin MA, Guillouzo A (2013) Hepatology 57:1518–1529
Tohen M, Vieta E (2009) Bipolar Disord 2:45–54
Morak-Młodawska B, Jeleń M (2007) Pol Merkur Lek 23:459–461
Shin SY, Kim CG, Kim SH, Kim YS, Lim Y, Lee YH (2010) Exp Mol Med 42:395–405
Liperoti R, Pedone C, Corsonello A (2008) Curr Neuropharmacol 6:117–124
Drucker AM, Rosen CF (2011) Drug Saf 34:821–837
Lasic D, Cvitanovic MZ, Uglešic B, Višic V, Hlevnjak I (2011) Psychiatr Danub 23:194–197
Shahzad S, Suleman MI, Shahab H, Mazour I, Kaur A, Rudzinskiy P, Lippmann S (2002) Psychosomatics 43:354–359
Subashini K, Rao VA (2004) Indian J Pharmacol 36:323–324
Toler SM (2004) Exp Biol Med (Maywood) 229:607–615
Wójcikowski J, Boksa J, Daniel WA (2010) Biochem Pharmacol 80:1252–1259
Daniel W (1995) Pol J Pharmacol 47:367–379
Chetty M, Pillay VL, Moodley SV, Miller R (1996) Eur Neuropsychopharmacol 2:85–91
Chetty M, Gouws E, Miller R, Moodley SV (1999) Eur Neuropsychopharmacol 9(1–2):77–82
Abernathy CO, Lukacs L, Zimmerman HJ (1977) Proc Soc Exp Biol Med 155:474–478
Tavoloni N, Boyer JL (1980) J Pharmacol Exp Ther 214:269–274
Wójcikowski J, Pichard-Garcia L, Maurel P, Daniel WA (2003) Br J Pharmacol 138:1465–1474
Wójcikowski J, Pichard-Garcia L, Maurel P, Daniel WA (2004) Eur Neuropsychopharmacol 14:199–208
Wójcikowski J, Maurel P, Daniel WA (2006) Drug Metab Dispos 34:471–476
Chetty M, Miller R, Moodley SV (1994) Eur J Clin Pharmacol 46:523–526
Kot M, Daniel WA (2008) Biochem Pharmacol 76:543–551
Blomberg MRA, Borowski T, Himo F, Liao RZ, Siegbahn PEM (2014) Chem Rev 114:3601–3658
Schrǒder D, Shaik S, Schwarz H (2000) Acc Chem Res 33:139–145
Baciocchi EBM, Gerini MF, Lanzalunga O (2005) J Org Chem 70:5144
Guengerich FP, Yun CH, Macdonald TL (1996) J Biol Chem 271:27321–27329
Jurva U, Bissel P, Isin EM, Igarashi K, Kuttab S, Castagnoli N (2005) J Am Chem Soc 127:12368–12377
Li CS, Wu W, Kumar D, Shaik S (2006) J Am Chem Soc 128:394–395
Chen H, de Groot MJ, Vermeulen NPE, Hanzlik RP (1997) J Org Chem 62:8227–8230
Chen ZQ, Kang Y, Zhang CH, Tao J, Xue Y (2015) Theor Chem Acc 134:110
Shaik S, Cohen S, Wang Y, Chen H, Kumar D, Thiel W (2010) Chem Rev 110:949–1017
Sansen S, Yano JK, Reynald RL, Schoch GA, Griffin KJ, Stout CD, Johnson EF (2007) J Biol Chem 282:14348–14355
Tao J, Kang Y, Xue ZY, Wang YT, Zhang Y, Chen Q, Chen ZQ, Xue Y (2015) J Mol Graph Model 61:123–132
Kwiecien RA, Molinié R, Paneth P, Silvestre V, Lebreton J, Robins RJ (2011) Arch Biochem Biophys 510:35–41
Li DM, Wang Y, Yang CL, Han KL (2009) Dalton Trans 14:291–297
Schyman PUD, Wang Y, Shaik S (2010) J Phys Chem B 114:7078–7089
Kang Y, Tao J, Xue ZY, Zhang Y, Chen ZQ, Xue Y (2016) Tetrahedron 72:2858–2867
Zhang Q, Bell R, Truong TN (1995) J Phys Chem 99:592–599
Reed AE, Schleyer PR (1990) J Am Chem Soc 112:1434–1445
Mennucci B (2012) WIREs Comput Mol Sci 2:386–404
Schutz CN, Warshel A (2001) Proteins 44:400–417
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JJA, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam NJ, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas O, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2013) Gaussian 09 revision D01. Gaussian Inc, Wallingford
Bach RD, Dmitrenko O (2003) J Phys Chem B 107:12851–12861
Wang Y, Kumar D, Yang CL, Han KL, Shaik S (2007) J Phys Chem B 111:7700–7710
Acknowledgments
This work was supported by grants from National Natural Science Foundation of China (Grant No. 21203153), Science and Technology Department of Sichuan Province (Grant No. 2011JY0136) and Department of Education of Sichuan Province (Grant No. 12ZA174) and China West Normal University (Grant No. 11B002).
Author information
Authors and Affiliations
Corresponding author
Additional information
Zhiyu Xue and Yan Zhang authors have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xue, Z., Zhang, Y., Tao, J. et al. Theoretical elucidation of the metabolic mechanisms of phenothiazine neuroleptic chlorpromazine catalyzed by cytochrome P450 isoenzyme 1A2. Theor Chem Acc 135, 218 (2016). https://doi.org/10.1007/s00214-016-1943-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00214-016-1943-4