Abstract
Key message
This study provides high-quality variation data of diverse radish genotypes. Genome-wide SNP comparison along with RNA-seq analysis identified candidate genes related to domestication that have potential as trait-related markers for genetics and breeding of radish.
Abstract
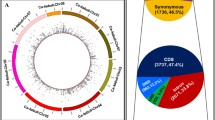
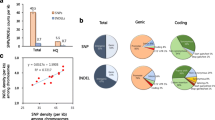
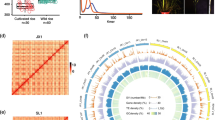
Radish (Raphanus sativus L.) is an annual root vegetable crop that also encompasses diverse wild species. Radish has a long history of domestication, but the origins and selective sweep of cultivated radishes remain controversial. Here, we present comprehensive whole-genome resequencing analysis of radish to explore genomic variation between the radish genotypes and to identify genetic bottlenecks due to domestication in Asian cultivars. High-depth resequencing and multi-sample genoty** analysis of ten cultivated and seven wild accessions obtained 4.0 million high-quality homozygous single-nucleotide polymorphisms (SNPs)/insertions or deletions. Variation analysis revealed that Asian cultivated radish types are closely related to wild Asian accessions, but are distinct from European/American cultivated radishes, supporting the notion that Asian cultivars were domesticated from wild Asian genotypes. SNP comparison between Asian genotypes identified 153 candidate domestication regions (CDRs) containing 512 genes. Network analysis of the genes in CDRs functioning in plant signaling pathways and biochemical processes identified group of genes related to root architecture, cell wall, sugar metabolism, and glucosinolate biosynthesis. Expression profiling of the genes during root development suggested that domestication-related selective advantages included a main taproot with few branched lateral roots, reduced cell wall rigidity and favorable taste. Overall, this study provides evolutionary insights into domestication-related genetic selection in radish as well as identification of gene candidates with the potential to act as trait-related markers for background selection of elite lines in molecular breeding.







Similar content being viewed by others
References
Abbo S, Pinhasi van-Oss R, Gopher A, Saranga Y, Ofner I, Peleg Z (2014) Plant domestication versus crop evolution: a conceptual framework for cereals and grain legumes. Trends Plant Sci 19:351–360
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Baute G, Kane N, Grassa C, Lai Z, Rieseberg L (2015) Genome scans reveal candidate domestication and improvement genes in cultivated sunflower, as well as post-domestication introgression with wild relatives. New Phytol 206:830–838
Becker B (1962) Rettich und Radies. Handb Pflanzenzücht 6:23–78
Browning S, Browning B (2007) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81:1084–1097
Bruex A, Kainkaryam R, Wieckowski Y, Kang Y, Bernhardt C, **a Y, Zheng X, Wang J, Lee M, Benfey P, Woolf P, Schiefelbein J (2012) A gene regulatory network for root epidermis cell differentiation in Arabidopsis. PLoS Genet 8:e1002446
Bus A, Hecht J, Huettel B, Reinhardt R, Stich B (2012) High-throughput polymorphism detection and genoty** in Brassica napus using next-generation RAD sequencing. BMC Genom 13:281
Cao K, Zheng Z, Wang L, Liu X, Zhu G, Fang W, Cheng S, Zeng P, Chen C, Wang X, **e M, Zhong X, Wang X, Zhao P, Bian C, Zhu Y, Zhang J, Ma G, Chen C, Li Y, Hao F, Li Y, Huang G, Li Y, Li H, Guo J, Xu X, Wang J (2014) Comparative population genomics reveals the domestication history of the peach, Prunus persica, and human influences on perennial fruit crops. Genome Biol 15:415
Chen H, Patterson N, Reich D (2010) Population differentiation as a test for selective sweeps. Genome Res 20:393–402
Chung W-H, Jeong N, Kim J, Lee W, Lee Y-G, Lee S-H, Yoon W, Kim J-H, Choi I-Y, Choi H-K, Moon J-K, Kim N, Jeong S-C (2014) Population structure and domestication revealed by high-depth resequencing of Korean cultivated and wild soybean genomes. DNA Res 21:153–167
Cingolani P, Platts A, Wang IL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6:80–92
Ellstrand N, Marshall D (1985) The impact of domestication on distribution of allozyme variation within and among cultivars of radish, Raphanus sativus L. Theor Appl Genet 69:393–398
Firon N, LaBonte D, Villordon A, Kfir Y, Solis J, Lapis E, Perlman T, Doron-Faigenboim A, Hetzroni A, Althan L, Nadir L (2013) Transcriptional profiling of sweet potato (Ipomoea batatas) roots indicates down-regulation of lignin biosynthesis and up-regulation of starch biosynthesis at an early stage of storage root formation. BMC Genom 14:460
George R, Evans D (1981) A classification of winter radish cultivars. Euphytica 30:483–492
Guevara D, El-Kereamy A, Yaish M, Mei-Bi Y, Rothstein S (2014) Functional characterization of the rice UDP-glucose 4-epimerase 1, OsUGE1: a potential role in cell wall carbohydrate partitioning during limiting nitrogen conditions. PLoS One 9:e96158
Hu Y, Zhong R, Wr Morrison, Ye Z (2003) The Arabidopsis RHD3 gene is required for cell wall biosynthesis and actin organization. Planta 217:912–921
Huang X, Lu T, Han B (2013) Resequencing rice genomes: an emerging new era of rice genomics. Trends Genet 29:225–232
Hufford M, Xu X, van Heerwaarden J, Pyhäjärvi T, Chia J, Cartwright R, Elshire R, Glaubitz J, Guill K, Kaeppler S, Lai J, Morrell P, Shannon L, Song C, Springer N, Swanson-Wagner R, Tiffin P, Wang J, Zhang G, Doebley J, McMullen M, Ware D, Buckler E, Yang S, Ross-Ibarra J (2012) Comparative population genomics of maize domestication and improvement. Nat Genet 44:808–811
Iorizzo M, Senalik D, Ellison S, Grzebelus D, Cavagnaro P, Allender C, Brunet J, Spooner D, Van Deynze A, Simon P (2013) Genetic structure and domestication of carrot (Daucus carota subsp. sativus) (Apiaceae). Am J Bot 100:930–938
Jang G, Lee J, Rastogi K, Park S, Oh S, Lee J (2015) Cytokinin-dependent secondary growth determines root biomass in radish (Raphanus sativus L.). J Exp Bot 66:4607–4619
Jeong Y-M, Kim N, Ahn B, Oh M, Chung W-H, Chung H, Jeong S, Lim K-B, Hwang Y-J, Kim G-B, Baek S, Choi S-B, Hyung D-J, Lee S-W, Sohn S-H, Kwon S-J, ** M, Seol Y-J, Chae W, Choi K, Park B-S, Yu H-J, Mun J-H (2016) Elucidating the triplicated ancestral genome structure of radish based on chromosome-level comparison with the Brassica genomes. Theor Appl Genet 129:1357–1372
Kaneko Y, Matsuzawa Y (1993) Radish (Raphanus sativus L.). Pergamon Press Ltd., Oxford
Kerk N, Ceserani T, Tausta S, Sussex I, Nelson T (2003) Laser capture microdissection of cells from plant tissues. Plant Physiol 132:27–35
Kitashiba H, Li F, Hirakawa H, Kawanabe T, Zou Z, Hasegawa Y, Tonosaki K, Shirasawa S, Fukushima A, Yokoi S, Takahata Y, Kakizaki T, Ishida M, Okamoto S, Sakamoto K, Shirasawa K, Tabata S, Nishio T (2014) Draft sequences of the radish (Raphanus sativus L.) genome. DNA Res 21:481–490
Kloosterman B, Vorst O, Hall R, Visser R, Bachem C (2005) Tuber on a chip: differential gene expression during potato tuber development. Plant Biotechnol J 3:505–519
Kopta T, Pokluda R (2013) Yields, quality and nutritional parameters of radish (Raphanus sativus) cultivars when grown in organically in Czech Republic. Hort Sci 40:16–21
Langmead B, Salzberg S (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Li X, Zeng R, Liao H (2016) Improving crop nutrient efficiency through root architecture modifications. J Integr Plant Biol 58:193–202
Maldonado Dos Santos J, Valliyodan B, Joshi T, Khan S, Liu Y, Wang J, Vuong T, Oliveira M, Marcelino-Guimarães F, Xu D, Nguyen H, Abdelnoor R (2016) Evaluation of genetic variation among Brazilian soybean cultivars through genome resequencing. BMC Genom 17:110
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo M (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Mitsui Y, Shimomura M, Komatsu K, Namiki N, Shibata-Hatta M, Imai M, Katayose Y, Mukai Y, Kanamori H, Kurita K, Kagami T, Wakatsuki A, Ohyanagi H, Ikawa H, Minaka N, Nakagawa K, Shiwa Y, Sasaki T (2015) The radish genome and comprehensive gene expression profile of tuberous root formation and development. Sci Rep 5:10835
Moghe G, Hufnagel D, Tang H, **ao Y, Dworkin I, Town C, Conner J, Shiu S (2014) Consequences of whole-genome triplication as revealed by comparative genomic analyses of the wild radish Raphanus raphanistrum and three other Brassicaceae species. Plant Cell 26:1925–1937
Mun J-H, Chung H, Chung W-H, Oh M, Jeong Y-M, Kim N, Ahn B, Park B-S, Park S, Lim K-B, Hwang Y-J, Yu H-J (2015) Construction of a reference genetic map of Raphanus sativus based on genoty** by whole-genome resequencing. Theor Appl Genet 128:259–272
Olsen K, Wendel J (2013) A bountiful harvest: genomic insights into crop domestication phenotypes. Annu Rev Plant Biol 64:47–70
Paradis E, Claude J, Strimmer K (2004) APE: analysis of phylogenetics and evolution in R language. Bioinformatics 20:289–290
Park S, Yu H-J, Mun J-H, Lee S (2010) Genome-wide discovery of DNA polymorphism in Brassica rapa. Mol Genet Genomics 283:135–145
Passardi F, Penel C, Dunand C (2004) Performing the paradoxical: how plant peroxidases modify the cell wall. Trends Plant Sci 9:534–540
Patel R, Jain M (2012) NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS One 7:e30619
Ralph P, Coop G (2010) Parallel adaptation: one or many waves of advance of an advantageous allele? Genetics 186:647–668
Remans T, Nacry P, Pervent M, Girin T, Tillard P, Lepetit M, Gojon A (2006) A central role for the nitrate transporter NRT2.1 in the integrated morphological and physiological responses of the root system to nitrogen limitation in Arabidopsis. Plant Physiol 140:909–921
Rong J, Lammers Y, Strasburg J, Schidlo N, Ariyurek Y, de Jong T, Klinkhamer P, Smulders M, Vrieling K (2014) New insights into domestication of carrot from root transcriptome analyses. BMC Genom 15:895
Rouhier H, Usuda H (2001) Spatial and temporal distribution of sucrose synthase in the radish hypocotyl in relation to thickening growth. Plant Cell Physiol 42:583–593
Ruzin S (1999) Plant microtechnique and microscopy. Oxford University Press, New York
Seifert G, Barber C, Wells B, Dolan L, Roberts K (2002) Galactose biosynthesis in Arabidopsis: genetic evidence for substrate channeling from UDP-D-galactose into cell wall polymers. Curr Biol 12:1840–1845
Sergeeva L, Keurentjes J, Bentsink L, Vonk J, van der Plas L, Koornneef M, Vreugdenhil D (2006) Vacuolar invertase regulates elongation of Arabidopsis thaliana roots as revealed by QTL and mutant analysis. Proc Natl Acad Sci USA 103:2994–2999
Sirju-Charran G, Wickham L (1988) The development of alternative storage sink sites in sweet potato Ipomoea batatas. Ann Bot 61:99–102
Sturm A, Tang G (1999) The sucrose-cleaving enzymes of plants are crucial for development, growth and carbon partitioning. Trends Plant Sci 4:401–407
Sun P, **ao X, Duan L, Guo Y, Qi J, Liao D, Zhao C, Liu Y, Zhou L, Li X (2015) Dynamic transcriptional profiling provides insights into tuberous root development in Rehmannia glutinosa. Front Plant Sci 6:396
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Ting F, Wren M (1980) Storage organ development in radish (Raphanus sativus L.). 1. A comparison of development in seedlings and rooted cuttings of two contrasting varieties. Ann Bot 46:267–276
Usuda H, Demura T, Shimogawara K, Fukuda H (1999) Development of sink capacity of the “storage root” in a radish cultivar with a high ratio of “storage root” to shoot. Plant Cell Physiol 40:369–377
Warwick S (2011) Brassicaceae in Agriculture. In: Schmidt R, Bancroft I (eds) Genetics and Genomics of the Brassicaceae. Springer, Gatersleben, pp 33–66
Xu M, **e W, Huang M (2012a) Overexpression of PeRHD3 alters the root architecture in Populus. Biochem Biophys Res Commun 424:239–244
Xu X, Liu X, Ge S, Jensen J, Hu F, Li X, Dong Y, Gutenkunst R, Fang L, Huang L, Li J, He W, Zhang G, Zheng X, Zhang F, Li Y, Yu C, Kristiansen K, Zhang X, Wang J, Wright M, McCouch S, Nielsen R, Wang J, Wang W (2012b) Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat Biotechnol 30:105–111
Yamagishi H (2004) Assessment of cytoplasmic polymorphisms by PCR-RFLP of the mitochondrial orfB region in wild and cultivated radishes (Raphanus). Plant Breed 123:141–144
Yamagishi H, Terachi T (2003) Multiple origins of cultivated radishes as evidenced by a comparison of the structural variations in mitochondrial DNA of Raphanus. Genome 46:89–94
Yamane K, Lu N, Ohnishi O (2005) Chloroplast DNA variations of cultivated radish and its wild relatives. Plant Sci 168:627–634
Yamane K, Lu N, Ohnishi O (2009) Multiple origins and high genetic diversity of cultivated radish inferred from polymorphism in chloroplast simple sequence repeats. Breed Sci 59:55–65
Zheng D, Han X, An Y, Guo H, **a X, Yin W (2013) The nitrate transporter NRT2.1 functions in the ethylene response to nitrate deficiency in Arabidopsis. Plant Cell Environ 36:1328–1337
Zhou Z, Jiang Y, Wang Z, Gou Z, Lyu J, Li W, Yu Y, Shu L, Zhao Y, Ma Y, Fang C, Shen Y, Liu T, Li C, Li Q, Wu M, Wang M, Wu Y, Dong Y, Wan W, Wang X, Ding Z, Gao Y, **ang H, Zhu B, Lee S, Wang W, Tian Z (2015) Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat Biotechnol 33:408–414
Acknowledgments
This work was supported by grants from the Next-Generation Biogreen21 program (PJ01108601 to JHM and PJ01108602 to HJY) and the National Academy of Agricultural Science (PJ009795 to JHM), Rural Development Administration, Korea. We appreciate Genebank Information Center, RDA, Korea and National Institute of Agricultural Sciences, Japan for providing seeds of radish accessions.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical standards
The authors declare that the experiments complied with current laws of the country in which they were performed.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by I. AP Parkin.
N. Kim and Y.-M. Jeong contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, N., Jeong, YM., Jeong, S. et al. Identification of candidate domestication regions in the radish genome based on high-depth resequencing analysis of 17 genotypes. Theor Appl Genet 129, 1797–1814 (2016). https://doi.org/10.1007/s00122-016-2741-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2741-z