Abstract
Histone modifying enzymes play critical roles in many key cellular processes and are appealing proteins for targeting by small molecules in disease. However, while the functions of histone modifying enzymes are often linked to epigenetic regulation of the genome, an emerging theme is that these enzymes often also act by non-catalytic and/or non-epigenetic mechanisms. SETD2 (Set2 in yeast) is best known for associating with the transcription machinery and methylating histone H3 on lysine 36 (H3K36) during transcription. This well-characterized molecular function of SETD2 plays a role in fine-tuning transcription, maintaining chromatin integrity, and mRNA processing. Here we give an overview of the various molecular functions and mechanisms of regulation of H3K36 methylation by Set2/SETD2. These fundamental insights are important to understand SETD2’s role in disease, most notably in cancer in which SETD2 is frequently inactivated. SETD2 also methylates non-histone substrates such as α-tubulin which may promote genome stability and contribute to the tumor-suppressor function of SETD2. Thus, to understand its role in disease, it is important to understand and dissect the multiple roles of SETD2 within the cell. In this review we discuss how histone methylation by Set2/SETD2 has led the way in connecting histone modifications in active regions of the genome to chromatin functions and how SETD2 is leading the way to showing that we also have to look beyond histones to truly understand the physiological role of an ‘epigenetic’ writer enzyme in normal cells and in disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
All molecular processes in the eukaryotic cell that involve transactions with genomic DNA have to deal with chromatin—the complex of DNA and histone proteins. In general, wrap** DNA around histone octamers to form nucleosomes acts as a barrier for processes that require DNA as a template such as transcription, replication and DNA repair. Indeed, histones need to be removed at least temporarily to allow access to the DNA for these processes (for reviews see [72, 131, 153]). At the same time, nucleosomes also act as an important docking platform for a myriad of factors that regulate DNA transactions. A critical layer of control for these processes is the post-translational modification of histone proteins. Histone post-translational modifications (PTMs) can directly influence the structure of chromatin, for example by neutralizing the positive charge of histone proteins, or act as a docking site for so called chromatin ‘reader’ proteins (for reviews see [24, 141, 176, 223]). Already early on it has been hypothesized that histone PTMs directly control chromatin processes and that a specific combination of histone PTMs can be viewed as a ‘code’ that specifies the function of a DNA region [177]. In the past decade, genome-wide maps for many histone PTMs in different cell types have been generated, which has indeed confirmed that specific histone PTMs often correlate with a DNA element that is in a particular state (i.e. active promoter, active enhancer, site of DNA damage) [10, 174]. However, correlation does not necessarily mean causation. One of the objectives in understanding chromatin is therefore to determine the functional roles of histone PTMs.
A large part of our understanding of chromatin modifying (‘writer’) enzymes comes from research on histone methyltransferases in budding yeast. In Saccharomyces cerevisiae, the SET domain-containing protein Set2 methylates histone H3 on lysine 36 (H3K36). This site is located at the base of the N-terminal tail of H3 and can be either mono-, di-, or trimethylated by Set2 [178]. H3K36 methylation was one of the first PTMs for which a clear function was found in transcription. Initial reports found Set2 to be enriched on the coding sequences of active genes suggesting a role in transcription elongation [164, 130]. In the absence of SETD2/H3K36me3, MORF4L1 might still be able to target PTB to exons by binding to H3K36me2, given its in vitro binding properties. However, because H3K36me2 is only enriched at the 5’ end of active gene bodies in humans [57], H3K36me2 is most likely not sufficient to promote PTB localization to the same target exons as H3K36me3.
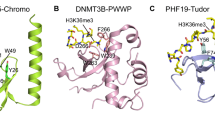
Another factor that influences splicing through SETD2 activity is Zinc Finger MYND-Type Containing 11 (ZMYND11), which is a chromatin reader protein that has been reported to associate with spliceosome components and regulate intron retention [70]. ZMYND11 binds specifically to the replication-independent ‘gap-filler’ histone variant H3.3 when trimethylated at K36 (H3.3K36me3) through its tandem PHD-, bromo-, and PWWP (PBP) domain, which specifically recognizes both H3K36me3 as well as the S31 residue that is unique to H3.3 compared to canonical H3 [70, 209, 210, 221]. H3.3K36M also inhibits the H3K36 dimethyltransferase NSD2 (also known as MMSET) in a manner analogous to SETD2 and therefore also affects global H3K36me2 levels [57].
In chondroblastoma, H3.3K36M has been reported to contribute to tumor development by promoting colony formation, and inhibiting apoptosis and chondrocyte differentiation [57, 128]. Interestingly, H3.3K36M also leads to a redistribution of H3K27me3 away from developmentally silenced genes to regions normally enriched in H3K36me3, which may contribute to the derepression of PRC2 target genes that prevent differentiation [128]. In vitro, H3K36me3 nucleosomes are a poor substrate for PRC2 [216], providing a possible explanation for this redistribution of H3K27me3 in cells lacking H3K36me3. PRC2-mediated silencing is not strongly impaired in Drosophila H3K36R cells [140] so future studies are required to determine if defective PRC2-mediated gene repression is a general feature of cells lacking H3K36me3.
Other mutations that have been found in H3.3 (predominantly in H3F3A) are H3.3G34R/V in osteosarcoma [8] and glioblastoma [165, 202], [179], and H3.3G34W/L in giant cell tumor of the bone (GCTB; [8]. Unlike H3.3K36M, these H3.3G34 mutations do not affect global H3K36me3 levels and only inhibit SETD2 in cis [111]. H3.3G34 is involved in the binding of SETD2 to H3, fitting in a small pocket in the SET domain, and any substitution with a bulky amino acid residue blocks the interaction [209, 210, 221]. It is currently not clear how H3.3G34 mutations are mechanistically involved in tumor development. A similarity between H3.3K36M and H3.3G34 mutations is that they tend to occur in tumors found in children and young adults. In glioblastoma, H3.3G34R is associated with a developmental expression signature that includes genes that block differentiation (such as self-renewal genes; [18], which is reminiscent of H3.3K36M-mutant chondroblastoma [128]. A common theme might therefore be that H3.3 mutations that affect SETD2 function maintain precursor cells in a pluripotent state and block differentiation during development, although the exact mechanisms behind this process are likely different given that global H3K36me3 levels are unaltered in H3.3G34 mutant cells. In addition, H3.3G34W cells derived from GCTB patients were reported to have mRNA splicing defects which may contribute to tumorigenesis [121]. Expression of H3.3G34R (but not H3.3G34V), as well as H3.3K36M, also impairs DNA repair through HR [127, 154, 208] indicating that genomic instability might be a shared pathway contributing to tumorigenesis in H3.3 mutant and SETD2 deficient cells.
To summarize, mutations in H3.3 genes and SETD2 are found in distinct cancer types and both affect SETD2 activity but can do so in distinct manners (Fig. 6). Besides H3K36me3, SETD2 loss affects the methylation of non-histone substrates of SETD2, whereas H3.3 mutations can also affect other modification states of H3K36 (such as H3K36ac, H3K36me1, and -me2) either by directly preventing the modification or by in-trans inhibition of other writers such as NSD2.
Connection between mutations in SETD2 and histone H3.3 affecting H3K36 methylation. A SETD2 allele can be lost by one-copy deletion of the short arm of chromosome 3, which is a frequent event in ccRCC. Mono-allelic loss of SETD2 does not appear to affect global H3K36me3 levels in ccRCC indicating that SETD2 is haplo-sufficient for H3K36me3. However, α-tubulin methylation on K40 is lost upon mono-allelic SETD2 inactivation suggesting that SETD2-mediated maintenance of genomic stability through tubulin methylation might be frequently perturbed in ccRCC. Mutations in H3.3 (found in chondroblastoma, brain tumors and osteosarcoma among others) can either inhibit SETD2 in-cis (H3.3G34R/V) or in-trans (H3.3K36M). It is currently unknown to what extent H3.3K36M affects the methylation of non-histone substrates of SETD2 such as α-tubulin
Overexpression of the H3K9me3/H3K36me3-demethylase KDM4A leads to genome instability
SETD2’s function can also be affected in cancer through misregulated expression of the demethylase KDM4A [69], which demethylates both H3K36me3 and H3K9me3 [100]. KDM4A is either deleted or overexpressed (predominantly through gene amplification) in several types of cancer including lung, breast, ovarian, and head and neck cancer [11, 20, 133]. KDM4A promotes S-phase progression and regulates replication timing [19, 20] and its function in cancer is best understood in the context of its overexpression (for a detailed review please see [107, 214]. Interestingly, KDM4A overexpression results in the (extrachromosomal) amplification of chromosome 1q12, through site-specific re-replication during a single cell cycle, and 1q12 amplification also correlates with KDM4A overexpression in tumor samples [20]. Chromosome 1q12 gain mediated by KDM4A overexpression depends on the catalytic activity of KDM4A, and can also be induced by expressing either H3.3K9M or H3.3K36M [20]. This suggests that both H3K36me3 and H3K9me3 prevent 1q12 re-replication during S-phase, although the exact mechanism remains to be determined. KDM4A also has a negative role in DNA repair, inhibiting the recruitment of 53BP1 to DNA damage sites, but this role is independent of its catalytic activity [132] suggesting it is not related to SETD2’s positive role in DNA repair. It is currently unknown if KDM4A further contributes to tumorigenesis by antagonizing SETD2’s role in promoting mRNA processing. Furthermore, it remains to be determined if KDM4A can demethylate non-histone substrates of SETD2 such as α-tubulin, which might be an additional pathway through which KDM4A negatively regulates genome stability. Thus, even though it is clear that both SETD2 loss and KDM4A overexpression perturb H3K36me3 levels, it is not yet entirely clear how much overlap there is in the mechanisms contributing to tumorigenesis in SETD2 deficient versus KDM4A overexpressing tumors.
Conclusion
The SETD2/Set2 enzymes have been a prime example of how different approaches in different model systems have led the way in unravelling the molecular role and regulation of a chromatin modifying system associated with RNA polymerase moving along genes. SETD2/Set2 is a key enzyme in the cell involved in a broad range of genome-associated processes. The studies on SETD2/Set2 and H3K36 methylation showcase that teasing apart the various functions requires perturbing not only the writer itself, but also the other domains of the enzymes, the opposing demethylation activity, and the substrate lysine. Moving beyond chromatin, the story of SETD2 emphasizes the importance of knowledge about non-histone substrates of so called ‘epigenetic writers’. An emerging theme in chromatin biology is that non-catalytic functions or activities towards non-histone substrates of epigenetic enzymes need to be considered to fully understand the physiological role of these enzymes. Looking forward, it will be important to develop tools to identify substrates of SETD2 in different cellular contexts in an unbiased way and to perturb SETD2 functions in a substrate-specific manner, e.g. by mutation of a substrate lysine, or by isolation of separation-of-function mutations. With the current advances in genome engineering and proteomics it can be expected that more SETD2 surprises will be discovered and that the function of SETD2 in normal cells and in disease will be further unraveled at a molecular level.
Availability of data and material
Not applicable.
References
Ahmad K, Henikoff S (2002) The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol Cell 9(6):1191–1200. https://doi.org/10.1016/S1097-2765(02)00542-7
Almeida SFd, Grosso AR, Koch F, Fenouil R, Carvalho S, Andrade J, Levezinho H et al (2011) Splicing enhances recruitment of methyltransferase HYPB/Setd2 and methylation of histone H3 Lys36. Nat Struct Mol Biol 18(9):977–83. https://doi.org/10.1038/nsmb.2123
Aubert Y, Egolf S, Capell BC (2019) The unexpected noncatalytic roles of histone modifiers in development and disease. Trends Genet 35(9):645–657. https://doi.org/10.1016/j.tig.2019.06.004
Audebert S, Koulakoff A, Berwald-Netter Y, Gros F, Denoulet P, Eddé B (1994) Developmental regulation of polyglutamylated alpha- and beta-tubulin in mouse brain neurons. J Cell Sci 107(Pt 8):2313–2322. https://doi.org/10.1242/jcs.107.8.2313
Awad S, Hassan AH (2008) The Swi2/Snf2 bromodomain is important for the full binding and remodeling activity of the SWI/SNF complex on H3- and H4-acetylated nucleosomes. Ann NY Acad Sci 1138:366–375. https://doi.org/10.1196/annals.1414.038
Ball MP, Li JB, Gao Y, Lee J-H, LeProust EM, Park I-H, **e B, Daley GQ, Church GM (2009) Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat Biotechnol 27(4):361–368. https://doi.org/10.1038/nbt.1533
Baubec T, Colombo DF, Wirbelauer C, Schmidt J, Burger L, Krebs AR, Akalin A, Schübeler D (2015) Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520(7546):243–247. https://doi.org/10.1038/nature14176
Behjati S, Tarpey PS, Presneau N, Scheipl S, Pillay N, Van Loo P, Wedge DC et al (2013) Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 45(12):1479–1482. https://doi.org/10.1038/ng.2814
Bell O, Wirbelauer C, Hild M, Scharf AND, Schwaiger M, MacAlpine DM, Zilbermann F et al (2007) Localized H3K36 methylation states define histone H4K16 acetylation during transcriptional elongation in Drosophila. EMBO J 26(24):4974–4984. https://doi.org/10.1038/sj.emboj.7601926
Bernstein BE, Stamatoyannopoulos JA, Costello JF, Ren B, Milosavljevic A, Meissner A, Kellis M et al (2010) The NIH roadmap epigenomics map** consortium. Nat Biotechnol 28(10):1045–1048. https://doi.org/10.1038/nbt1010-1045
Berry WL, Shin S, Lightfoot SA, Janknecht R (2012) Oncogenic features of the JMJD2A histone demethylase in breast cancer. Int J Oncol 41(5):1701–1706. https://doi.org/10.3892/ijo.2012.1618
Bewick AJ, Vogel KJ, Moore AJ, Schmitz RJ (2017) Evolution of DNA Methylation across Insects. Mol Biol Evol 34(3):654–665. https://doi.org/10.1093/molbev/msw264
Bhattacharya S, Lange JJ, Levy M, Florens L, Washburn MP, Workman JL (2021) The disordered regions of the methyltransferase SETD2 govern its function by regulating its proteolysis and phase separation. J Biol Chem 297(3):101075. https://doi.org/10.1016/j.jbc.2021.101075
Bhattacharya S, Levy MJ, Zhang N, Li H, Florens L, Washburn MP, Workman JL (2021) The methyltransferase SETD2 couples transcription and splicing by engaging MRNA processing factors through its SHI domain. Nat Commun 12(1):1–16. https://doi.org/10.1038/s41467-021-21663-w
Bhattacharya S, Workman JL (2020) Regulation of SETD2 stability is important for the fidelity of H3K36me3 deposition. Epigenet Chromatin 13(1):40. https://doi.org/10.1186/s13072-020-00362-8
Bilokapic S, Halic M (2019) Nucleosome and ubiquitin position Set2 to methylate H3K36. Nat Commun 10(1):3795. https://doi.org/10.1038/s41467-019-11726-4
Bintu L, Kopaczynska M, Hodges C, Lubkowska L, Kashlev M, Bustamante C (2011) The elongation rate of RNA polymerase determines the fate of transcribed nucleosomes. Nat Struct Mol Biol 18(12):1394–1399. https://doi.org/10.1038/nsmb.2164
Bjerke L, Mackay A, Nandhabalan M, Burford A, Jury A, Popov S, Bax DA et al (2013) Histone H3.3. Mutations drive pediatric glioblastoma through upregulation of MYCN. Cancer Discov 3(5):512–519. https://doi.org/10.1158/2159-8290.CD-12-0426
Black JC, Allen A, Van Rechem C, Forbes E, Longworth M, Tschöp K, Rinehart C et al (2010) Conserved antagonism between JMJD2A/KDM4A and HP1γ during cell cycle progression. Mol Cell 40(5):736–748. https://doi.org/10.1016/j.molcel.2010.11.008
Black JC, Manning AL, Van Rechem C, Kim J, Ladd B, Cho J, Pineda CM et al (2013) KDM4A lysine demethylase induces site-specific copy gain and rereplication of regions amplified in tumors. Cell 154(3):541–555. https://doi.org/10.1016/j.cell.2013.06.051
Bodakuntla S, Schnitzler A, Villablanca C, Gonzalez-Billault C, Bieche I, Janke C, Magiera MM (2020) Tubulin polyglutamylation is a general traffic-control mechanism in hippocampal neurons. J Cell Sci 133(3):jcs241802. https://doi.org/10.1242/jcs.241802
Booth V, Koth CM, Edwards AM, Arrowsmith CH (2000) Structure of a conserved domain common to the transcription factors TFIIS, elongin A, and CRSP70 *. J Biol Chem 275(40):31266–31268. https://doi.org/10.1074/jbc.M002595200
Bortvin A, Winston F (1996) Evidence that Spt6p controls chromatin structure by a direct interaction with histones. Science (New York, NY) 272(5267):1473–1476. https://doi.org/10.1126/science.272.5267.1473
Bowman GD, Poirier MG (2015) Post-translational modifications of histones that influence nucleosome dynamics. Chem Rev 115(6):2274–2295. https://doi.org/10.1021/cr500350x
Brown CE, Howe L, Sousa K, Alley SC, Carrozza MJ, Tan S, Workman JL (2001) Recruitment of HAT complexes by direct activator interactions with the ATM-related Tra1 subunit. Science (New York, NY) 292(5525):2333–37. https://doi.org/10.1126/science.1060214
Bruzzone MJ, Grünberg S, Kubik S, Zentner GE, Shore D (2018) Distinct patterns of histone acetyltransferase and mediator deployment at yeast protein-coding genes. Genes Dev 32(17–18):1252–1265. https://doi.org/10.1101/gad.312173.118
Buratowski S (2009) Progression through the RNA polymerase II CTD cycle. Mol Cell 36(4):541–546. https://doi.org/10.1016/j.molcel.2009.10.019
Buschbeck M, Hake SB (2017) Variants of core histones and their roles in cell fate decisions, development and cancer. Nat Rev Mol Cell Biol 18(5):299–314. https://doi.org/10.1038/nrm.2016.166
Cancer Genome Atlas Research Network (2013) Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499(7456):43–49. https://doi.org/10.1038/nature12222
Carrozza MJ, Li B, Florens L, Suganuma T, Swanson SK, Lee KK, Shia W-J et al (2005) Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123(4):581–592. https://doi.org/10.1016/j.cell.2005.10.023
Carvalho S, Raposo AC, Martins FB, Grosso AR, Chaitanya Sridhara S, Rino J, Carmo-Fonseca M, de Almeida SF (2013) Histone methyltransferase SETD2 coordinates FACT recruitment with nucleosome dynamics during transcription. Nucleic Acids Res 41(5):2881–2893. https://doi.org/10.1093/nar/gks1472
Carvalho S, Vítor AC, Sridhara SC, Martins FB, Raposo AC, Desterro JMP, Ferreira J, de Almeida SF (2014) SETD2 is required for DNA double-strand break repair and activation of the P53-mediated checkpoint. Elife 3(May):e02482. https://doi.org/10.7554/eLife.02482
Chang C-F, Chu P-C, Wu P-Y, Yu M-Y, Lee J-Y, Tsai M-D, Chang M-S (2015) PHRF1 promotes genome integrity by modulating non-homologous end-joining. Cell Death Dis 6(4):e1716. https://doi.org/10.1038/cddis.2015.81
Chatterjee N, Sinha D, Lemma-Dechassa M, Tan S, Shogren-Knaak MA, Bartholomew B (2011) Histone H3 tail acetylation modulates ATP-dependent remodeling through multiple mechanisms. Nucleic Acids Res 39(19):8378–8391. https://doi.org/10.1093/nar/gkr535
Chen K, Liu J, Liu S, **a M, Zhang X, Han D, Jiang Y, Wang C, Cao X (2017) Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell 170(3):492-506.e14. https://doi.org/10.1016/j.cell.2017.06.042
Chiang Y-C, Park I-Y, Terzo EA, Tripathi DN, Mason FM, Fahey CC, Karki M et al (2018) SETD2 haploinsufficiency for microtubule methylation is an early driver of genomic instability in renal cell carcinoma. Can Res 78(12):3135–3146. https://doi.org/10.1158/0008-5472.CAN-17-3460
Choi JK, Howe LJ (2009) Histone acetylation: truth of consequences? This paper is one of a selection of papers published in this special issue, entitled CSBMCB’s 51st annual meeting—epigenetics and chromatin dynamics, and has undergone the journal’s usual peer review process. Biochem Cell Biol 87(1):139–50. https://doi.org/10.1139/O08-112
Chu Y, Sutton A, Sternglanz R, Prelich G (2006) The Bur1 cyclin-dependent protein kinase is required for the normal pattern of histone methylation BySet2. Mol Cell Biol 26(8):3029–3038. https://doi.org/10.1128/MCB.26.8.3029-3038.2006
Cornett EM, Ferry L, Defossez P-A, Rothbart SB (2019) Lysine methylation regulators moonlighting outside the epigenome. Mol Cell 75(6):1092–1101. https://doi.org/10.1016/j.molcel.2019.08.026
Cramer P (2019) Organization and regulation of gene transcription. Nature 573(7772):45–54. https://doi.org/10.1038/s41586-019-1517-4
de Cubas AA, Rathmell WK (2018) Epigenetic modifiers: activities in renal cell carcinoma. Nat Rev Urol 15(10):599–614. https://doi.org/10.1038/s41585-018-0052-7
Dalgliesh GL, Furge K, Greenman C, Chen L, Bignell G, Butler A, Davies H et al (2010) Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463(7279):360–363. https://doi.org/10.1038/nature08672
Daugaard M, Baude A, Fugger K, Povlsen LK, Beck H, Sørensen CS, Petersen NHT et al (2012) LEDGF (P75) promotes DNA-End resection and homologous recombination. Nat Struct Mol Biol 19(8):803–810. https://doi.org/10.1038/nsmb.2314
Deal RB, Henikoff JG, Henikoff S (2010) Genome-wide kinetics of nucleosome turnover determined by metabolic labeling of histones. Science 328(5982):1161–1164. https://doi.org/10.1126/science.1186777
Diebold M-L, Koch M, Loeliger E, Cura V, Winston F, Cavarelli J, Romier C (2010) The structure of an Iws1/Spt6 complex reveals an interaction domain conserved in TFIIS, elongin A and Med26. EMBO J 29(23):3979–3991. https://doi.org/10.1038/emboj.2010.272
Dion MF, Kaplan T, Kim M, Buratowski S, Friedman N, Rando OJ (2007) Dynamics of replication-independent histone turnover in budding yeast. Science (New York NY) 315(5817):1405–1408. https://doi.org/10.1126/science.1134053
Dover J, Schneider J, Tawiah-Boateng MA, Wood A, Dean K, Johnston M, Shilatifard A (2002) Methylation of histone H3 by COMPASS requires ubiquitination of histone H2B by Rad6. J Biol Chem 277(32):28368–28371. https://doi.org/10.1074/jbc.C200348200
Dronamraju R, Jha DK, Eser U, Adams AT, Dominguez D, Choudhury R, Chiang Y-C et al (2018) Set2 methyltransferase facilitates cell cycle progression by maintaining transcriptional fidelity. Nucleic Acids Res 46(3):1331–1344. https://doi.org/10.1093/nar/gkx1276
Drouin S, Laramée L, Jacques P-É, Forest A, Bergeron M, Robert F (2010) DSIF and RNA polymerase II CTD phosphorylation coordinate the recruitment of Rpd3S to actively transcribed genes. PLoS Genet 6(10):e1001173. https://doi.org/10.1371/journal.pgen.1001173
Du H-N, Fingerman IM, Briggs SD (2008) Histone H3 K36 methylation is mediated by a trans-histone methylation pathway involving an interaction between Set2 and histone H4. Genes Dev 22(20):2786–2798. https://doi.org/10.1101/gad.1700008
Duns G, van den Berg E, van Duivenbode I, Osinga J, Hollema H, Hofstra RMW, Kok K (2010) Histone methyltransferase gene SETD2 is a novel tumor suppressor gene in clear cell renal cell carcinoma. Can Res 70(11):4287–4291. https://doi.org/10.1158/0008-5472.CAN-10-0120
Edmunds JW, Mahadevan LC, Clayton AL (2008) Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J 27(2):406–420. https://doi.org/10.1038/sj.emboj.7601967
Egloff S, Murphy S (2008) Cracking the RNA polymerase II CTD code. Trends Genet 24(6):280–288. https://doi.org/10.1016/j.tig.2008.03.008
Eick D, Geyer M (2013) The RNA polymerase II carboxy-terminal domain (CTD) code. Chem Rev 113(11):8456–8490. https://doi.org/10.1021/cr400071f
Eram MS, Kuznetsova E, Li F, Lima-Fernandes E, Kennedy S, Chau I, Arrowsmith CH, Schapira M, Vedadi M (2015) Kinetic characterization of human histone H3 lysine 36 methyltransferases, ASH1L and SETD2. Biochem Biophys Acta 1850(9):1842–1848. https://doi.org/10.1016/j.bbagen.2015.05.013
Faber PW, Barnes GT, Srinidhi J, Chen J, Gusella JF, MacDonald ME (1998) Huntingtin interacts with a family of WW domain proteins. Hum Mol Genet 7(9):1463–1474. https://doi.org/10.1093/hmg/7.9.1463
Fang D, Gan H, Lee J-H, Han J, Wang Z, Riester SM, ** L et al (2016) The histone H3.3K36M mutation reprograms the epigenome of chondroblastomas. Science 352(6291):1344–1348. https://doi.org/10.1126/science.aae0065
Ferrari P, Strubin M (2015) Uncoupling histone turnover from transcription-associated histone H3 modifications. Nucleic Acids Res 43(8):3972–3985. https://doi.org/10.1093/nar/gkv282
Formosa T (2012) The role of FACT in making and breaking nucleosomes. Biochem Biophys Acta 1819(3–4):247–255. https://doi.org/10.1016/j.bbagrm.2011.07.009
Fuchs SM, Kizer KO, Braberg H, Krogan NJ, Strahl BD (2012) RNA Polymerase II carboxyl-terminal domain phosphorylation regulates protein stability of the Set2 methyltransferase and histone H3 Di- and trimethylation at lysine 36. J Biol Chem 287(5):3249–3256. https://doi.org/10.1074/jbc.M111.273953
Gaertig J, Wloga D (2008) Chapter 4 Ciliary tubulin and its post-translational modifications. In: Current topics in developmental biology. Ciliary Function in Mammalian Development, vol 85. Academic Press, pp 83–113. https://doi.org/10.1016/S0070-2153(08)00804-1
Gao Y-G, Yang H, Zhao J, Jiang Y-J, Hong-Yu Hu (2014) Autoinhibitory structure of the WW domain of HYPB/SETD2 regulates its interaction with the proline-rich region of huntingtin. Structure 22(3):378–386. https://doi.org/10.1016/j.str.2013.12.005
Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E, Martinez P et al (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366(10):883–892. https://doi.org/10.1056/NEJMoa1113205
Gill JK, Maffioletti A, García-Molinero V, Stutz F, Soudet J (2020) Fine chromatin-driven mechanism of transcription interference by antisense noncoding transcription. Cell Rep 31(5):107612. https://doi.org/10.1016/j.celrep.2020.107612
Goldberg AD, Banaszynski LA, Noh K-M, Lewis PW, Elsaesser SJ, Stadler S, Dewell S et al (2010) Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140(5):678–691. https://doi.org/10.1016/j.cell.2010.01.003
Gopalakrishnan R, Marr SK, Kingston RE, Winston F (2019) A conserved genetic interaction between Spt6 and Set2 regulates H3K36 methylation. Nucleic Acids Res 47(8):3888–3903. https://doi.org/10.1093/nar/gkz119
Govind CK, Qiu H, Ginsburg DS, Ruan C, Hofmeyer K, Cuihua Hu, Swaminathan V, Workman JL, Li B, Hinnebusch AG (2010) Phosphorylated Pol II CTD recruits multiple HDACs, including Rpd3C(S), for methylation-dependent deacetylation of ORF nucleosomes. Mol Cell 39(2):234–246. https://doi.org/10.1016/j.molcel.2010.07.003
Grosso AR, Leite AP, Carvalho S, Matos MR, Martins FB, Vítor AC, Desterro JMP, Carmo-Fonseca M, de Almeida SF (2015) Pervasive transcription read-through promotes aberrant expression of oncogenes and RNA chimeras in renal carcinoma. ELife. https://doi.org/10.7554/eLife.09214
Guerra-Calderas L, González-Barrios R, Herrera LA, Cantú de León D, Soto-Reyes E (2015) The role of the histone demethylase KDM4A in cancer. Cancer Genet 208(5):215–24. https://doi.org/10.1016/j.cancergen.2014.11.001
Guo R, Zheng L, Park JW, Lv R, Chen H, Jiao F, Wenqi Xu et al (2014) BS69/ZMYND11 reads and connects histone H3.3 lysine 36 trimethylation decorated chromatin to regulated pre-MRNA processing. Mol Cell 56(2):298–310. https://doi.org/10.1016/j.molcel.2014.08.022
Hakimi AA, Ostrovnaya I, Reva B, Schultz N, Chen Y-B, Gonen M, Liu H et al (2013) Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin Cancer Res 19(12):3259–3267. https://doi.org/10.1158/1078-0432.CCR-12-3886
Hauer MH, Gasser SM (2017) Chromatin and nucleosome dynamics in DNA damage and repair. Genes Dev 31(22):2204–2221. https://doi.org/10.1101/gad.307702.117
Ho TH, Park IY, Zhao H, Tong P, Champion MD, Yan H, Monzon FA et al (2016) High-resolution profiling of histone H3 lysine 36 trimethylation in metastatic renal cell carcinoma. Oncogene 35(12):1565–1574. https://doi.org/10.1038/onc.2015.221
Hodges C, Bintu L, Lubkowska L, Kashlev M, Bustamante C (2009) Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II. Science 325(5940):626–628. https://doi.org/10.1126/science.1172926
Hödl M, Basler K (2012) Transcription in the absence of histone H3.2 and H3K4 methylation. Curr Biol 22(23):2253–2257. https://doi.org/10.1016/j.cub.2012.10.008
Holmes RK, Tuck AC, Zhu C, Dunn-Davies HR, Kudla G, Clauder-Munster S, Granneman S, Steinmetz LM, Guthrie C, Tollervey D (2015) Loss of the yeast SR protein Npl3 alters gene expression due to transcription readthrough. PLoS Genet. https://doi.org/10.1371/journal.pgen.1005735
Howe FS, Fischl H, Murray SC, Mellor J (2017) Is H3K4me3 instructive for transcription activation? BioEssays 39(1):e201600095. https://doi.org/10.1002/bies.201600095
Hsieh F-K, Kulaeva OI, Patel SS, Dyer PN, Luger K, Reinberg D, Studitsky VM (2013) Histone chaperone FACT action during transcription through chromatin by RNA polymerase II. Proc Natl Acad Sci 110(19):7654–7659. https://doi.org/10.1073/pnas.1222198110
Hsieh JJ, Le VH, Oyama T, Ricketts CJ, Ho TH, Cheng EH (2018) Chromosome 3p loss-orchestrated VHL, HIF, and epigenetic deregulation in clear cell renal cell carcinoma. J Clin Oncol 36(36):3533–3539. https://doi.org/10.1200/JCO.2018.79.2549
Hsin J-P, Manley JL (2012) The RNA polymerase II CTD coordinates transcription and RNA processing. Genes Dev 26(19):2119–2137. https://doi.org/10.1101/gad.200303.112
Huang Y, Liya Gu, Li G-M (2018) H3K36me3-mediated mismatch repair preferentially protects actively transcribed genes from mutation. J Biol Chem 293(20):7811–7823. https://doi.org/10.1074/jbc.RA118.002839
Jamai A, Puglisi A, Strubin M (2009) Histone chaperone Spt16 promotes redeposition of the original H3–H4 histones evicted by elongating RNA polymerase. Mol Cell 35(3):377–383. https://doi.org/10.1016/j.molcel.2009.07.001
Janke C (2014) The tubulin code: molecular components, readout mechanisms, and functions. J Cell Biol 206(4):461–472. https://doi.org/10.1083/jcb.201406055
Janna A, Davarinejad H, Joshi M, Couture J-F (2020) Structural paradigms in the recognition of the nucleosome core particle by histone lysine methyltransferases. Front Cell Develop Biol 8:600. https://doi.org/10.3389/fcell.2020.00600
Jeronimo C, Poitras C, Robert F (2019) Histone recycling by FACT and Spt6 during transcription prevents the scrambling of histone modifications. Cell Rep 28(5):1206-1218.e8. https://doi.org/10.1016/j.celrep.2019.06.097
Jha DK, Strahl BD (2014) An RNA polymerase II-coupled function for histone H3K36 methylation in checkpoint activation and DSB repair. Nat Commun 5:3965. https://doi.org/10.1038/ncomms4965
Jimeno-González S, Payán-Bravo L, Muñoz-Cabello AM, Guijo M, Gutierrez G, Prado F, Reyes JC (2015) Defective histone supply causes changes in RNA polymerase II elongation rate and cotranscriptional Pre-MRNA splicing. Proc Natl Acad Sci 112(48):14840–14845. https://doi.org/10.1073/pnas.1506760112
Joshi AA, Struhl K (2005) Eaf3 chromodomain interaction with methylated H3–K36 links histone deacetylation to Pol II elongation. Mol Cell 20(6):971–978. https://doi.org/10.1016/j.molcel.2005.11.021
Josling GA, Selvarajah SA, Petter M, Duffy MF (2012) The role of bromodomain proteins in regulating gene expression. Genes 3(2):320–343. https://doi.org/10.3390/genes3020320
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K et al (2021) Highly accurate protein structure prediction with alphafold. Nature 596(7873):583–589. https://doi.org/10.1038/s41586-021-03819-2
Kallappagoudar S, Yadav RK, Lowe BR, Partridge JF (2015) Histone H3 mutations—a special role for H3.3 in tumorigenesis? Chromosoma 124(2):177–189. https://doi.org/10.1007/s00412-015-0510-4
Kanu N, Grönroos E, Martinez P, Burrell RA, Yi Goh X, Bartkova J, Maya-Mendoza A et al (2015) SETD2 loss-of-function promotes renal cancer branched evolution through replication stress and impaired DNA repair. Oncogene 34(46):5699–5708. https://doi.org/10.1038/onc.2015.24
Kassem S, Ferrari P, Hughes AL, Soudet J, Rando OJ, Strubin M (2020) Histone exchange is associated with activator function at transcribed promoters and with repression at histone loci. Sci Adv 6(36):0333. https://doi.org/10.1126/sciadv.abb0333
Kearns S, Mason FM, Rathmell WK, Park IY, Walker C, Verhey KJ, Cianfrocco MA (2021) Molecular determinants for α-tubulin methylation by SETD2. J Biol Chem 297(1):100898. https://doi.org/10.1016/j.jbc.2021.100898
Keogh M-C, Kurdistani SK, Morris SA, Ahn SH, Podolny V, Collins SR, Schuldiner M et al (2005) Cotranscriptional Set2 methylation of histone H3 lysine 36 recruits a repressive Rpd3 complex. Cell 123(4):593–605. https://doi.org/10.1016/j.cell.2005.10.025
Kim JH, Lee BB, Young Mi O, Zhu C, Steinmetz LM, Lee Y, Kim WK, Lee SB, Buratowski S, Kim TS (2016) Modulation of MRNA and LncRNA expression dynamics by the Set2–Rpd3S pathway. Nat Commun. https://doi.org/10.1038/ncomms13534
Kim S, Kim H, Fong N, Erickson B, Bentley DL (2011) Pre-MRNA splicing is a determinant of histone H3K36 methylation. Proc Natl Acad Sci 108(33):13564–13569. https://doi.org/10.1073/pnas.1109475108
Kizer KO, Phatnani HP, Shibata Y, Hall H, Greenleaf AL, Strahl BD (2005) A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3 K36 methylation with transcript elongation. Mol Cell Biol 25(8):3305–3316. https://doi.org/10.1128/MCB.25.8.3305-3316.2005
Klein BJ, Krajewski K, Restrepo S, Lewis PW, Strahl BD, Kutateladze TG (2018) Recognition of cancer mutations in histone H3K36 by epigenetic writers and readers. Epigenetics 13(7):683–692. https://doi.org/10.1080/15592294.2018.1503491
Klose RJ, Yamane K, Bae Y, Zhang D, Erdjument-Bromage H, Tempst P, Wong J, Zhang Yi (2006) The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine 9 and lysine 36. Nature 442(7100):312–316. https://doi.org/10.1038/nature04853
Koenning M, Wang X, Karki M, Jangid MK, Kearns S, Tripathi DN, Cianfrocco M et al (2021) Neuronal SETD2 activity links microtubule methylation to an anxiety-like phenotype in mice. Brain J Neurol 144(8):2527–2540. https://doi.org/10.1093/brain/awab200
Krogan NJ, Kim M, Ahn SH, Zhong G, Kobor MS, Cagney G, Emili A, Shilatifard A, Buratowski S, Greenblatt JF (2002) RNA polymerase II elongation factors of saccharomyces cerevisiae: a targeted proteomics approach. Mol Cell Biol 22(20):6979–6992. https://doi.org/10.1128/mcb.22.20.6979-6992.2002
Kubo T, Yanagisawa H, Yagi T, Hirono M, Kamiya R (2010) Tubulin polyglutamylation regulates axonemal motility by modulating activities of inner-arm dyneins. Curr Biol 20(5):441–445. https://doi.org/10.1016/j.cub.2009.12.058
Kulaeva OI, Gaykalova DA, Pestov NA, Golovastov VV, Vassylyev DG, Artsimovitch I, Studitsky VM (2009) Mechanism of chromatin remodeling and recovery during passage of RNA polymerase II. Nat Struct Mol Biol 16(12):1272–1278. https://doi.org/10.1038/nsmb.1689
Kulaeva OI, Hsieh F-K, Studitsky VM (2010) RNA polymerase complexes cooperate to relieve the nucleosomal barrier and evict histones. Proc Natl Acad Sci 107(25):11325–11330. https://doi.org/10.1073/pnas.1001148107
Laribee RN, Krogan NJ, **ao T, Shibata Y, Hughes TR, Greenblatt JF, Strahl BD (2005) BUR kinase selectively regulates H3 K4 trimethylation and H2B ubiquitylation through recruitment of the PAF elongation complex. Curr Biol 15(16):1487–1493. https://doi.org/10.1016/j.cub.2005.07.028
Lee DH, Kim GW, Jeon YH, Yoo J, Lee SW, Kwon SH (2020) Advances in histone demethylase KDM4 as cancer therapeutic targets. FASEB J 34(3):3461–3484. https://doi.org/10.1096/fj.201902584R
Lee KK, Workman JL (2007) Histone acetyltransferase complexes: one size doesn’t fit all. Nat Rev Mol Cell Biol 8(4):284–295. https://doi.org/10.1038/nrm2145
LeRoy G, Oksuz O, Descostes N, Aoi Y, Ganai RA, Kara HO, Jia-Ray Y et al (2019) LEDGF and HDGF2 relieve the nucleosome-induced barrier to transcription in differentiated cells. Sci Adv 5(10):3068. https://doi.org/10.1126/sciadv.aay3068
Leung CS, Douglass SM, Morselli M, Obusan MB, Pavlyukov MS, Pellegrini M, Johnson TL (2019) H3K36 methylation and the chromodomain protein Eaf3 are required for proper cotranscriptional spliceosome assembly. Cell Rep 27(13):3760-3769.e4. https://doi.org/10.1016/j.celrep.2019.05.100
Lewis PW, Müller MM, Koletsky MS, Cordero F, Lin S, Banaszynski LA, Garcia BA, Muir TW, Becher OJ, David Allis C (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science (New York, NY) 340(6134):857–61. https://doi.org/10.1126/science.1232245
Li B, Gogol M, Carey M, Lee D, Seidel C, Workman JL (2007) Combined action of PHD and chromo domains directs the Rpd3S HDAC to transcribed chromatin. Science (New York, NY) 316(5827):1050–54. https://doi.org/10.1126/science.1139004
Li B, Howe LeAnn, Anderson S, Yates JR, Workman JL (2003) The Set2 histone methyltransferase functions through the phosphorylated carboxyl-terminal domain of RNA polymerase II. J Biol Chem 278(11):8897–8903. https://doi.org/10.1074/jbc.M212134200
Li B, Jackson J, Simon MD, Fleharty B, Gogol M, Seidel C, Workman JL, Shilatifard A (2009) Histone H3 lysine 36 dimethylation (H3K36me2) is sufficient to recruit the Rpd3s histone deacetylase complex and to repress spurious transcription. J Biol Chem 284(12):7970–7976. https://doi.org/10.1074/jbc.M808220200
Li F, Mao G, Tong D, Huang J, Liya Gu, Yang W, Li G-M (2013) The histone mark H3K36me3 regulates human DNA mismatch repair through its interaction with MutSα. Cell 153(3):590–600. https://doi.org/10.1016/j.cell.2013.03.025
Li J, Ahn JH, Wang GG (2019) Understanding histone H3 lysine 36 methylation and its deregulation in disease. Cell Mol Life Sci 76(15):2899–2916. https://doi.org/10.1007/s00018-019-03144-y
Li J, Duns G, Westers H, Sijmons R, van den Berg A, Kok K (2016) SETD2: an epigenetic modifier with tumor suppressor functionality. Oncotarget 7(31):50719–50734. https://doi.org/10.18632/oncotarget.9368
Li L, Wang Y (2017) Cross-talk between the H3K36me3 and H4K16ac histone epigenetic marks in DNA double-strand break repair. J Biol Chem 292(28):11951–11959. https://doi.org/10.1074/jbc.M117.788224
Li M, Phatnani HP, Guan Z, Sage H, Greenleaf AL, Zhou P (2005) Solution structure of the Set2-Rpb1 interacting domain of human Set2 and its interaction with the hyperphosphorylated C-terminal domain of Rpb1. Proc Natl Acad Sci USA 102(49):17636–17641. https://doi.org/10.1073/pnas.0506350102
Lickwar CR, Rao B, Shabalin AA, Nobel AB, Strahl BD, Lieb JD (2009) The Set2/Rpd3S pathway suppresses cryptic transcription without regard to gene length or transcription frequency. PLoS One 4(3):e4886. https://doi.org/10.1371/journal.pone.0004886
Lim J, Park JH, Baude A, Yoo Y, Lee YK, Schmidt CR, Park JB et al (2017) The histone variant H3.3 G34W substitution in giant cell tumor of the bone link chromatin and RNA processing. Sci Rep 7(1):13459. https://doi.org/10.1038/s41598-017-13887-y
Lindstrom DL, Squazzo SL, Muster N, Burckin TA, Wachter KC, Emigh CA, McCleery JA, Yates JR, Hartzog GA (2003) Dual roles for Spt5 in Pre-MRNA processing and transcription elongation revealed by identification of Spt5-associated proteins. Mol Cell Biol 23(4):1368–1378. https://doi.org/10.1128/MCB.23.4.1368-1378.2003
Ling Y, Smith AJ, Morgan GT (2006) A sequence Motif conserved in diverse nuclear proteins identifies a protein interaction domain utilised for nuclear targeting by human TFIIS. Nucleic Acids Res 34(8):2219–2229. https://doi.org/10.1093/nar/gkl239
Liu L, Guo R, Zhang X, Liang Y, Kong F, Wang J, Zhonghua Xu (2017) Loss of SETD2, but not H3K36me3, correlates with aggressive clinicopathological features of clear cell renal cell carcinoma patients. Biosci Trends 11(2):214–220. https://doi.org/10.5582/bst.2016.01228
Liu Y, Zhang Y, Xue H, Cao Mi, Bai G, Zongkai Mu, Yao Y, Sun S, Fang D, Huang J (2021) Cryo-EM structure of SETD2/Set2 methyltransferase bound to a nucleosome containing oncohistone mutations. Cell Discovery 7(1):1–12. https://doi.org/10.1038/s41421-021-00261-6
Lowe BR, Maxham LA, Hamey JJ, Wilkins MR, Partridge JF (2019) Histone H3 mutations: an updated view of their role in chromatin deregulation and cancer. Cancers 11(5):E660. https://doi.org/10.3390/cancers11050660
Lowe BR, Yadav RK, Henry RA, Schreiner P, Matsuda A, Fernandez AG, Finkelstein D et al (2021) “Surprising phenotypic diversity of cancer-associated mutations of Gly 34 in the histone H3 Tail” edited by Jerry L Workman, Jessica K Tyler, and Jerry L Workman. ELife 10:65369. https://doi.org/10.7554/eLife.65369
Lu C, Jain SU, Hoelper D, Bechet D, Molden RC, Ran L, Murphy D et al (2016) Histone H3K36 mutations promote sarcomagenesis through altered histone methylation landscape. Science (New York, NY) 352(6287):844–49. https://doi.org/10.1126/science.aac7272
Lu M, Zhao B, Liu M, Le Wu, Li Y, Zhai Y, Shen X (2021) Pan-cancer analysis of SETD2 mutation and its association with the efficacy of immunotherapy. Npj Precis Oncol 5(1):1–6. https://doi.org/10.1038/s41698-021-00193-0
Luco RF, Pan Q, Tominaga K, Blencowe BJ, Pereira-Smith OM, Misteli T (2010) Regulation of alternative splicing by histone modifications. Science (New York, NY) 327(5968):996–1000. https://doi.org/10.1126/science.1184208
MacAlpine DM, Almouzni G (2013) Chromatin and DNA replication. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a010207
Mallette FA, Mattiroli F, Cui G, Young LC, Hendzel MJ, Mer G, Sixma TK, Richard S (2012) RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J 31(8):1865–1878. https://doi.org/10.1038/emboj.2012.47
Mallette FA, Richard S (2012) JMJD2A promotes cellular transformation by blocking cellular senescence through transcriptional repression of the tumor suppressor CHD5. Cell Rep 2(5):1233–1243. https://doi.org/10.1016/j.celrep.2012.09.033
Marnef A, Cohen S, Legube G (2017) Transcription-coupled DNA double-strand break repair: active genes need special care. J Mol Biol 429(9):1277–1288. https://doi.org/10.1016/j.jmb.2017.03.024
Martin BJE, Amour JB, Kuzmin A, Jensen KN, Cheng Liu Z, Lorincz M, Howe LJ (2021) Transcription shapes genome-wide histone acetylation patterns. Nat Commun 12(1):210. https://doi.org/10.1038/s41467-020-20543-z
Masumoto H, Hawke D, Kobayashi R, Verreault A (2005) A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436(7048):294–298. https://doi.org/10.1038/nature03714
Maunakea AK, Nagarajan RP, Bilenky M, Ballinger TJ, D’Souza C, Fouse SD, Johnson BE et al (2010) Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature 466(7303):253–257. https://doi.org/10.1038/nature09165
McDonald SM, Close D, **n H, Formosa T, Hill CP (2010) Structure and biological importance of the Spn1-Spt6 interaction, and its regulatory role in nucleosome binding. Mol Cell 40(5):725–735. https://doi.org/10.1016/j.molcel.2010.11.014
Medvedeva YA, Khamis AM, Kulakovskiy IV, Wail Ba-Alawi Md, Bhuyan SI, Kawaji H, Lassmann T et al (2014) Effects of cytosine methylation on transcription factor binding sites. BMC Genom 15(1):119. https://doi.org/10.1186/1471-2164-15-119
Meers MP, Henriques T, Lavender CA, McKay DJ, Strahl BD, Duronio RJ, Adelman K, Matera AG (2017) Histone gene replacement reveals a post-transcriptional role for H3K36 in maintaining metazoan transcriptome fidelity. ELife. https://doi.org/10.7554/eLife.23249
Michael AK, Thomä NH (2021) Reading the chromatinized genome. Cell 184(14):3599–3611. https://doi.org/10.1016/j.cell.2021.05.029
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38(1):23–38. https://doi.org/10.1038/npp.2012.112
Morgan MAJ, Shilatifard A (2020) Reevaluating the roles of histone-modifying enzymes and their associated chromatin modifications in transcriptional regulation. Nat Genet 52(12):1271–1281. https://doi.org/10.1038/s41588-020-00736-4
Muhar M, Ebert A, Neumann T, Umkehrer C, Jude J, Wieshofer C, Rescheneder P et al (2018) SLAM-Seq defines direct gene-regulatory functions of the BRD4-MYC Axis. Science (New York, NY) 360(6390):800–805. https://doi.org/10.1126/science.aao2793
Murawska M, Greenstein RA, Schauer T, Olsen KCF, Ng H, Ladurner AG, Al-Sady B, Braun S (2021) The histone chaperone FACT facilitates heterochromatin spreading by regulating histone turnover and H3K9 methylation states. Cell Rep 37(5):109944. https://doi.org/10.1016/j.celrep.2021.109944
Neri F, Rapelli S, Krepelova A, Incarnato D, Parlato C, Basile G, Maldotti M, Anselmi F, Oliviero S (2017) Intragenic DNA methylation prevents spurious transcription initiation. Nature 543(7643):72–77. https://doi.org/10.1038/nature21373
Osman S, Cramer P (2020) Structural biology of RNA polymerase II transcription: 20 years on. Annu Rev Cell Dev Biol 36:1–34. https://doi.org/10.1146/annurev-cellbio-042020-021954
Pai C-C, Deegan RS, Subramanian L, Gal C, Sarkar S, Blaikley EJ, Walker C et al (2014) A histone H3K36 chromatin switch coordinates DNA double-strand break repair pathway choice. Nat Commun 5:4091. https://doi.org/10.1038/ncomms5091
Pai C-C, Hsu K-F, Durley SC, Keszthelyi A, Kearsey SE, Rallis C, Folkes LK et al (2019) An essential role for DNTP homeostasis following CDK-induced replication stress. J Cell Sci 132(6):226969. https://doi.org/10.1242/jcs.226969
Pai C-C, Kishkevich A, Deegan RS, Keszthelyi A, Folkes L, Kearsey SE, De León N et al (2017) Set2 methyltransferase facilitates DNA replication and promotes genotoxic stress responses through MBF-dependent transcription. Cell Rep 20(11):2693–2705. https://doi.org/10.1016/j.celrep.2017.08.058
Papillon-Cavanagh S, Chao Lu, Gayden T, Mikael LG, Bechet D, Karamboulas C, Ailles L et al (2017) Impaired H3K36 methylation defines a subset of head and neck squamous cell carcinomas. Nat Genet 49(2):180–185. https://doi.org/10.1038/ng.3757
Park Y, Powell RT, Tripathi DN, Dere R, Ho TH, Lynne Blasius T, Chiang Y-C et al (2016) Dual chromatin and cytoskeletal remodeling by SETD2. Cell 166(4):950. https://doi.org/10.1016/j.cell.2016.07.005
Petesch SJ, Lis JT (2012) Overcoming the nucleosome barrier during transcript elongation. Trends Genet 28(6):285–294. https://doi.org/10.1016/j.tig.2012.02.005
Pfister SX, Ahrabi S, Zalmas L-P, Sarkar S, Aymard F, Bachrati CZ, Helleday T et al (2014) SETD2-dependent histone H3K36 trimethylation is required for homologous recombination repair and genome stability. Cell Rep 7(6):2006–2018. https://doi.org/10.1016/j.celrep.2014.05.026
Pfister SX, Markkanen E, Jiang Y, Sarkar S, Woodcock M, Orlando G, Mavrommati I et al (2015) Inhibiting WEE1 selectively kills histone H3K36me3-deficient cancers by DNTP starvation. Cancer Cell 28(5):557–568. https://doi.org/10.1016/j.ccell.2015.09.015
Pinto D, Pagé V, Fisher RP, Tanny JC (2021) New connections between ubiquitylation and methylation in the co-transcriptional histone modification network. Curr Genet 67(5):695–705. https://doi.org/10.1007/s00294-021-01196-x
Pradeepa MM, Sutherland HG, Ule J, Grimes GR, Bickmore WA (2012) Psip1/Ledgf P52 binds methylated histone H3K36 and splicing factors and contributes to the regulation of alternative splicing. PLoS Genet 8(5):e1002717. https://doi.org/10.1371/journal.pgen.1002717
Radman-Livaja M, Verzijlbergen KF, Weiner A, van Welsem T, Friedman N, Rando OJ, van Leeuwen F (2011) Patterns and mechanisms of ancestral histone protein inheritance in budding yeast. PLoS Biol 9(6):e1001075. https://doi.org/10.1371/journal.pbio.1001075
Rawal Y, Chereji RV, Qiu H, Ananthakrishnan S, Govind CK, Clark DJ, Hinnebusch AG (2018) SWI/SNF and RSC cooperate to reposition and evict promoter nucleosomes at highly expressed genes in yeast. Genes Dev 32(9–10):695–710. https://doi.org/10.1101/gad.312850.118
Reim NI, Chuang J, Jain D, Alver BH, Park PJ, Winston F (2020) The conserved elongation factor Spn1 is required for normal transcription, histone modifications, and splicing in Saccharomyces cerevisiae. Nucleic Acids Res 48(18):10241–10258. https://doi.org/10.1093/nar/gkaa745
Ruan C, Lee C-H, Cui H, Li S, Li B (2015) Nucleosome contact triggers conformational changes of Rpd3S driving high affinity H3K36me nucleosome engagement. Cell Rep 10(2):204–215. https://doi.org/10.1016/j.celrep.2014.12.027
Rufiange A, Jacques P-E, Bhat W, Robert F, Nourani A (2007) Genome-wide replication-independent histone H3 exchange occurs predominantly at promoters and implicates H3 K56 acetylation and Asf1. Mol Cell 27(3):393–405. https://doi.org/10.1016/j.molcel.2007.07.011
Santos-Rosa H, Millán-Zambrano G, Han N, Leonardi T, Klimontova M, Nasiscionyte S, Pandolfini L, Tzelepis K, Bartke T, Kouzarides T (2021) Methylation of histone H3 at lysine 37 by Set1 and Set2 prevents spurious DNA replication. Mol Cell 81(13):2793-2807.e8. https://doi.org/10.1016/j.molcel.2021.04.021
Schaft D, Roguev A, Kotovic KM, Shevchenko A, Sarov M, Shevchenko A, Neugebauer KM, Francis Stewart A (2003) The histone 3 lysine 36 methyltransferase, SET2, is involved in transcriptional elongation. Nucleic Acids Res 31(10):2475–2482. https://doi.org/10.1093/nar/gkg372
Schwartzentruber J, Korshunov A, Liu X-Y, Jones DTW, Pfaff E, Jacob K, Sturm D et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231. https://doi.org/10.1038/nature10833
Seervai RNH, Jangid RK, Karki M, Tripathi DN, Jung SY, Kearns SE, Verhey KJ et al (2020) The huntingtin-interacting protein SETD2/HYPB is an actin lysine methyltransferase. Sci Adv. https://doi.org/10.1126/sciadv.abb7854
Selvam K, Plummer DA, Mao P, Wyrick JJ (2022) Set2 histone methyltransferase regulates transcription coupled-nucleotide excision repair in yeast. PLoS Genet 18(3):e1010085. https://doi.org/10.1371/journal.pgen.1010085
Separovich RJ, Wilkins MR (2021) Ready, SET, go: post-translational regulation of the histone lysine methylation network in budding yeast. J Biol Chem 297(2):100939. https://doi.org/10.1016/j.jbc.2021.100939
Sessa A, Fagnocchi L, Mastrototaro G, Massimino L, Zaghi M, Indrigo M, Cattaneo S et al (2019) SETD5 regulates chromatin methylation state and preserves global transcriptional fidelity during brain development and neuronal wiring. Neuron 104(2):271-289.e13. https://doi.org/10.1016/j.neuron.2019.07.013
Shah MA, Denton EL, Arrowsmith CH, Lupien M, Schapira M (2014) A global assessment of cancer genomic alterations in epigenetic mechanisms. Epigenet Chromat 7(1):29. https://doi.org/10.1186/1756-8935-7-29
Simon JM, Hacker KE, Darshan Singh A, Brannon R, Parker JS, Weiser M, Ho TH et al (2014) Variation in chromatin accessibility in human kidney cancer links H3K36 methyltransferase loss with widespread RNA processing defects. Genome Res 24(2):241–250. https://doi.org/10.1101/gr.158253.113
Skucha A, Ebner J, Grebien F (2019) Roles of SETD2 in leukemia-transcription, DNA-damage, and beyond. Int J Mol Sci 20(5):E1029. https://doi.org/10.3390/ijms20051029
Smolle M, Venkatesh S, Gogol MM, Li H, Zhang Y, Florens L, Washburn MP, Workman JL (2012) Chromatin remodelers Isw1 and Chd1 maintain chromatin structure during transcription by preventing histone exchange. Nat Struct Mol Biol 19(9):884–892. https://doi.org/10.1038/nsmb.2312
Snyder MP, Gingeras TR, Moore JE, Weng Z, Gerstein MB, Ren B, Hardison RC et al (2020) Perspectives on ENCODE. Nature 583(7818):693–698. https://doi.org/10.1038/s41586-020-2449-8
Sorenson MR, Jha DK, Ucles SA, Flood DM, Strahl BD, Stevens SW, Kress TL (2016) Histone H3K36 methylation regulates pre-MRNA splicing in Saccharomyces cerevisiae. RNA Biol 13(4):412–426. https://doi.org/10.1080/15476286.2016.1144009
Speranzini V, Pilotto S, Sixma TK, Mattevi A (2016) Touch, act and Go: landing and operating on nucleosomes. EMBO J 35(4):376–88. https://doi.org/10.1525/embj.201593377
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403(6765):41–45. https://doi.org/10.1038/47412
Strahl BD, Grant PA, Briggs SD, Sun Z-W, Bone JR, Caldwell JA, Mollah S et al (2002) Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol Cell Biol 22(5):1298–1306. https://doi.org/10.1128/mcb.22.5.1298-1306.2002
Sumrall AL, **u J, Eschbacher JM, Mittal S, Gatalica Z, Pandey MK, Phuphanich S, Korn WM, Fuller GN, Heimberger AB (2019) Mutations of H3.3 and H3.1 in a large cohort of glioma tumors. J Clin Oncol 37:13540–13540. https://doi.org/10.1200/JCO.2019.37.15_suppl.e13540
Sun X-J, Wei Ju, **n-Yan Wu, Ming Hu, Wang L, Wang H-H, Zhang Q-H, Chen S-J, Huang Q-H, Chen Z (2005) Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J Biol Chem 280(42):35261–35271. https://doi.org/10.1074/jbc.M504012200
Sun Z, Zhang Y, Jia J, Fang Y, Tang Y, Hongfei Wu, Fang D (2020) H3K36me3, message from chromatin to DNA damage repair. Cell Biosci 10(1):9. https://doi.org/10.1186/s13578-020-0374-z
Sun Z-W, David Allis C (2002) Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418(6893):104–108. https://doi.org/10.1038/nature00883
Suryavanshi S, Eddé B, Fox LA, Guerrero S, Hard R, Hennessey T, Kabi A et al (2010) Tubulin glutamylation regulates ciliary motility by altering inner dynein arm activity. Curr Biol 20(5):435–440. https://doi.org/10.1016/j.cub.2009.12.062
Svensson JP, Shukla M, Menendez-Benito V, Norman-Axelsson U, Audergon P, Sinha I, Tanny JC, Allshire RC, Ekwall K (2015) A Nucleosome turnover map reveals that the stability of histone H4 Lys20 methylation depends on histone recycling in transcribed chromatin. Genome Res 25(6):872–883. https://doi.org/10.1101/gr.188870.114
Tang L, Nogales E, Ciferri C (2010) Structure and function of SWI/SNF chromatin remodeling complexes and mechanistic implications for transcription. Prog Biophys Mol Biol 102(2–3):122–128. https://doi.org/10.1016/j.pbiomolbio.2010.05.001
Tran K, Green EM (2019) SET domains and stress: uncovering new functions for yeast Set4. Curr Genet 65(3):643–648. https://doi.org/10.1007/s00294-018-0917-6
Valencia AM, Kadoch C (2019) Chromatin regulatory mechanisms and therapeutic opportunities in cancer. Nat Cell Biol 21(2):152–161. https://doi.org/10.1038/s41556-018-0258-1
Valencia-Sánchez MI, De Ioannes P, Wang M, Truong DM, Lee R, Armache J-P, Boeke JD, Armache K-J (2021) Regulation of the Dot1 histone H3K79 methyltransferase by histone H4K16 acetylation. Science. https://doi.org/10.1126/science.abc6663
Venkatesh S, Smolle M, Li H, Gogol MM, Saint M, Kumar S, Natarajan K, Workman JL (2012) Set2 methylation of histone H3 lysine 36 suppresses histone exchange on transcribed genes. Nature 489(7416):452–455. https://doi.org/10.1038/nature11326
Villaseñor R, Baubec T (2021) Regulatory mechanisms governing chromatin organization and function. Curr Opin Cell Biol Cell Nucl 70:10–17. https://doi.org/10.1016/j.ceb.2020.10.015
Vlaming H, Molenaar TM, van Welsem T, Poramba-Liyanage DW, Smith DE, Velds A, Hoekman L et al (2016) Direct screening for chromatin status on DNA barcodes in yeast delineates the regulome of H3K79 methylation by Dot1 Edited by Cynthia Wolberger. ELife 5:e18919. https://doi.org/10.7554/eLife.18919
Vlaming H, van Welsem T, de Graaf EL, David Ontoso AF, Altelaar M, San-Segundo PA, Heck AJR, van Leeuwen F (2014) Flexibility in crosstalk between H2B ubiquitination and H3 methylation in vivo. EMBO Rep 15(10):1077–1084. https://doi.org/10.1525/embr.201438793
Vlaming H, McLean CM, Korthout T, Alemdehy MF, Hendriks S, Lancini C, Palit S et al (2019) Conserved crosstalk between histone deacetylation and H3K79 methylation generates DOT1L-dose dependency in HDAC1-deficient thymic lymphoma. EMBO J 38(14):e101564. https://doi.org/10.15252/embj.2019101564
Wagner EJ, Carpenter PB (2012) Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13(2):115–126. https://doi.org/10.1038/nrm3274
Wang J, Liu Li, Yang Qu, Wei ** Yu, **a QB, **ong Y, Long Q, Jiejie Xu, Guo J (2016) Prognostic value of SETD2 expression in patients with metastatic renal cell carcinoma treated with tyrosine kinase inhibitors. J Urol 196(5):1363–1370. https://doi.org/10.1016/j.juro.2016.06.010
Wang Yi, Niu Y, Li B (2015) Balancing Acts of SRI and an Auto-inhibitory domain specify Set2 function at transcribed chromatin. Nucleic Acids Res 43(10):4881–4892. https://doi.org/10.1093/nar/gkv393
Weinberg DN, Papillon-Cavanagh S, Chen H, Yue Y, Chen X, Rajagopalan KN, Horth C et al (2019) The histone Mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature 573(7773):281–286. https://doi.org/10.1038/s41586-019-1534-3
Wen H, Li Y, ** Y, Jiang S, Stratton S, Peng D, Tanaka K et al (2014) ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression. Nature 508(7495):263–268. https://doi.org/10.1038/nature13045
Wojcik F, Dann GP, Beh LY, Debelouchina GT, Hofmann R, Muir TW (2018) Functional crosstalk between histone H2B ubiquitylation and H2A modifications and variants. Nat Commun 9(1):1394. https://doi.org/10.1038/s41467-018-03895-5
Wood A, Schneider J, Dover J, Johnston M, Shilatifard A (2005) The Bur1/Bur2 complex is required for histone H2B monoubiquitination by Rad6/Bre1 and histone methylation by compass. Mol Cell 20(4):589–599. https://doi.org/10.1016/j.molcel.2005.09.010
Wood K, Tellier M, Murphy S (2018) DOT1L and H3K79 methylation in transcription and genomic stability. Biomolecules. https://doi.org/10.3390/biom8010011
Wu G, Broniscer A, McEachron TA, Charles Lu, Paugh BS, Becksfort J, Chunxu Qu et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44(3):251–253. https://doi.org/10.1038/ng.1102
**ao C, Fan T, Tian He, Zheng Y, Zhou Z, Li S, Li C, He J (2021) H3K36 trimethylation-mediated biological functions in cancer. Clin Epigenetics 13(1):199. https://doi.org/10.1186/s13148-021-01187-2
**ao T, Hall H, Kizer KO, Shibata Y, Hall MC, Borchers CH, Strahl BD (2003) Phosphorylation of RNA polymerase II CTD regulates H3 methylation in yeast. Genes Dev 17(5):654–663. https://doi.org/10.1101/gad.1055503
**e Y, Sahin M, Sinha S, Wang Y, Nargund AM, Lyu Y, Han S et al (2022) SETD2 loss perturbs the kidney cancer epigenetic landscape to promote metastasis and engenders actionable dependencies on histone chaperone complexes. Nat Cancer 3(2):188–202. https://doi.org/10.1038/s43018-021-00316-3
Xu W, Li J, Rong B, Zhao B, Wang M, Dai R, Chen Q et al (2020) DNMT3A reads and connects histone H3K36me2 to DNA methylation. Protein Cell 11(2):150–154. https://doi.org/10.1007/s13238-019-00672-y
Yaakov G, Jonas F, Barkai N (2021) Measurement of histone replacement dynamics with genetically encoded exchange timers in yeast. Nat Biotechnol 39(11):1434–1443. https://doi.org/10.1038/s41587-021-00959-8
Yadav RK, Jablonowski CM, Fernandez AG, Lowe BR, Henry RA, Finkelstein D, Barnum KJ et al (2017) Histone H3G34R mutation causes replication stress, homologous recombination defects and genomic instability in S. Pombe. Elife 6:e27406. https://doi.org/10.7554/eLife.27406
Yang H, Kwon CS, Choi Y, Lee D (2016) Both H4K20 Mono-methylation and H3K56 acetylation mark transcription-dependent histone turnover in fission yeast. Biochem Biophys Res Commun 476(4):515–521. https://doi.org/10.1016/j.bbrc.2016.05.155
Yang S, Zheng X, Chao Lu, Guo-Min Li C, Allis D, Li H (2016) Molecular basis for oncohistone H3 recognition by SETD2 methyltransferase. Genes Dev 30(14):1611–1616. https://doi.org/10.1101/gad.284323.116
Yoh SM, Cho H, Pickle L, Evans RM, Jones KA (2007) The Spt6 SH2 domain binds Ser2-P RNAPII to direct Iws1-dependent MRNA splicing and export. Genes Dev 21(2):160–174. https://doi.org/10.1101/gad.1503107
Yoh SM, Lucas JS, Jones KA (2008) The Iws1:Spt6:CTD complex controls cotranscriptional MRNA biosynthesis and HYPB/Setd2-mediated histone H3K36 methylation. Genes Dev 22(24):3422–3434. https://doi.org/10.1101/gad.1720008
Youdell ML, Kizer KO, Kisseleva-Romanova E, Fuchs SM, Duro E, Strahl BD, Mellor J (2008) Roles for Ctk1 and Spt6 in regulating the different methylation states of histone H3 lysine 36. Mol Cell Biol 28(16):4915–4926. https://doi.org/10.1128/MCB.00001-08
Young NL, Dere R (2021) Mechanistic insights into KDM4A driven genomic instability. Biochem Soc Trans 49(1):93–105. https://doi.org/10.1042/BST20191219
Yuan H, Han Y, Wang X, Li Ni, Liu Q, Yin Y, Wang H et al (2020) SETD2 restricts prostate cancer metastasis by integrating EZH2 and AMPK signaling pathways. Cancer Cell 38(3):350-365.e7. https://doi.org/10.1016/j.ccell.2020.05.022
Yuan W, Mo Xu, Huang C, Liu N, Chen S, Zhu B (2011) H3K36 methylation antagonizes PRC2-mediated H3K27 methylation. J Biol Chem 286(10):7983–7989. https://doi.org/10.1074/jbc.M110.194027
Zhang H, Gao L, Anandhakumar J, Gross DS (2014) Uncoupling transcription from covalent histone modification. PLoS Genet. https://doi.org/10.1371/journal.pgen.1004202
Zhang P, Jiamu Du, Sun B, Dong X, Guoliang Xu, Zhou J, Huang Q, Liu Q, Hao Q, Ding J (2006) Structure of human MRG15 chromo domain and its binding to Lys36-methylated histone H3. Nucleic Acids Res 34(22):6621–6628. https://doi.org/10.1093/nar/gkl989
Zhang **, Huang Y, Shi X (2015) Emerging roles of lysine methylation on non-histone proteins. Cell Mol Life Sci 72(22):4257–4272. https://doi.org/10.1007/s00018-015-2001-4
Zhang Yi (2003) Transcriptional regulation by histone ubiquitination and deubiquitination. Genes Dev 17(22):2733–2740. https://doi.org/10.1101/gad.1156403
Zhang Y, Shan C-M, Wang J, Bao K, Tong L, Jia S (2017) Molecular basis for the role of oncogenic histone mutations in modulating H3K36 methylation. Sci Rep 7(1):43906. https://doi.org/10.1038/srep43906
Zhang Y-L, Sun J-W, **e Y-Y, Zhou Y, Liu P, Song J-C, Chun-Hui Xu et al (2018) Setd2 deficiency impairs hematopoietic stem cell self-renewal and causes malignant transformation. Cell Res 28(4):476–490. https://doi.org/10.1038/s41422-018-0015-9
Zhao S, David Allis C, Wang GG (2021) The language of chromatin modification in human cancers. Nat Rev Cancer 21(7):413–430. https://doi.org/10.1038/s41568-021-00357-x
Zhu Q, Yang Q, **aopeng Lu, Wang H, Tong L, Li Z, Liu Ge et al (2021) SETD2-mediated H3K14 trimethylation promotes ATR activation and stalled replication fork restart in response to DNA replication stress. Proc Natl Acad Sci USA 118(23):e2011278118. https://doi.org/10.1073/pnas.2011278118
Acknowledgements
We thank Elzo de Wit, Marlize van Breugel and Willem-Jan de Leeuw for carefully reading the manuscript and for valuable suggestions.
Funding
This work was supported by funding from The Dutch Research Council (NWO-VICI-016.130.627 to FvL) and the Dutch Cancer Society (KWF-NKI2018-1/11490 to FvL). The funders had no role in writing of the manuscript or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
TMM and FvL evaluated the literature and wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Molenaar, T.M., van Leeuwen, F. SETD2: from chromatin modifier to multipronged regulator of the genome and beyond. Cell. Mol. Life Sci. 79, 346 (2022). https://doi.org/10.1007/s00018-022-04352-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04352-9