Abstract
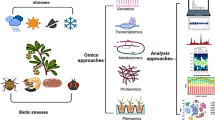
This book chapter discusses the application of integrated OMIC approaches for bioenergy crops. OMIC technologies such as genomics, transcriptomics, proteomics, and metabolomics have been increasingly used to identify genetic factors and metabolic pathways that play important roles in the growth and development of bioenergy crops. By integrating multiple OMIC datasets, researchers can gain a more comprehensive understanding of the molecular mechanisms underlying plant growth and development as well as responses to environmental stresses. The chapter provides an overview of the latest advances in integrated OMIC approaches for bioenergy crops, including the development of high-throughput sequencing platforms, bioinformatic tools, and computational models for data analysis. It also highlights some of the key challenges in applying these approaches, such as data integration and interpretation and the need for further validation of findings through experimental approaches. Furthermore, the chapter showcases some case studies that illustrate the use of integrated OMIC approaches in bioenergy crop research. These case studies cover a range of bioenergy crops, including switchgrass, corn, and sugarcane, and highlight the potential of OMIC approaches to improve plant growth, yield, and quality as well as to enhance stress tolerance and resilience. Overall, this book chapter provides an in-depth exploration of the current state of integrated OMIC approaches for bioenergy crops and highlights their potential for advancing the development of sustainable bioenergy production systems.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Brunner AM, Nilsson O (2004) Revisiting tree maturation and floral initiation in the poplar functional genomics era. New Phytol 164:43–51
Cao P, Zhao Y, Wu F, **n D, Liu C, Wu X, Lv J, Chen Q, Qi Z (2022) Multi-omics techniques for soybean molecular breeding. Int J Mol Sci 23:4994
Cardoso-Silva CB, Costa EA, Mancini MC, Balsalobre TWA, Canesin LEC, Pinto LR, Carneiro MS, Garcia AAF, de Souza AP, Vicentini R (2014) De novo assembly and transcriptome analysis of contrasting sugarcane varieties. PLoS One 9:e88462
Casler MD, Tobias CM, Kaeppler SM, Buell CR, Wang ZY, Cao P, Schmutz J, Ronald P (2011) The switchgrass genome: tools and strategies. Plant Genome 4
Chung JN (2013) Grand challenges in bioenergy and biofuel research: engineering and technology development, environmental impact, and sustainability, vol 1. Frontiers Media SA, p 4
Deshmukh R, Sonah H, Patil G, Chen W, Prince S, Mutava R, Vuong T, Valliyodan B, Nguyen HT (2014) Integrating omic approaches for abiotic stress tolerance in soybean. Front Plant Sci 5:244
Frigon JC, Guiot SR (2010) Biomethane production from starch and lignocellulosic crops: a comparative review. Biofuels Bioprod Biorefin 4:447–458
Gomez-Cabrero D, Abugessaisa I, Maier D, Teschendorff A, Merkenschlager M, Gisel A, Ballestar E, Bongcam-Rudloff E, Conesa A, Tegnér J (2014) Data integration in the era of omics: current and future challenges. BMC Syst Biol 8:1–10
Guarnieri MT, Nag A, Smolinski SL, Darzins A, Seibert M, Pienkos PT (2011) Examination of triacylglycerol biosynthetic pathways via De Novo transcriptomic and proteomic analyses in an Unsequenced microalga. PLoS One 6:e25851
Hassan SS, Williams GA, Jaiswal AK (2019) Moving towards the second generation of lignocellulosic biorefineries in the EU: drivers, challenges, and opportunities. Renew Sust Energ Rev 101:590–599
Hu H, Dai M, Yao J, **ao B, Li X, Zhang Q, **ong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci 103:12987–12992
Jiang Z, Zhang H, Jiao P, Wei X, Liu S, Guan S, Ma Y (2022) The integration of metabolomics and transcriptomics provides new insights for the identification of genes key to auxin synthesis at different growth stages of maize. Int J Mol Sci 23:13195
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363
Koçar G, Civaş N (2013) An overview of biofuels from energy crops: current status and future prospects. Renew Sust Energ Rev 28:900–916
Lemus R, Lal R (2005) Bioenergy crops and carbon sequestration. Crit Rev Plant Sci 24:1–21
Li P, Ponnala L, Gandotra N, Wang L, Si Y, Tausta SL, Kebrom TH, Provart N, Patel R, Myers CR, Reidel EJ, Turgeon R, Liu P, Sun Q, Nelson T, Brutnell TP (2010) The developmental dynamics of the maize leaf transcriptome. Nat Genet 42:1060–1067
Lu F, Romay MC, Glaubitz JC, Bradbury PJ, Elshire RJ, Wang T, Li Y, Li Y, Semagn K, Zhang X (2015) High-resolution genetic map** of maize pan-genome sequence anchors. Nat Commun 6:1–8
Ma X-F, Jensen E, Alexandrov N, Troukhan M, Zhang L, Thomas-Jones S, Farrar K, Clifton-Brown J, Donnison I, Swaller T (2012) High resolution genetic map** by genome sequencing reveals genome duplication and tetraploid genetic structure of the diploid Miscanthus sinensis. PLoS One 7:e33821
Ma J, Wan D, Duan B, Bai X, Bai Q, Chen N, Ma T (2019) Genome sequence and genetic transformation of a widely distributed and cultivated poplar. Plant Biotechnol J 17:451–460
Meng J, Wang B, He G, Wang Y, Tang X, Wang S, Ma Y, Fu C, Chai G, Zhou G (2019) Metabolomics integrated with transcriptomics reveals redirection of the phenylpropanoids metabolic flux in Ginkgo biloba. J Agric Food Chem 67:3284–3291
Miao J, Feng Q, Li Y, Zhao Q, Zhou C, Lu H, Fan D, Yan J, Lu Y, Tian Q (2021) Chromosome-scale assembly and analysis of biomass crop miscanthus lutarioriparius genome. Nat Commun 12:2458
Misra N, Panda P, Parida B, Mishra B (2016) Way forward to achieve sustainable and cost-effective biofuel production from microalgae: a review. Int J Environ Sci Technol 13:2735–2756
Misra BB, Langefeld C, Olivier M, Cox LA (2019) Integrated omics: tools, advances and future approaches. J Mol Endocrinol 62:R21–R45
Mouratiadou I, Stella T, Gaiser T, Wicke B, Nendel C, Ewert F, van der Hilst F (2020) Sustainable intensification of crop residue exploitation for bioenergy: opportunities and challenges. GCB Bioenergy 12:71–89
Poudel S, Giannone RJ, Rodriguez M, Raman B, Martin MZ, Engle NL, Mielenz JR, Nookaew I, Brown SD, Tschaplinski TJ (2017) Integrated omics analyses reveal the details of metabolic adaptation of clostridium thermocellum to lignocellulose-derived growth inhibitors released during the deconstruction of switchgrass. Biotechnol Biofuels 10:1–14
Ramos-Madrigal J, Smith BD, Moreno-Mayar JV, Gopalakrishnan S, Ross-Ibarra J, Gilbert MTP, Wales N (2016) Genome sequence of a 5,310-year-old maize cob provides insights into the early stages of maize domestication. Curr Biol 26:3195–3201
Reid WV, Ali MK, Field CB (2020) The future of bioenergy. Glob Chang Biol 26:274–286
Riaño-Pachón DM, Mattiello L (2017) Draft genome sequencing of the sugarcane hybrid SP80-3280. F1000Research 6
Sablok G, Fu Y, Bobbio V, Laura M, Rotino GL, Bagnaresi P, Allavena A, Velikova V, Viola R, Loreto F, Li M, Varotto C (2014) Fuelling genetic and metabolic exploration of C3 bioenergy crops through the first reference transcriptome of Arundo donax L. Plant Biotechnol J 12:554–567
Sharma MK, Sharma R, Cao P, Jenkins J, Bartley LE, Qualls M, Grimwood J, Schmutz J, Rokhsar D, Ronald PC (2012) A genome-wide survey of switchgrass genome structure and organization. PLoS One 7:e33892
Sheng J, Yan M, Wang J, Zhao L, Zhou F, Hu Z, ** S, Diao Y (2021) The complete chloroplast genome sequences of five miscanthus species, and comparative analyses with other grass plastomes. Ind Crop Prod 162:113248
Slade R, Bauen A, Gross R (2014) Global bioenergy resources. Nat Clim Change 4:99–105
Stamenković OS, Siliveru K, Veljković VB, Banković-Ilić IB, Tasić MB, Ciampitti IA, Đalović IG, Mitrović PM, Sikora VŠ, Prasad PV (2020) Production of biofuels from sorghum. Renew Sust Energ Rev 124:109769
Thirugnanasambandam PP, Hoang NV, Henry RJ (2018) The challenge of analyzing the sugarcane genome. Front Plant Sci 9:616
Tiedge K, Li X, Merrill AT, Davisson D, Chen Y, Yu P, Tantillo DJ, Last RL, Zerbe P (2022) Comparative transcriptomics and metabolomics reveal specialized metabolite drought stress responses in switchgrass (Panicum virgatum). New Phytol 236:1393–1408
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Déjardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjärvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leplé JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouzé P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P et al (2006) The genome of black cottonwood, Populus trichocarpa (torr. & gray). Science 313:1596–1604
Valentine J, Clifton-Brown J, Hastings A, Robson P, Allison G, Smith P (2012) Food vs. fuel: the use of land for lignocellulosic ‘next generation’energy crops that minimize competition with primary food production. GCB Bioenergy 4:1–19
Vilanova C, Porcar M (2016) Are multi-omics enough? Nat Microbiol 1:1–2
Yuan JS, Tiller KH, Al-Ahmad H, Stewart NR, Stewart CN (2008) Plants to power: bioenergy to fuel the future. Trends Plant Sci 13:421–429
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mahmood, A. et al. (2023). Integrated OMIC Approaches for Bioenergy Crops. In: Aasim, M., Baloch, F.S., Nadeem, M.A., Habyarimana, E., Ahmed, S., Chung, G. (eds) Biotechnology and Omics Approaches for Bioenergy Crops. Springer, Singapore. https://doi.org/10.1007/978-981-99-4954-0_4
Download citation
DOI: https://doi.org/10.1007/978-981-99-4954-0_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-4953-3
Online ISBN: 978-981-99-4954-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)