Abstract
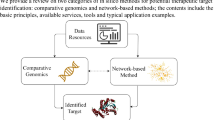
Drug discovery has predominantly followed the concept of “one drug, one target, one disease” for an extended period, case in point, to design chemical entities that can specifically bind to one key target related to a specific disease [1]. Collateral pharmacology aims to develop drugs that earmark multiple proteins or networks connected to diseases. It also demonstrates the possibility of finding multi-target and multi-component drugs that earmark disease-related networks at the system level [2]. Network pharmacology research integrates the data of various public databases, high-throughput screening (HTS), genome-wide association studies (GWAS), and large-scale omics (such as genomics, transcriptomics, metabonomics, and proteomics) to construct a network prediction or inference model. The analysis of the complex biological pathways influenced by drug therapy at different biological levels (molecules, cells, tissues, organs, and phenotypes) has given a boost to cognition of the biological mechanisms of complex diseases, the systemic mechanism of the impact of drugs, and the development of multi-target, multi-component drugs. This chapter selects some exceptional results of network pharmacology in the R&D and application of modern drugs in recent years, and analyzes the results from the dimensions of research purpose, data source, analysis index and algorithm, analysis results, experimental verification, and main conclusion, as a means to introduce the principal research contents, ideas, and procedures of frontier research in network pharmacology, for readers.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kaufmann SH. Paul Ehrlich: founder of chemotherapy. Nat Rev Drug Discov. 2008;7(5):373.
Hopkins AL. Network pharmacology: the next paradigm in drug discovery. Nat Chem Biol. 2008;4(11):682.
Oliver S. Guilt-by-association goes global. Nature. 2000;403(6770):601–2.
Wu X, Jiang R, Zhang MQ, et al. Network-based global inference of human disease genes. Mol Syst Biol. 2008;4:1.
Lin L, Yang T, Fang L, et al. Gene gravity-like algorithm for disease gene prediction based on phenotype-specific network. BMC Syst Biol. 2017;11(1):121.
Zhang B, Gaiteri C, Bodea L-G, et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell. 2013;153(3):707–20.
Mani KM, Lefebvre C, Wang K, et al. A systems biology approach to prediction of oncogenes and molecular perturbation targets in B-cell lymphomas. Mol Syst Biol. 2008;4:1.
Menche J, Sharma A, Kitsak M, et al. Uncovering disease-disease relationships through the incomplete interactome. Science. 2015;347(6224):1257601.
Chen C, Meng Q, **a Y, et al. The transcription factor POU3F2 regulates a gene coexpression network in brain tissue from patients with psychiatric disorders. Sci Transl Med. 2018;10(472):eaat8178.
Torrey EF, Webster M, Knable M, et al. The stanley foundation brain collection and neuropathology consortium. Schizophr Res. 2000;44(2):151–5.
Gibbs JR, van der Brug MP, Hernandez DG, et al. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet. 2010;6:5.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 2008;9(1):559.
Aten JE, Fuller TF, Lusis AJ, et al. Using genetic markers to orient the edges in quantitative trait networks: the NEO software. BMC Syst Biol. 2008;2(1):34.
de Leeuw CA, Mooij JM, Heskes T, et al. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:4.
Lee PH, O'dushlaine C, Thomas B, et al. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics. 2012;28(13):1797–9.
Purcell SM, Moran JL, Fromer M, et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature. 2014;506(7487):185–90.
Genovese G, Fromer M, Stahl EA, et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat Neurosci. 2016;19(11):1433.
Li J, Cai T, Jiang Y, et al. Genes with de novo mutations are shared by four neuropsychiatric disorders discovered from NPdenovo database. Mol Psychiatry. 2016;21(2):290–7.
Bass JIF, Sahni N, Shrestha S, et al. Human gene-centered transcription factor networks for enhancers and disease variants. Cell. 2015;161(3):661–73.
Kheradpour P, Kellis M. Systematic discovery and characterization of regulatory motifs in ENCODE TF binding experiments. Nucleic Acids Res. 2014;42(5):2976–87.
Guo Y, Bao C, Ma D, et al. Network-based combinatorial crispr-cas9 screens identify synergistic modules in human cells. ACS Synth Biol. 2019;8(3):482–90.
Schäfer H, Struck B, Feldmann E, et al. TGF-β1-dependent L1CAM expression has an essential role in macrophage-induced apoptosis resistance and cell migration of human intestinal epithelial cells. Oncogene. 2013;32(2):180–9.
Bassik MC, Kampmann M, Lebbink RJ, et al. A systematic mammalian genetic interaction map reveals pathways underlying ricin susceptibility. Cell. 2013;152(4):909–22.
Kampmann M, Bassik MC, Weissman JS. Integrated platform for genome-wide screening and construction of high-density genetic interaction maps in mammalian cells. Proc Natl Acad Sci. 2013;110(25):E2317–26.
Bandyopadhyay S, Mehta M, Kuo D, et al. Rewiring of genetic networks in response to DNA daMage. Science. 2010;330(6009):1385–9.
Lamb J, Crawford ED, Peck D, et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 2006;313(5795):1929–35.
Subramanian A, Narayan R, Corsello SM, et al. A Next generation connectivity map: l1000 platform and the first 1,000,000 profiles. Cell. 2017;171(6):1437–52.
Iwata M, Sawada R, Iwata H, et al. Elucidating the modes of action for bioactive compounds in a cell-specific manner by large-scale chemically-induced transcriptomics. Sci Rep. 2017;7:40164.
Guney E, Menche J, Vidal M, et al. Network-based in silico drug efficacy screening. Nat Commun. 2016;7:10331.
Wei W-Q, Cronin RM, Xu H, et al. Development and evaluation of an ensemble resource linking medications to their indications. J Am Med Inform Assoc. 2013;20(5):954–61.
Sartor MA, Ade A, Wright Z, et al. Metab2MeSH: annotating compounds with medical subject headings. Bioinformatics. 2012;28(10):1408–10.
Iorio F, Bosotti R, Scacheri E, et al. Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc Natl Acad Sci. 2010;107(33):14621–6.
Kruskal JB. On the shortest spanning subtree of a graph and the traveling salesman problem. Proc Am Math Soc. 1956;7(1):48–50.
Diaconis P, Graham RL. Spearman’s footrule as a measure of disarray. J R Stat Soc. 1977;39(2):262–8.
Lin S. Space oriented rank-based data integration. Stat Appl Genet Mol Biol. 2010;9:1.
Frey BJ, Dueck D. Clustering by passing messages between data points. Science. 2007;315(5814):972–6.
Csermely P, Agoston V, Pongor S. The efficiency of multi-target drugs: the network approach might help drug design. Trends Pharmacol Sci. 2005;26(4):178–82.
Keith CT, Borisy AA, Stockwell BR. Multicomponent therapeutics for networked systems. Nat Rev Drug Discov. 2005;4(1):71–8.
Korcsmáros T, Szalay MS, Böde C, et al. How to design multi-target drugs: target search options in cellular networks. Expert Opin Drug Discovery. 2007;2(6):799–808.
Ramsay RR, Popovic-Nikolic MR, Nikolic K, et al. A perspective on multi-target drug discovery and design for complex diseases. Clin Transl Med. 2018;7(1):3.
Meng H, Liu Y, Lai L. Diverse ways of perturbing the human arachidonic acid metabolic network to control inflammation. Acc Chem Res. 2015;48(8):2242–50.
Keane H, Ryan BJ, Jackson B, et al. Protein-protein interaction networks identify targets which rescue the MPP+ cellular model of Parkinson’s disease. Sci Rep. 2015;5(1):1–12.
Casas AI, Hassan AA, Larsen SJ, et al. From single drug targets to synergistic network pharmacology in ischemic stroke. Proc Natl Acad Sci. 2019;116(14):7129–36.
Kotlyar M, Pastrello C, Sheahan N, et al. Integrated interactions database: tissue-specific view of the human and model organism interactomes. Nucleic Acids Res. 2016;44(D1):D536–41.
Wishart DS, Feunang YD, Marcu A, et al. HMDB 4.0: the human metabolome database for. Nucleic Acids Res. 2018;46(D1):D608–17.
Li YH, Yu CY, Li XX, et al. Therapeutic target database update 2018: enriched resource for facilitating bench-to-clinic research of targeted therapeutics. Nucleic Acids Res. 2018;46(D1):D1121–7.
Wishart DS, Tzur D, Knox C, et al. HMDB: the human metabolome database. Nucleic Acids Res. 2007;35(suppl_1):D521–6.
Sidders B, Karlsson A, Kitching L, et al. Network-based drug discovery: coupling network pharmacology with phenotypic screening for neuronal excitability. J Mol Biol. 2018;430(18):3005–15.
Jamieson DG, Moss A, Kennedy M, et al. The pain interactome: connecting pain-specific protein interactions. Pain. 2014;155(11):2243–52.
Razick S, Magklaras G, Donaldson IM. iRefIndex: a consolidated protein interaction database with provenance. BMC Bioinf. 2008;9(1):405.
Pushpakom S, Iorio F, Eyers PA, et al. Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov. 2019;18(1):41–58.
Zhang Y, Cheng X, Zhou W. Drug reorientation: an important application field of network pharmacology. Chin J Pharmacol Toxicol. 2012;26(6):779–86.
Cheng F, Liu C, Jiang J, et al. Prediction of drug-target interactions and drug repositioning via network-based inference. PLoS Comput Biol. 2012;8(5):e1002503.
Iorio F, Isacchi A, di Bernardo D, et al. Identification of small molecules enhancing autophagic function from drug network analysis. Autophagy. 2010;6(8):1204–5.
Luo H, Wang J, Li M, et al. Drug repositioning based on comprehensive similarity measures and bi-random walk algorithm. Bioinformatics. 2016;32(17):2664–71.
Cheng F, Desai RJ, Handy DE, et al. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat Commun. 2018;9(1):1–12.
Rolland T, Taşan M, Charloteaux B, et al. A proteome-scale map of the human interactome network. Cell. 2014;159(5):1212–26.
Rual J-F, Venkatesan K, Hao T, et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature. 2005;437(7062):1173–8.
Cheng F, Jia P, Wang Q, et al. Quantitative network map** of the human kinome interactome reveals new clues for rational kinase inhibitor discovery and individualized cancer therapy. Oncotarget. 2014;5(11):3697.
Keshava Prasad T, Goel R, Kandasamy K, et al. Human protein reference database—2009 update. Nucleic Acids Res. 2009;37(suppl_1):D767–72.
Hu J, Rho H-S, Newman RH, et al. PhosphoNetworks: a database for human phosphorylation networks. Bioinformatics. 2014;30(1):141–2.
Hornbeck PV, Zhang B, Murray B, et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 2015;43(D1):D512–20.
Lu C-T, Huang K-Y, Su M-G, et al. DbPTM 3.0: an informative resource for investigating substrate site specificity and functional association of protein post-translational modifications. Nucleic Acids Res. 2013;41(D1):D295–305.
Dinkel H, Chica C, Via A, et al. Phospho. ELM: a database of phosphorylation sites—update 2011. Nucleic Acids Res. 2010;39(suppl_1):D261–7.
Oughtred R, Stark C, Breitkreutz B-J, et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019;47(D1):D529–41.
Cowley MJ, Pinese M, Kassahn KS, et al. PINA v2. 0: mining interactome modules. Nucleic Acids Res. 2012;40(D1):D862–5.
Chatr-Aryamontri A, Ceol A, Palazzi LM, et al. MINT: the Molecular INTeraction database. Nucleic Acids Res. 2007;35(suppl_1):D572–4.
Orchard S, Ammari M, Aranda B, et al. The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 2014;42(D1):D358–63.
Breuer K, Foroushani AK, Laird MR, et al. InnateDB: systems biology of innate immunity and beyond—recent updates and continuing curation. Nucleic Acids Res. 2013;41(D1):D1228–33.
Meyer MJ, Das J, Wang X, et al. INstruct: a database of high-quality 3D structurally resolved protein interactome networks. Bioinformatics. 2013;29(12):1577–9.
Fazekas D, Koltai M, Türei D, et al. SignaLink 2–a signaling pathway resource with multi-layered regulatory networks. BMC Syst Biol. 2013;7(1):7.
Bodenreider O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 2004;32(suppl_1):D267–70.
Hamosh A, Scott AF, Amberger JS, et al. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2005;33(suppl_1):D514–7.
Davis AP, Grondin CJ, Lennon-Hopkins K, et al. The comparative toxicogenomics database’s 10th year anniversary: update 2015. Nucleic Acids Res. 2015;43(D1):D914–20.
Yu W, Gwinn M, Clyne M, et al. A navigator for human genome epidemiology. Nat Genet. 2008;40(2):124–5.
Piñero J, Bravo À, Queralt-Rosinach N, et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2016;2016:gkw943.
Landrum MJ, Lee JM, Riley GR, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42(D1):D980–5.
Welter D, Macarthur J, Morales J, et al. The NHGRI GWAS catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014;42(D1):D1001–6.
Li MJ, Liu Z, Wang P, et al. GWASdb v2: an update database for human genetic variants identified by genome-wide association studies. Nucleic Acids Res. 2016;44(D1):D869–76.
Denny JC, Bastarache L, Ritchie MD, et al. Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat Biotechnol. 2013;31(12):1102.
Wishart DS, Feunang YD, Guo AC, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46(D1):D1074–82.
Hernandez-Boussard T, Whirl-Carrillo M, Hebert JM, et al. The pharmacogenetics and pharmacogenomics knowledge base: accentuating the knowledge. Nucleic Acids Res. 2007;36(suppl_1):D913–8.
Bento AP, Gaulton A, Hersey A, et al. The ChEMBL bioactivity database: an update. Nucleic Acids Res. 2014;42(D1):D1083–90.
Gilson MK, Liu T, Baitaluk M, et al. BindingDB in 2015: a public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res. 2016;44(D1):D1045–53.
Pawson AJ, Sharman JL, Benson HE, et al. The IUPHAR/BPS Guide to pharmacology: an expert-driven knowledgebase of drug targets and their ligands. Nucleic Acids Res. 2014;42(D1):D1098–106.
Consortium G. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348(6235):648–60.
Wang S, Verpillat P, Rassen J, et al. Transparency and reproducibility of observational cohort studies using large healthcare databases. Clin Pharmacol Therap. 2016;99(3):325–32.
Zhang T-T, Xue R, Wang X, et al. Network-based drug repositioning: a novel strategy for discovering potential antidepressants and their mode of action. Eur Neuropsychopharmacol. 2018;28(10):1137–50.
Zhao S, Li S. Network-based relating pharmacological and genomic spaces for drug target identification. PLoS One. 2010;5:7.
Cheng F, Kovács IA, Barabási A-L. Network-based prediction of drug combinations. Nat Commun. 2019;10(1):1–11.
Li W, Mao X, Guo Q, et al. Application of network pharmacology research strategy in combinatorial drug research. E-j Transl Med. 2018;5(3):3–16.
Wang Y-Y, Xu K-J, Song J, et al. Exploring drug combinations in genetic interaction network. BMC Bioinf. 2012;2017:S7.
Cheng F, Zhao Z. Machine learning-based prediction of drug–drug interactions by integrating drug phenotypic, therapeutic, chemical, and genomic properties. J Am Med Inform Assoc. 2014;21(e2):e278–86.
Chandrasekaran S, Cokol-Cakmak M, Sahin N, et al. Chemogenomics and orthology-based design of antibiotic combination therapies. Mol Syst Biol. 2016;12:5.
Li S, Zhang B, Zhang N. Network target for screening synergistic drug combinations with application to traditional Chinese medicine. BMC Syst Biol. 2011;5(S1):S10.
He L, Wennerberg K, Aittokallio T, et al. TIMMA-R: an R package for predicting synergistic multi-targeted drug combinations in cancer cell lines or patient-derived samples. Bioinformatics. 2015;31(11):1866–8.
Huang L, Li F, Sheng J, et al. DrugComboRanker: drug combination discovery based on target network analysis. Bioinformatics. 2014;30(12):i228–36.
Gaulton A, Bellis LJ, BENTO AP, et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012;40(D1):D1100–7.
LIU T, LIN Y, WEN X, et al. BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 2007;35(suppl_1):D198–201.
Tatonetti NP, Patrick PY, Daneshjou R, et al. Data-driven prediction of drug effects and interactions. Sci Transl Med. 2012;4(125):125ra131.
SUN Y, SHENG Z, MA C, et al. Combining genomic and network characteristics for extended capability in predicting synergistic drugs for cancer. Nat Commun. 2015;6(1):1–10.
Liu Y, Wei Q, Yu G, et al. DCDB 2.0: a major update of the drug combination database. Database. 2014;2014:124.
Sun Y, Sheng Z, Ma C, et al. Combining genomic and network characteristics for extended capability in predicting synergistic drugs for cancer. Nat Commun. 2015;28(6):8481.
Liu Y, Wei Q, Yu G, Gai W, Li Y, Chen X. DCDB 2.0: a major update of the drug combination database. Database (Oxford). 2014;2014:bau124.
Bansal M, Yang J, Karan C, et al. A community computational challenge to predict the activity of pairs of compounds. Nat Biotechnol. 2014;32(12):1213–22.
Barretina J, Caponigro G, Stransky N, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483(7391):603–7.
Zhao J, Zhang XS, Zhang S. Predicting cooperative drug effects through the quantitative cellular profiling of response to individual drugs. CPT Pharmacometrics Syst Pharmacol. 2014;3(2):e102.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Zhang, W., Zhao, J. (2021). Network Pharmacology and Modern Drug R&D Cases. In: Li, S. (eds) Network Pharmacology. Springer, Singapore. https://doi.org/10.1007/978-981-16-0753-0_6
Download citation
DOI: https://doi.org/10.1007/978-981-16-0753-0_6
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-0752-3
Online ISBN: 978-981-16-0753-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)