Abstract
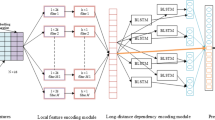
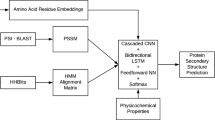
The growth of biological databases has been increased in the past decades makes a protein structure prediction is one of the most important and challenging problems in Bioinformatics. The recent growth of Neural Network that have been shown promising result and also became indispensable tools in Bioinformatics. Using Convolutional Neural Network and Recurrent Neural Network with Long-Short Term Memory to predicting the secondary structure, our experiment shows a high result for RNN LSTM with accuracy 88.74% and loss rate 3.64%. In addition, for the CNN methods with an accuracy 87.74% and loss rate 3.85% prove that the latest technology of neural network also effective to be applied in secondary structure prediction of the proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chen K, Kurgan LA (2012) Neural networks in bioinformatics. Handbook of natural computing, pp 565–583. Springer, Berlin, Heidelberg
Anfinsen CB, Haber E, Sela M, White FH Jr (1961) The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain. Proc Natl Acad Sci U S A 47(9):1309–1314
Zhang B, Li J, Lü Q (2018) Prediction of 8-state protein secondary structures by a novel deep learning architecture. BMC Bioinformatics. 19:293
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers: Original Res Biomolecules 22(12):2577–2637
Wang S, Peng J, Ma J, Xu J (2016) Protein secondary structure prediction using deep convolutional neural fields. Sci Rep 6:18962
Yang Y, Gao J, Wang J, Heffernan R, Hanson J, Paliwal K et al (2016) Sixty-five years of the long march in protein secondary structure prediction: the final stretch? Brief Bioinform 19(3):482–494
Protein secondary structure (2019). https://www.kaggle.com/alfrandom/protein-secondary-structure
Karplus K, Barrett C, Hughey R (1998) Hidden Markov models for detecting remote protein homologies. Bioinformatics (Oxford, England) 14(10):846–856
Ward JJ, McGuffin LJ, Buxton BF, Jones DT (2003) Secondary structure prediction with support vector machines. Bioinformatics 19(13):1650–1655
Babaei S, Geranmayeh A, Seyyedsalehi SA (2010) Protein secondary structure prediction using modular reciprocal bidirectional recurrent neural networks. Comput Methods Program Biomed 100(3):237–247
Prostruct (2019). https://github.com/vmalepati1/Prostruct
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2019R1F1A1058394), and the MSIP (Ministry of Science, ICT & Future Planning), Korea, under the National Program for Excellence in SW) (2018-0-01865) supervised by the IITP (Institute for Information & communications Technology Promotion).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Listio, S.W.P., Elbasani, E., Oh, TJ., Kim, B., Kim, JD. (2021). Deep Learning-Based Experimentation for Predicting Secondary Structure of Amino Acid Sequence. In: Park, J.J., Fong, S.J., Pan, Y., Sung, Y. (eds) Advances in Computer Science and Ubiquitous Computing. Lecture Notes in Electrical Engineering, vol 715. Springer, Singapore. https://doi.org/10.1007/978-981-15-9343-7_8
Download citation
DOI: https://doi.org/10.1007/978-981-15-9343-7_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-9342-0
Online ISBN: 978-981-15-9343-7
eBook Packages: Computer ScienceComputer Science (R0)