Abstract
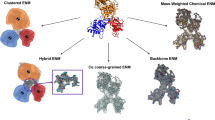
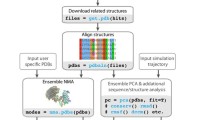
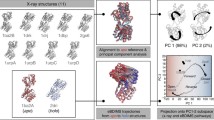
Large biomolecules are involved in many important biological processes. It would be difficult to use large-scale atomistic molecular dynamics (MD) simulations to study the functional motions of these systems because of the computational expense. Therefore various coarse-grained (CG) approaches have attracted rapidly growing interest, which enable simulations of large biomolecules over longer effective timescales than all-atom MD simulations. The first issue in CG modeling is to construct CG maps from atomic structures. In this chapter, we review the recent development of a novel and systematic method for constructing CG representations of arbitrarily complex biomolecules, in order to preserve large-scale and functionally relevant essential dynamics (ED) at the CG level. In this ED-CG scheme, the essential dynamics can be characterized by principal component analysis (PCA) on a structural ensemble, or elastic network model (ENM) of a single atomic structure. Validation and applications of the method cover various biological systems, such as multi-domain proteins, protein complexes, and even biomolecular machines. The results demonstrate that the ED-CG method may serve as a very useful tool for identifying functional dynamics of large biomolecules at the CG level.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dror RO, Dirks RM, Grossman JP, Xu HF, Shaw DE (2012) Biomolecular simulation: a computational microscope for molecular biology. Annu Rev Biophys 41:429–452
Tozzini V (2005) Coarse-grained models of proteins. Curr Opin Struct Biol 15:144–150
Ayton GS, Noid WG, Voth GA (2007) Multiscale modeling of biomolecular systems: in serial and in parallel. Curr Opin Struct Biol 17:192–198
Murtola T, Bunker A, Vattulainen I, Deserno M, Karttunen M (2009) Multiscale modeling of emergent materials: biological and soft matter. Phys Chem Chem Phys 11:1869–1892
Saunders MG, Voth GA (2012) Coarse-graining of multiprotein assemblies. Curr Opin Struct Biol 22:144–150
Voth GA (2009) Coarse-graining of condensed phase and biomolecular systems. CRC Press-Taylor & Francis Group, Boca Raton 2009
Marrink SJ, Risselada HJ, Yefimov S, Tieleman DP, de Vries AH (2007) The MARTINI force field: Coarse grained model for biomolecular simulations. J Phys Chem B 111:7812–7824
Monticelli L, Kandasamy SK, Periole X, Larson RG, Tieleman DP, Marrink SJ (2008) The MARTINI coarse-grained force field: extension to proteins. J Chem Theory Comput 4:819–834
Liwo A, He Y, Scheraga HA (2011) Coarse-grained force field: general folding theory. Phys Chem Chem Phys 13:16890–16901
Atilgan AR, Durell SR, Jernigan RL, Demirel MC, Keskin O, Bahar I (2001) Anisotropy of fluctuation dynamics of proteins with an elastic network model. Biophys J 80:505–515
Bahar I, Rader AJ (2005) Coarse-grained normal mode analysis in structural biology. Curr Opin Struct Biol 15:586–592
Bahar I, Lezon TR, Yang LW, Eyal E (2010) Global dynamics of proteins: bridging between structure and function. Annu Rev Biophys 39:23–42
Arkhipov A, Freddolino PL, Schulten K (2006) Stability and dynamics of virus capsids described by coarse-grained modeling. Structure 14:1767–1777
Murtola T, Kupiainen M, Falck E, Vattulainen I (2007) Conformational analysis of lipid molecules by self-organizing maps. J Chem Phys 126:17
Gohlke H, Thorpey MF (2006) A natural coarse graining for simulating large biomolecular motion. Biophys J 91:2115–2120
Stepanova M (2007) Dynamics of essential collective motions in proteins: theory. Phys Rev E 76:16
Gfeller D, De Los Rios P (2008) Spectral coarse graining and synchronization in oscillator networks. Phys Rev Lett 100:4
Zhang Z, Lu L, Noid WG, Krishna V, Pfaendtner J, Voth GA (2008) A systematic methodology for defining coarse-grained sites in large biomolecules. Biophys J 95:5073–5083
Zhang Z, Wriggers W (2008) Coarse-graining protein structures with local multivariate features from molecular dynamics. J Phys Chem B 112:14026–14035
Jang H, Na S, Eom K (2009) Multiscale network model for large protein dynamics. J Chem Phys 131:10
Potestio R, Pontiggia F, Micheletti C (2009) Coarse-grained description of protein internal dynamics: an optimal strategy for decomposing proteins in rigid subunits. Biophys J 96:4993–5002
Zhang Z, Pfaendtner J, Grafmüller A, Voth GA (2009) Defining coarse-grained representations of large biomolecules and biomolecular complexes from elastic network models. Biophys J 97:2327–2337
Zhang Z, Voth GA (2010) Coarse-grained representations of large biomolecular complexes from low-resolution structural data. J Chem Theory Comput 6:2990–3002
Amadei A, Linnsen ABM, Berendsen HJC (1993) Essential dynamics of proteins. Proteins 17:412–425
Kitao A, Go N (1999) Investigating protein dynamics in collective coordinate space. Curr Opin Struct Biol 9:164–169
Berendsen HJC, Hayward S (2000) Collective protein dynamics in relation to function. Curr Opin Struct Biol 10:165–169
de Groot BL, van Aalten DMF, Scheek RM, Amadei A, Vriend G, Berendsen HJC (1997) Prediction of protein conformational freedom from distance constraints. Proteins 29:240–251
Brooks B, Karplus M (1983) Harmonic dynamics of proteins: normal modes and fluctuations in bovine pancreatic trypsin inhibitor. Proc Natl Acad Sci USA 80:6571–6575
Cui Q, Bahar I (eds) (2006) Normal mode analysis: theory and applications to biological and chemical systems. Chapman & Hall/CRC, London
Hinsen K (2009) Physical arguments for distance-weighted interactions in elastic network models for proteins. Proc Natl Acad Sci USA 106:E128–E128
Yang L, Song G, Jernigan RL (2009) Protein elastic network models and the ranges of cooperativity. Proc Natl Acad Sci USA 106:12347–12352
Yesylevskyy SO, Kharkyanen VN, Demchenko AP (2006) Dynamic protein domains: Identification, interdependence, and stability. Biophys J 91:670–685
Flory PJ, Gordon M, McCrum NG (1976) Statistical thermodynamics of random networks [and discussion]. Proc R Soc Lond A Math Phys Sci 351:351–380
Kirkpatrick S, Gelatt Jr CD, Vecchi MP (1983) Optimization by simulated annealing. Science 220:671–680
Metropolis N, Rosenbluth AW, Rosenbluth MN, Teller AH, Teller E (1953) Equation of state calculations by fast computing machines. J Chem Phys 21:1087–1092
Krishna V, Ayton GS, Voth GA (2010) Role of protein interactions in defining HIV-1 viral capsid shape and stability: a coarse-grained analysis. Biophys J 98:18–26
Chu JW, Voth GA (2005) Allostery of actin filaments: molecular dynamics simulations and coarse-grained analysis. Proc Natl Acad Sci USA 102:13111–13116
Chu JW, Voth GA (2006) Coarse-grained modeling of the actin filament derived from atomistic-scale simulations. Biophys J 90:1572–1582
Fan J, Saunders MG, Voth GA (2012) Coarse-graining provides insights on the essential nature of heterogeneity in Actin filaments. Biophys J 103:1334–1342
Kabsch W, Mannherz HG, Suck D, Pai EF, Holmes KC (1990) Atomic structure of the actin: DNase I complex. Nature 347:37–44
Graceffa P, Dominguez R (2003) Crystal structure of monomeric actin in the ATP state. Structural basis of nucleotide-dependent actin dynamics. J Biol Chem 278:34172–34180
Steitz TA (2008) A structural understanding of the dynamic ribosome machine. Nat Rev Mol Cell Biol 9:242–253
Schmeing TM, Ramakrishnan V (2009) What recent ribosome structures have revealed about the mechanism of translation. Nature 461:1234–1242
Yonath A (2009) Large facilities and the evolving ribosome, the cellular machine for genetic-code translation. J R Soc Interface 6:S575–S585
Gabashvili IS, Agrawal RK, Spahn CMT, Grassucci RA, Svergun DI, Frank J, Penczek P (2000) Solution structure of the E. coli 70S ribosome at 11.5 Å resolution. Cell 100:537–549
Yusupov MM, Yusupova GZ, Baucom A, Lieberman K, Earnest TN, Cate JHD, Noller HF (2001) Crystal structure of the ribosome at 5.5 Å resolution. Science 292:883–896
Sanbonmatsu KY, Tung CS (2007) High performance computing in biology: multimillion atom simulations of nanoscale systems. J Struct Biol 157:470–480
Sanbonmatsu KY (2012) Computational studies of molecular machines: the ribosome. Curr Opin Struct Biol 22:168–174
Zhang Z, Sanbonmatsu KY, Voth GA (2011) Key intermolecular interactions in the E. coli 70S ribosome revealed by coarse-grained analysis. J Am Chem Soc 133:16828–16838
Lyman E, Pfaendtner J, Voth GA (2008) Systematic multiscale parameterization of heterogeneous elastic network models of proteins. Biophys J 95:4183–4192
Frank J, Agrawal RK (2000) A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406:318–322
Poornam GP, Matsumoto A, Ishida H, Hayward S (2009) A method for the analysis of domain movements in large biomolecular complexes. Proteins 76:201–212
Amadei A, Ceruso MA, Di Nola A (1999) On the convergence of the conformational coordinates basis set obtained by the essential dynamics analysis of prtoeins’ molecular dynamics simulations. Proteins Struc Func Genet 36:419–424
Sinitskiy AV, Saunders MG, Voth GA (2012) Optimal number of coarse-grained sites in different components of large biomolecular complexes. J Phys Chem B 116:8363–8374
Humphrey W, Dalke A, Schulten K (1996) VMD: Visual molecular dynamics. J Mol Graphics 14:33–38
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Shanghai Jiaotong University Press, Shanghai and Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Zhang, Z. (2015). Systematic Methods for Defining Coarse-Grained Maps in Large Biomolecules. In: Wei, D., Xu, Q., Zhao, T., Dai, H. (eds) Advance in Structural Bioinformatics. Advances in Experimental Medicine and Biology, vol 827. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-9245-5_4
Download citation
DOI: https://doi.org/10.1007/978-94-017-9245-5_4
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-9244-8
Online ISBN: 978-94-017-9245-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)