Abstract
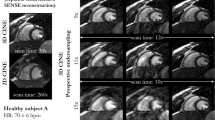
Cine cardiac MRI is routinely acquired for the assessment of cardiac health, but the imaging process is slow and typically requires several breath-holds to acquire sufficient k-space profiles to ensure good image quality. Several undersampling-based reconstruction techniques have been proposed during the last decades to speed up cine cardiac MRI acquisition. However, the undersampling factor is commonly fixed to conservative values before acquisition to ensure diagnostic image quality, potentially leading to unnecessarily long scan times. In this paper, we propose an end-to-end quality-aware cine short-axis cardiac MRI framework that combines image acquisition and reconstruction with downstream tasks such as segmentation, volume curve analysis and estimation of cardiac functional parameters. The goal is to reduce scan time by acquiring only a fraction of k-space data to enable the reconstruction of images that can pass quality control checks and produce reliable estimates of cardiac functional parameters. The framework consists of a deep learning model for the reconstruction of 2D+t cardiac cine MRI images from undersampled data, an image quality-control step to detect good quality reconstructions, followed by a deep learning model for bi-ventricular segmentation, a quality-control step to detect good quality segmentations and automated calculation of cardiac functional parameters. To demonstrate the feasibility of the proposed approach, we perform simulations using a cohort of selected participants from the UK Biobank (n = 270), 200 healthy subjects and 70 patients with cardiomyopathies. Our results show that we can produce quality-controlled images in a scan time reduced from 12 to 4 s per slice, enabling reliable estimates of cardiac functional parameters such as ejection fraction within 5% mean absolute error.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bustin, A., Fuin, N., Botnar, R.M., Prieto, C.: From compressed-sensing to artificial intelligence-based cardiac MRI reconstruction. Front. Cardiovasc. Med. 7, 17 (2020)
Menchón-Lara, R.M., Simmross-Wattenberg, F., Casaseca-de-la Higuera, P., Martín-Fernández, M., Alberola-López, C.: Reconstruction techniques for cardiac cine MRI. Insights Imaging 10(1), 1–16 (2019)
Schlemper, J., Caballero, J., Hajnal, J.V., Price, A.N., Rueckert, D.: A deep cascade of convolutional neural networks for dynamic MR image reconstruction. IEEE Trans. Med. Imaging 37(2), 491–503 (2017)
Hammernik, K., et al.: Learning a variational network for reconstruction of accelerated MRI data. Magn. Reson. Med. 79(6), 3055–3071 (2018)
Petersen, S.E., et al.: UK biobank’s cardiovascular magnetic resonance protocol. J. Cardiovasc. Magn. Reson. 18(1), 8 (2015)
Petersen, S.E., et al.: Reference ranges for cardiac structure and function using cardiovascular magnetic resonance (CMR) in caucasians from the UK biobank population cohort. J. Cardiovasc. Magn. Reson. 19(1), 1–19 (2017)
Haldar, J.P.: Low-rank modeling of local \( k \)-space neighborhoods (loraks) for constrained MRI. IEEE Trans. Med. Imaging 33(3), 668–681 (2013)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 770–778 (2016)
Chen, C., et al.: Improving the generalizability of convolutional neural network-based segmentation on CMR images. Front. Cardiovasc. Med. 7, 105 (2020)
Robinson, R., et al.: Automated quality control in image segmentation: application to the UK biobank cardiovascular magnetic resonance imaging study. J. Cardiovasc. Magn. Reson. 21(1), 1–14 (2019)
Galati, F., Zuluaga, M.A.: Efficient model monitoring for quality control in cardiac image segmentation. In: Ennis, D.B., Perotti, L.E., Wang, V.Y. (eds.) FIMH 2021. LNCS, vol. 12738, pp. 101–111. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-78710-3_11
Robinson, R., et al.: Real-time prediction of segmentation quality. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11073, pp. 578–585. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00937-3_66
Acknowledgement
This work was funded by the Engineering and Physical Sciences Research Council (EPSRC) programme grant ‘SmartHeart’ (EP/P001009/1) and supported by the Wellcome/EPSRC Centre for Medical Engineering [WT 203148/Z/16/Z]. The research was supported by the National Institute for Health Research (NIHR) Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust and King’s College London. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. This work was also supported by Health Data Research UK, an initiative funded by UK Research and Innovation, Department of Health and Social Care (England) and the devolved administrations, and leading medical research charities. This research has been conducted using the UK Biobank Resource under Application Number 17806.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this paper
Cite this paper
Machado, I. et al. (2022). Quality-Aware Cine Cardiac MRI Reconstruction and Analysis from Undersampled K-Space Data. In: Puyol Antón, E., et al. Statistical Atlases and Computational Models of the Heart. Multi-Disease, Multi-View, and Multi-Center Right Ventricular Segmentation in Cardiac MRI Challenge. STACOM 2021. Lecture Notes in Computer Science(), vol 13131. Springer, Cham. https://doi.org/10.1007/978-3-030-93722-5_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-93722-5_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-93721-8
Online ISBN: 978-3-030-93722-5
eBook Packages: Computer ScienceComputer Science (R0)